-Search query
-Search result
Showing 1 - 50 of 51 items for (author: ahel & j)

EMDB-50229: 
Cryo-tomogram of FIB-milled vegetatively growing yeast cell with mitochondria

EMDB-50230: 
Cryo-tomogram of FIB-milled pre-meiotic yeast cell with mitochondria

EMDB-50231: 
Cryo-tomogram of FIB-milled meiotic yeast cell containing mitochondria with filaments
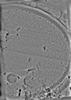
EMDB-50232: 
Cryo-tomogram of FIB-milled yeast spore with mitochondria

EMDB-50233: 
Cryo-tomogram of FIB-milled meiotic yeast cell containing mitochondria with filament arrays
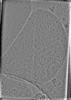

EMDB-50234: 
Cryo-tomogram of purified meiotic yeast mitochondria with Ald4 filaments

EMDB-19548: 
cryoEM structure of Acs1 filament determined by FilamentID

EMDB-19549: 
cryoEM structure of the central Ald4 filament determined by FilamentID

EMDB-19550: 
cryoEM structure of purified Acs1 filament from meiotic yeast cells
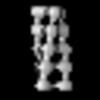
EMDB-19551: 
cryo sub-tomogram average of Acs1 filament from spread meiotic yeast spheroplasts
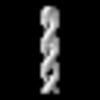
EMDB-19552: 
cryo sub-tomogram average of Ald4 filaments from purified meiotic yeast mitochondria
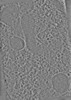
EMDB-19553: 
cryo-tomogram of FIB-milled meiotic yeast cell containing filaments in mitochondria
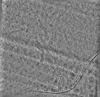

EMDB-19554: 
cryo-tomogram of FIB-milled meiotic yeast cell containing filaments in the nucleus
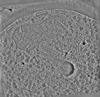
EMDB-19555: 
cryo-tomogram of FIB-milled meiotic yeast cell containing filaments in the cytoplasm
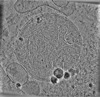
EMDB-19556: 
cryo-tomogram of purified meiotic yeast mitochondrion containing Ald4 filaments
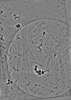
EMDB-19557: 
cryo-tomogram of spread meiotic yeast spheroplast containing Acs1 filaments

EMDB-19558: 
cryo-tomogram of spread starved yeast spheroplast containing filaments

PDB-8rwj: 
cryoEM structure of Acs1 filament determined by FilamentID

PDB-8rwk: 
cryoEM structure of the central Ald4 filament determined by FilamentID

EMDB-15365: 
Vairimorpha necatrix 20S proteasome from spores

EMDB-15366: 
Vairimorpha necatrix 26S proteasome from sporoplasms

EMDB-15367: 
Vairimorpha necatrix 20S proteasome from sporoplasms

PDB-8adn: 
Vairimorpha necatrix 20S proteasome from spores

EMDB-12931: 
Mouse RNF213:UBE2L3 transthiolation intermediate, chemically stabilized

EMDB-12932: 
Mouse RNF213, with mixed nucleotides bound

PDB-7oik: 
Mouse RNF213:UBE2L3 transthiolation intermediate, chemically stabilized

PDB-7oim: 
Mouse RNF213, with mixed nucleotides bound

EMDB-24444: 
Mouse GITR (mGITR) with DTA-1 Fab fragment

PDB-7rfp: 
Mouse GITR (mGITR) with DTA-1 Fab fragment

EMDB-21981: 
PARP2/HPF1_Nucleosome complex

EMDB-21982: 
PARP2/HPF1_Nucleosome complex

EMDB-21970: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

EMDB-21971: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

EMDB-21978: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

EMDB-21979: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

EMDB-21980: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

PDB-6wz5: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

PDB-6wz9: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

PDB-6x0l: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

PDB-6x0m: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

PDB-6x0n: 
Bridging of double-strand DNA break activates PARP2/HPF1 to modify chromatin

EMDB-10429: 
Mouse RNF213 wild type protein

EMDB-10430: 
Mouse RNF213 mutant R4753K modeling the Moyamoya-disease-related Human variant R4810K

PDB-6tax: 
Mouse RNF213 wild type protein

PDB-6tay: 
Mouse RNF213 mutant R4753K modeling the Moyamoya-disease-related Human variant R4810K

EMDB-20645: 
MicroED structure of a FIB-milled CypA Crystal

PDB-6u5g: 
MicroED structure of a FIB-milled CypA Crystal

EMDB-20628: 
Volume 1 for truncated dimeric Cytohesin-3 (Grp1; amino acids 14-399)

EMDB-20629: 
Best fitting antiparallel model for Volume 2 of truncated dimeric Cytohesin-3 (Grp1; amino acids 14-399)

PDB-6u3e: 
Best fitting antiparallel model for Volume 1 of truncated dimeric Cytohesin-3 (Grp1; amino acids 14-399)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
