[English] 日本語
 Yorodumi
Yorodumi- EMDB-51033: Cryo-electron tomogram of AL59 amyloids interacting with collagen VI -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron tomogram of AL59 amyloids interacting with collagen VI | |||||||||||||||
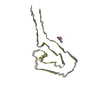
 Map data Map data | Tomogram of figure 1B, middle panel showing decorated AL59 fibril. Figure shows slice 110. Also Tomogram used for the segmentation shown in figure 6B. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | AL59 / Collagen IV / Amyloid / Protein Interaction / Light Chain Amyloidosis / PROTEIN BINDING | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | electron tomography / cryo EM | |||||||||||||||
 Authors Authors | Sicking K / Fernandez-Busnadiego R / Ricagno S | |||||||||||||||
| Funding support |  Germany, Germany,  United States, United States,  Italy, 4 items Italy, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Helical superstructures between amyloid and collagen in cardiac fibrils from a patient with AL amyloidosis. Authors: Tim Schulte / Antonio Chaves-Sanjuan / Valentina Speranzini / Kevin Sicking / Melissa Milazzo / Giulia Mazzini / Paola Rognoni / Serena Caminito / Paolo Milani / Chiara Marabelli / ...Authors: Tim Schulte / Antonio Chaves-Sanjuan / Valentina Speranzini / Kevin Sicking / Melissa Milazzo / Giulia Mazzini / Paola Rognoni / Serena Caminito / Paolo Milani / Chiara Marabelli / Alessandro Corbelli / Luisa Diomede / Fabio Fiordaliso / Luigi Anastasia / Carlo Pappone / Giampaolo Merlini / Martino Bolognesi / Mario Nuvolone / Rubén Fernández-Busnadiego / Giovanni Palladini / Stefano Ricagno /     Abstract: Systemic light chain (LC) amyloidosis (AL) is a disease where organs are damaged by an overload of a misfolded patient-specific antibody-derived LC, secreted by an abnormal B cell clone. The high LC ...Systemic light chain (LC) amyloidosis (AL) is a disease where organs are damaged by an overload of a misfolded patient-specific antibody-derived LC, secreted by an abnormal B cell clone. The high LC concentration in the blood leads to amyloid deposition at organ sites. Indeed, cryogenic electron microscopy (cryo-EM) has revealed unique amyloid folds for heart-derived fibrils taken from different patients. Here, we present the cryo-EM structure of heart-derived AL amyloid (AL59) from another patient with severe cardiac involvement. The double-layered structure displays a u-shaped core that is closed by a β-arc lid and extended by a straight tail. Noteworthy, the fibril harbours an extended constant domain fragment, thus ruling out the variable domain as sole amyloid building block. Surprisingly, the fibrils were abundantly concatenated with a proteinaceous polymer, here identified as collagen VI (COLVI) by immuno-electron microscopy (IEM) and mass-spectrometry. Cryogenic electron tomography (cryo-ET) showed how COLVI wraps around the amyloid forming a helical superstructure, likely stabilizing and protecting the fibrils from clearance. Thus, here we report structural evidence of interactions between amyloid and collagen, potentially signifying a distinct pathophysiological mechanism of amyloid deposits. #1:  Journal: Res Sq / Year: 2023 Journal: Res Sq / Year: 2023Title: Helical superstructures between amyloid and collagen VI in heart-derived fibrils from a patient with Light Chain Amyloidosis. Authors: Ricagno S / Schulte T / Chaves-Sanjuan A / Speranzini V / Sicking K / Mazzini G / Rognoni P / Caminito S / Milani P / Marabelli C / Corbelli A / Diomede L / Fiordaliso F / Anastasia L / ...Authors: Ricagno S / Schulte T / Chaves-Sanjuan A / Speranzini V / Sicking K / Mazzini G / Rognoni P / Caminito S / Milani P / Marabelli C / Corbelli A / Diomede L / Fiordaliso F / Anastasia L / Pappone C / merlini G / Bolognesi M / Nuvolone M / Fernandez-Busnadiego R / Palladini G | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_51033.map.gz emd_51033.map.gz | 739.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-51033-v30.xml emd-51033-v30.xml emd-51033.xml emd-51033.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_51033.png emd_51033.png | 207.8 KB | ||
| Filedesc metadata |  emd-51033.cif.gz emd-51033.cif.gz | 4.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-51033 http://ftp.pdbj.org/pub/emdb/structures/EMD-51033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51033 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_51033.map.gz / Format: CCP4 / Size: 800 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_51033.map.gz / Format: CCP4 / Size: 800 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tomogram of figure 1B, middle panel showing decorated AL59 fibril. Figure shows slice 110. Also Tomogram used for the segmentation shown in figure 6B. | ||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 9.25 Å | ||||||||||||||||||||||||||||||||
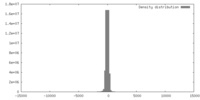
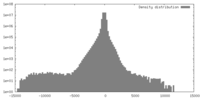
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Sample extracted from a patient of Light Chain Amyloidosis
| Entire | Name: Sample extracted from a patient of Light Chain Amyloidosis |
|---|---|
| Components |
|
-Supramolecule #1: Sample extracted from a patient of Light Chain Amyloidosis
| Supramolecule | Name: Sample extracted from a patient of Light Chain Amyloidosis type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: Heart Homo sapiens (human) / Tissue: Heart |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Details: water |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: 30mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.0 K / Instrument: FEI VITROBOT MARK IV |
| Sectioning | Other: NO SECTIONING |
| Fiducial marker | Manufacturer: Electron Microscopy Sciences / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 120.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 53000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Software - Name:  IMOD (ver. 4.11.15) / Number images used: 35 IMOD (ver. 4.11.15) / Number images used: 35 |
|---|
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
















