-検索条件
-検索結果
検索 (著者・登録者: deb & i)の結果808件中、1から50件目までを表示しています

EMDB-18137: 
HK68 cryo-EM structure achieved via rapid-spray and vitrification for grid preparation

EMDB-44552: 
Septin Hexameric Complex SEPT2/SEPT6/SEPT7 of Ciona intestinalis by Cryo-EM

EMDB-44555: 
Septin Tetrameric Complex SEPT7/SEPT9 of Ciona intestinalis by Cryo-EM

PDB-9bht: 
Septin Hexameric Complex SEPT2/SEPT6/SEPT7 of Ciona intestinalis by Cryo-EM

PDB-9bhw: 
Septin Tetrameric Complex SEPT7/SEPT9 of Ciona intestinalis by Cryo-EM

EMDB-17725: 
CryoEM reconstruction of Influenza A virus (HK68) hemagglutinin bound to an Affimer reagent
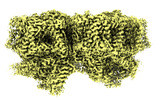
EMDB-41460: 
Structure of a mutated photosystem II complex reveals perturbation of the oxygen-evolving complex

PDB-8tow: 
Structure of a mutated photosystem II complex reveals perturbation of the oxygen-evolving complex

EMDB-18290: 
Cryo-EM structure of Cx26 gap junction K125E mutant in bicarbonate buffer (classification on hemichannel)

EMDB-18291: 
Cryo-EM structure of Cx26 solubilised in LMNG - hemichannel classification - NConst conformation

EMDB-18292: 
Cryo-EM structure of Cx26 solubilised in LMNG - Hemichannel classification NFlex conformation
EMDB-18293: 
Cryo-EM structure of Cx26 solubilised in LMNG: classification on subunit A; Nconst-mon conformation

EMDB-18294: 
Cryo-EM structure of Cx26 solubilised in LMNG: classification on subunit A; NFlex conformation

EMDB-18295: 
Cryo-EM reconstruction of Cx26 gap junction K125R mutant (D6 symmetry)
EMDB-18296: 
Cryo-EM reconstruction of Cx26 gap junction K125E mutant in HEPES buffer

EMDB-18297: 
Cryo-EM reconstruction of Cx26 gap junction WT in HEPES buffer

PDB-8q9z: 
Cryo-EM structure of Cx26 gap junction K125E mutant in bicarbonate buffer (classification on hemichannel)

PDB-8qa0: 
Cryo-EM structure of Cx26 solubilised in LMNG - hemichannel classification - NConst conformation

PDB-8qa1: 
Cryo-EM structure of Cx26 solubilised in LMNG - Hemichannel classification NFlex conformation

PDB-8qa2: 
Cryo-EM structure of Cx26 solubilised in LMNG: classification on subunit A; Nconst-mon conformation

PDB-8qa3: 
Cryo-EM structure of Cx26 solubilised in LMNG: classification on subunit A; NFlex conformation

EMDB-19014: 
PDCoV spike glycoprotein ectodomain in complex with the 22C10 antibody Fab fragment

EMDB-19015: 
Local refinement of the PDCoV spike glycoprotein ectodomain in complex with the 22C10 antibody Fab fragment

EMDB-19016: 
S1B domain of the PDCoV spike glycoprotein in complex with the 67B12 and 42H3 antibody Fab fragments

EMDB-19017: 
S1B domain of the PDCoV spike glycoprotein in complex with the 67B12 and 46E6 antibody Fab fragments

PDB-8r9w: 
PDCoV spike glycoprotein ectodomain in complex with the 22C10 antibody Fab fragment

PDB-8r9x: 
Local refinement of the PDCoV spike glycoprotein ectodomain in complex with the 22C10 antibody Fab fragment

PDB-8r9y: 
S1B domain of the PDCoV spike glycoprotein in complex with the 67B12 and 42H3 antibody Fab fragments

PDB-8r9z: 
S1B domain of the PDCoV spike glycoprotein in complex with the 67B12 and 46E6 antibody Fab fragments
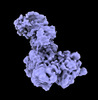
EMDB-40261: 
DDB1/CRBN in complex with ARV-471 and the ER ligand-binding domain

EMDB-41060: 
Cryo-EM structure of DRH-1 helicase and C-terminal domain bound to dsRNA

PDB-8t5s: 
Cryo-EM structure of DRH-1 helicase and C-terminal domain bound to dsRNA

EMDB-19005: 
structure of the GLMP/MFSD1 complex

EMDB-19006: 
Lysosomal peptide transporter

PDB-8r8q: 
Lysosomal peptide transporter

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement

PDB-8vye: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs

PDB-8vyf: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement

PDB-8vyg: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2

PDB-8tnp: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40

PDB-8tnq: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1

PDB-8tnr: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
