+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of Cx26 gap junction WT in HEPES buffer | |||||||||
 Map data Map data | Full map D6 averaged (unsharpened) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Gap junction Large Pore Channel Carbon dioxide sensitive / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationTransport of connexons to the plasma membrane / gap junction channel activity involved in cell communication by electrical coupling / gap junction-mediated intercellular transport / Oligomerization of connexins into connexons / Transport of connexins along the secretory pathway / gap junction assembly / connexin complex / gap junction / Gap junction assembly / gap junction channel activity ...Transport of connexons to the plasma membrane / gap junction channel activity involved in cell communication by electrical coupling / gap junction-mediated intercellular transport / Oligomerization of connexins into connexons / Transport of connexins along the secretory pathway / gap junction assembly / connexin complex / gap junction / Gap junction assembly / gap junction channel activity / endoplasmic reticulum-Golgi intermediate compartment / sensory perception of sound / transmembrane transport / cell-cell signaling / calcium ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
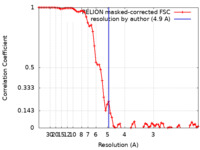
| Method | single particle reconstruction / cryo EM / Resolution: 4.9 Å | |||||||||
 Authors Authors | Brotherton DH / Savva CG / Cameron AD | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2024 Journal: Elife / Year: 2024Title: Structures of wild-type and a constitutively closed mutant of connexin26 shed light on channel regulation by CO. Authors: Deborah H Brotherton / Sarbjit Nijjar / Christos G Savva / Nicholas Dale / Alexander David Cameron /  Abstract: Connexins allow intercellular communication by forming gap junction channels (GJCs) between juxtaposed cells. Connexin26 (Cx26) can be regulated directly by CO. This is proposed to be mediated ...Connexins allow intercellular communication by forming gap junction channels (GJCs) between juxtaposed cells. Connexin26 (Cx26) can be regulated directly by CO. This is proposed to be mediated through carbamylation of K125. We show that mutating K125 to glutamate, mimicking the negative charge of carbamylation, causes Cx26 GJCs to be constitutively closed. Through cryo-EM we observe that the K125E mutation pushes a conformational equilibrium towards the channel having a constricted pore entrance, similar to effects seen on raising the partial pressure of CO. In previous structures of connexins, the cytoplasmic loop, important in regulation and where K125 is located, is disordered. Through further cryo-EM studies we trap distinct states of Cx26 and observe density for the cytoplasmic loop. The interplay between the position of this loop, the conformations of the transmembrane helices and the position of the N-terminal helix, which controls the aperture to the pore, provides a mechanism for regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18297.map.gz emd_18297.map.gz | 22.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18297-v30.xml emd-18297-v30.xml emd-18297.xml emd-18297.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18297_fsc.xml emd_18297_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_18297.png emd_18297.png | 76 KB | ||
| Filedesc metadata |  emd-18297.cif.gz emd-18297.cif.gz | 5.3 KB | ||
| Others |  emd_18297_half_map_1.map.gz emd_18297_half_map_1.map.gz emd_18297_half_map_2.map.gz emd_18297_half_map_2.map.gz | 22.3 MB 22.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18297 http://ftp.pdbj.org/pub/emdb/structures/EMD-18297 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18297 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18297 | HTTPS FTP |
-Related structure data
| Related structure data |  8q9zC  8qa0C  8qa1C  8qa2C  8qa3C C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18297.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18297.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map D6 averaged (unsharpened) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map D6 averaged
| File | emd_18297_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map D6 averaged | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map D6 averaged
| File | emd_18297_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map D6 averaged | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Connexin 26 WT in HEPES buffer pH 8.0
| Entire | Name: Connexin 26 WT in HEPES buffer pH 8.0 |
|---|---|
| Components |
|
-Supramolecule #1: Connexin 26 WT in HEPES buffer pH 8.0
| Supramolecule | Name: Connexin 26 WT in HEPES buffer pH 8.0 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Connexin 26
| Macromolecule | Name: Connexin 26 / type: protein_or_peptide / ID: 1 / Details: Connexin subunit / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MDWGTLQTIL GGVNKHSTSI GKIWLTVLFI FRIMILVVAA KEVWGDEQAD FVCNTLQPGC KNVCYDHYFP IS HIRLWAL QLIFVSTPAL LVAMHVAYRR HEKKRKFIKG EIKSEFKDIE EIKTQKVRIE GSLWWTYTSS IFFRV IFEA AFMYVFYVMY DGFSMQRLVK ...String: MDWGTLQTIL GGVNKHSTSI GKIWLTVLFI FRIMILVVAA KEVWGDEQAD FVCNTLQPGC KNVCYDHYFP IS HIRLWAL QLIFVSTPAL LVAMHVAYRR HEKKRKFIKG EIKSEFKDIE EIKTQKVRIE GSLWWTYTSS IFFRV IFEA AFMYVFYVMY DGFSMQRLVK CNAWPCPNTV DCFVSRPTEK TVFTVFMIAV SGICILLNVT ELCYLLIR Y CSGKSKKPVL VPR UniProtKB: Gap junction beta-2 protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: The buffer, except DDM and DTT, was prepared fresh from 10x stock on day of used. The basal buffer was filtered and degassed and DDM and DTT added. | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: LEICA EM GP Details: 3 microlitres protein applied to grid, blot time 3 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE / Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number real images: 2436 / Average exposure time: 60.0 sec. / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.7 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)