+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lysosomal peptide transporter | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MFS transporter / MFSD family / nutrition transporter / lysosomal transporter / outward open / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdipeptide transmembrane transport from lysosomal lumen to cytosol / dipeptide uniporter activity / protein localization to lysosome / lysosome / protein stabilization / lysosomal membrane / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / membrane / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.08 Å | |||||||||
 Authors Authors | Jungnickel KEJ / Loew C | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Cell Biol / Year: 2024 Journal: Nat Cell Biol / Year: 2024Title: MFSD1 with its accessory subunit GLMP functions as a general dipeptide uniporter in lysosomes. Authors: Katharina Esther Julia Jungnickel / Océane Guelle / Miharu Iguchi / Wentao Dong / Vadim Kotov / Florian Gabriel / Cécile Debacker / Julien Dairou / Isabelle McCort-Tranchepain / Nouf N ...Authors: Katharina Esther Julia Jungnickel / Océane Guelle / Miharu Iguchi / Wentao Dong / Vadim Kotov / Florian Gabriel / Cécile Debacker / Julien Dairou / Isabelle McCort-Tranchepain / Nouf N Laqtom / Sze Ham Chan / Akika Ejima / Kenji Sato / David Massa López / Paul Saftig / Ahmad Reza Mehdipour / Monther Abu-Remaileh / Bruno Gasnier / Christian Löw / Markus Damme /      Abstract: The lysosomal degradation of macromolecules produces diverse small metabolites exported by specific transporters for reuse in biosynthetic pathways. Here we deorphanized the major facilitator ...The lysosomal degradation of macromolecules produces diverse small metabolites exported by specific transporters for reuse in biosynthetic pathways. Here we deorphanized the major facilitator superfamily domain containing 1 (MFSD1) protein, which forms a tight complex with the glycosylated lysosomal membrane protein (GLMP) in the lysosomal membrane. Untargeted metabolomics analysis of MFSD1-deficient mouse lysosomes revealed an increase in cationic dipeptides. Purified MFSD1 selectively bound diverse dipeptides, while electrophysiological, isotope tracer and fluorescence-based studies in Xenopus oocytes and proteoliposomes showed that MFSD1-GLMP acts as a uniporter for cationic, neutral and anionic dipeptides. Cryoelectron microscopy structure of the dipeptide-bound MFSD1-GLMP complex in outward-open conformation characterized the heterodimer interface and, in combination with molecular dynamics simulations, provided a structural basis for its selectivity towards diverse dipeptides. Together, our data identify MFSD1 as a general lysosomal dipeptide uniporter, providing an alternative route to recycle lysosomal proteolysis products when lysosomal amino acid exporters are overloaded. | |||||||||
| History |
|
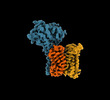
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19006.map.gz emd_19006.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19006-v30.xml emd-19006-v30.xml emd-19006.xml emd-19006.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19006.png emd_19006.png | 50.4 KB | ||
| Filedesc metadata |  emd-19006.cif.gz emd-19006.cif.gz | 6.8 KB | ||
| Others |  emd_19006_half_map_1.map.gz emd_19006_half_map_1.map.gz emd_19006_half_map_2.map.gz emd_19006_half_map_2.map.gz | 95.6 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19006 http://ftp.pdbj.org/pub/emdb/structures/EMD-19006 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19006 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19006 | HTTPS FTP |
-Related structure data
| Related structure data |  8r8qMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19006.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19006.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_19006_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19006_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GLMP-MFSD1
| Entire | Name: GLMP-MFSD1 |
|---|---|
| Components |
|
-Supramolecule #1: GLMP-MFSD1
| Supramolecule | Name: GLMP-MFSD1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Major facilitator superfamily domain-containing protein 1
| Macromolecule | Name: Major facilitator superfamily domain-containing protein 1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 55.765551 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEDEDGEDRA LLGGRREADS AVHGAPRALS ALCDPSRLAH RLVVLSLMCF LGFGSYFCYD NPAALQTQVK RDMQVNTTKF MLLYAWYSW PNVVLCFLGG FLIDRIFGIR WGTVIFSCFV CIGQVIFALG GIFNAFWLME LGRFVFGIGG ESLAVAQNTY A VSWFKGKE ...String: MEDEDGEDRA LLGGRREADS AVHGAPRALS ALCDPSRLAH RLVVLSLMCF LGFGSYFCYD NPAALQTQVK RDMQVNTTKF MLLYAWYSW PNVVLCFLGG FLIDRIFGIR WGTVIFSCFV CIGQVIFALG GIFNAFWLME LGRFVFGIGG ESLAVAQNTY A VSWFKGKE LNLVFGLQLS MARIGSTVNM NLMGWLYGKI EALLGSAGHM TLGVTLMIGC ITCIFSLICA LALAYLDRRA EK ILHKEQG KTGEVIKLRD IKDFSLPLIL VFVICVCYYV AVFPFIGLGK VFFMEKFRFS SQSASAINSI VYIISAPMSP LFG LLVDKT GKNIIWVLYA VAATLVSHMM LAFTFWNPWI AMCLLGFSYS LLACALWPMV AFIVPEHQLG TAYGFMQSIQ NLGL AVIAI LAGMILDSKG YLLLEVFFIA CVSLSLLAVV CLYLVNRAQG GNLNYSAKQR ERMKLSHPEI IILEVLFQGP SSGWS HPQF EKGGGSGGGS GGSAWSHPQF EK UniProtKB: Lysosomal dipeptide transporter MFSD1 |
-Macromolecule #2: Glycosylated lysosomal membrane protein
| Macromolecule | Name: Glycosylated lysosomal membrane protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.102711 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFRCWGPHWG WVPCAPTPWL LLSLLVCSAP FGLQGEETRQ VSMEVISGWP NPQNLLHIRA VGSNSTLHYV WSSLGPPAVV LVATNTTQS VLSVNWSLLL SPDPAGALMV LPKSSIQFSS ALVFTRLLEF DSTNASEGAQ PPGKPYPPYS LAKFSWNNIT N SLDLANLS ...String: MFRCWGPHWG WVPCAPTPWL LLSLLVCSAP FGLQGEETRQ VSMEVISGWP NPQNLLHIRA VGSNSTLHYV WSSLGPPAVV LVATNTTQS VLSVNWSLLL SPDPAGALMV LPKSSIQFSS ALVFTRLLEF DSTNASEGAQ PPGKPYPPYS LAKFSWNNIT N SLDLANLS ADFQGRPVDD PTGAFANGSL TFKVQAFSRS GRPAQPPRLL HTADVCQLEV ALVGASPRGN HSLFGLEVAT LG QGPDCPS VNERNSIDDE YAPAVFQLNQ LLWGSSPSGF MQWRPVAFSE EERARESALP CQASTLHSTL ASSLPHSPIV QAF FGSQNN FCAFNLTFGA PTGPGYWDQY YLCWSMLLGM GFPPVDIFSP LVLGIMAVAL GAPGLMFLGG GLFLLGSAGS AAGS GEF UniProtKB: Glycosylated lysosomal membrane protein |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.33 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. | |||||||||||||||
| Vitrification | Cryogen name: PROPANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 3193 / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
| Output model |  PDB-8r8q: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)