7RV1
 
 | | Crystal structure of the BCL6 BTB domain in complex with OICR-8826 | | Descriptor: | (E)-3-(7-(2-((3-chloropyridin-4-yl)amino)-2-oxoethyl)-3-(3-(1-methyl-1H-pyrazol-4-yl)prop-2-yn-1-yl)-4-oxo-4,7-dihydro-3H-pyrrolo[2,3-d]pyrimidin-5-yl)acrylamide, B-cell lymphoma 6 protein, DIMETHYL SULFOXIDE, ... | | Authors: | Kuntz, D.A, Prive, G.G. | | Deposit date: | 2021-08-18 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | Crystal structure of the BCL6 BTB domain in complex with OICR-8826
To Be Published
|
|
7RUY
 
 | | Crystal structure of the BCL6 BTB domain in complex with OICR-8388 | | Descriptor: | B-cell lymphoma 6 protein, FORMIC ACID, N-(3-chloropyridin-4-yl)-2-(2-{[(2,4-dimethoxyphenyl)methyl]amino}-4-oxo-3,4-dihydro-7H-pyrrolo[2,3-d]pyrimidin-7-yl)acetamide | | Authors: | Kuntz, D.A, Prive, G.G. | | Deposit date: | 2021-08-18 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Crystal structure of the BCL6 BTB domain in complex with OICR-8388
To Be Published
|
|
7T0U
 
 | | Crystal structure of the BCL6 BTB domain in complex with OICR-12387 | | Descriptor: | 3-chloro-5-{7-[2-({5-chloro-2-[(2S)-2-methyl-4-(oxetan-3-yl)piperazin-1-yl]pyridin-4-yl}amino)-2-oxoethyl]-3-methyl-4-oxo-2-(trifluoromethyl)-4,7-dihydro-3H-pyrrolo[2,3-d]pyrimidin-5-yl}-2-hydroxybenzamide, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Kuntz, D.A, Prive, G.G. | | Deposit date: | 2021-11-30 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Crystal structure of the BCL6 BTB domain in complex with OICR-12387
To Be Published
|
|
7T0T
 
 | | Crystal structure of the BCL6 BTB domain in complex with OICR-10562 | | Descriptor: | DIMETHYL SULFOXIDE, Isoform 2 of B-cell lymphoma 6 protein, N-[5-chloro-2-(morpholin-4-yl)pyridin-4-yl]-2-[5-(3-cyano-4-hydroxy-5-methylphenyl)-3-methyl-2-(1-methyl-1H-pyrazol-4-yl)-4-oxo-3,4-dihydro-7H-pyrrolo[2,3-d]pyrimidin-7-yl]acetamide | | Authors: | Kuntz, D.A, Prive, G.G. | | Deposit date: | 2021-11-30 | | Release date: | 2022-12-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the BCL6 BTB domain in complex with OICR-10562
To Be Published
|
|
7T58
 
 | | Crystal structure of Miz1 BTB domain | | Descriptor: | DIMETHYL SULFOXIDE, Zinc finger and BTB domain-containing protein 17 | | Authors: | Linhares, B.M, Cierpicki, T. | | Deposit date: | 2021-12-11 | | Release date: | 2022-12-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.05333114 Å) | | Cite: | Prediction of BTB domain ligandability guided by protein dynamics
To Be Published
|
|
7T0S
 
 | | Crystal structure of the BCL6 BTB domain in complex with OICR-11864 | | Descriptor: | 3-chloro-5-{1-[2-({5-chloro-2-[(3S)-3-methylmorpholin-4-yl]pyridin-4-yl}amino)-2-oxoethyl]-4-fluoro-1H-pyrrolo[2,3-b]pyridin-3-yl}-2-hydroxybenzamide, CHLORIDE ION, GLYCEROL, ... | | Authors: | Kuntz, D.A, Prive, G.G. | | Deposit date: | 2021-11-30 | | Release date: | 2022-12-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal structure of the BCL6 BTB domain in complex with OICR-11864
To Be Published
|
|
6I2M
 
 | |
5NLB
 
 | | Crystal structure of human CUL3 N-terminal domain bound to KEAP1 BTB and 3-box | | Descriptor: | Cullin-3, Kelch-like ECH-associated protein 1 | | Authors: | Adamson, R, Krojer, T, Pinkas, D.M, Bartual, S.G, Burgess-Brown, N.A, Borkowska, O, Chalk, R, Newman, J.A, Kopec, J, Dixon-Clarke, S.E, Mathea, S, Sethi, R, Velupillai, S, Mackinnon, S, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Bullock, A. | | Deposit date: | 2017-04-04 | | Release date: | 2017-04-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.45 Å) | | Cite: | Structural and biochemical characterization establishes a detailed understanding of KEAP1-CUL3 complex assembly.
Free Radic Biol Med, 204, 2023
|
|
4HXI
 
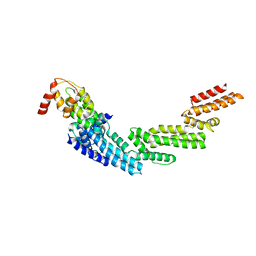 | | Crystal structure of KLHL3/Cul3 complex | | Descriptor: | Cullin-3, Kelch-like protein 3 | | Authors: | Ji, A.X, Prive, G.G. | | Deposit date: | 2012-11-10 | | Release date: | 2013-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.513 Å) | | Cite: | Crystal structure of KLHL3 in complex with Cullin3.
Plos One, 8, 2013
|
|
7MK2
 
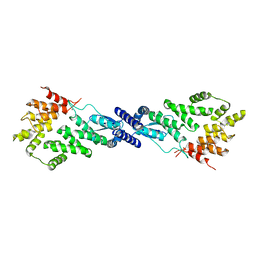 | | CryoEM Structure of NPR1 | | Descriptor: | Regulatory protein NPR1, ZINC ION | | Authors: | Kumar, S, Zhou, Y, Dillard, L, Borgnia, M, Bartesaghi, A, Zhou, P. | | Deposit date: | 2021-04-21 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural basis of NPR1 in activating plant immunity.
Nature, 605, 2022
|
|
7MK3
 
 | | Crystal structure of NPR1 | | Descriptor: | CHLORIDE ION, GLYCEROL, Regulatory protein NPR1, ... | | Authors: | Cheng, J, Wu, Q, Zhou, P. | | Deposit date: | 2021-04-21 | | Release date: | 2022-03-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | Structural basis of NPR1 in activating plant immunity.
Nature, 605, 2022
|
|
4AP2
 
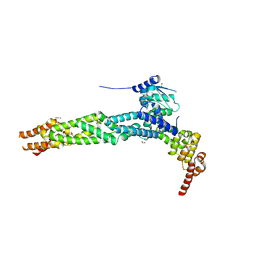 | | Crystal structure of the human KLHL11-Cul3 complex at 2.8A resolution | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, CULLIN-3, ... | | Authors: | Canning, P, Cooper, C.D.O, Krojer, T, Filippakopoulos, P, Ayinampudi, V, Arrowsmith, C.H, Edwards, A.M, Bountra, C, von Delft, F, Bullock, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2012-03-30 | | Release date: | 2012-05-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Basis for Cul3 Assembly with the Btb-Kelch Family of E3 Ubiquitin Ligases.
J.Biol.Chem., 288, 2013
|
|
4APF
 
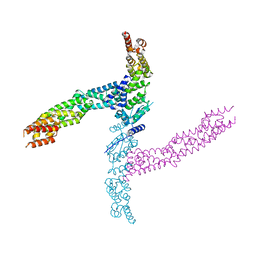 | | Crystal structure of the human KLHL11-Cul3 complex at 3.1A resolution | | Descriptor: | CULLIN 3, KELCH-LIKE PROTEIN 11 | | Authors: | Canning, P, Cooper, C.D.O, Krojer, T, Vollmar, M, Ugochukwu, E, Muniz, J.R.C, Ayinampudi, V, Savitsky, P, Arrowsmith, C.H, Edwards, A.M, Bountra, C, von Delft, F, Bullock, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2012-04-02 | | Release date: | 2012-05-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Basis for Cul3 Assembly with the Btb-Kelch Family of E3 Ubiquitin Ligases.
J.Biol.Chem., 288, 2013
|
|
8DWT
 
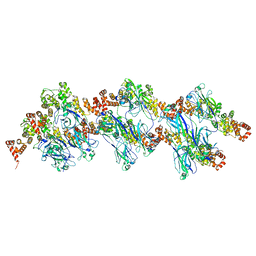 | | SPOP W22R Form 2 | | Descriptor: | Speckle-type POZ protein | | Authors: | Cuneo, M.J, Mittag, T, O'Flynn, B, Lo, Y.H. | | Deposit date: | 2022-08-02 | | Release date: | 2023-01-18 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (6.2 Å) | | Cite: | Higher-order SPOP assembly reveals a basis for cancer mutant dysregulation.
Mol.Cell, 83, 2023
|
|
8DWV
 
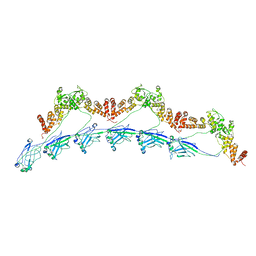 | | Full-length wild type SPOP | | Descriptor: | Speckle-type POZ protein | | Authors: | Cuneo, M.J, Mittag, T, O'Flynn, B, Lo, Y.H. | | Deposit date: | 2022-08-02 | | Release date: | 2023-01-18 | | Last modified: | 2023-03-22 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Higher-order SPOP assembly reveals a basis for cancer mutant dysregulation.
Mol.Cell, 83, 2023
|
|
8DWS
 
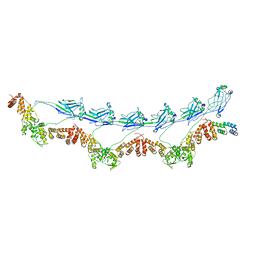 | | Full-length E47K SPOP | | Descriptor: | Speckle-type POZ protein | | Authors: | Cuneo, M.J, Mittag, T, O'Flynn, B, Lo, Y.H. | | Deposit date: | 2022-08-02 | | Release date: | 2023-01-18 | | Last modified: | 2023-03-22 | | Method: | ELECTRON MICROSCOPY (3.73 Å) | | Cite: | Higher-order SPOP assembly reveals a basis for cancer mutant dysregulation.
Mol.Cell, 83, 2023
|
|
8DWU
 
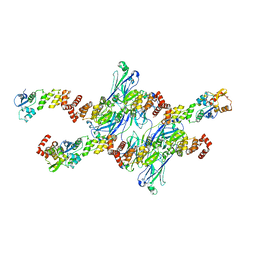 | | SPOP W22R Form 1 | | Descriptor: | Speckle-type POZ protein | | Authors: | Cuneo, M.J, Mittag, T, O'Flynn, B. | | Deposit date: | 2022-08-02 | | Release date: | 2023-01-18 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Higher-order SPOP assembly reveals a basis for cancer mutant dysregulation.
Mol.Cell, 83, 2023
|
|
3HU6
 
 | |
3I3N
 
 | | Crystal structure of the BTB-BACK domains of human KLHL11 | | Descriptor: | CHLORIDE ION, Kelch-like protein 11, THIOCYANATE ION | | Authors: | Murray, J.W, Cooper, C.D.O, Krojer, T, Mahajan, P, Salah, E, Keates, T, Savitsky, P, Pike, A.C.W, Roos, A, Muniz, J, von Delft, F, Bountra, C, Arrowsmith, C.H, Weigelt, J, Edwards, A, Knapp, S, Bullock, A, Structural Genomics Consortium (SGC) | | Deposit date: | 2009-06-30 | | Release date: | 2009-08-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the BTB-BACK domains of human KLHL11
To be Published
|
|
3HVE
 
 | |
3HQI
 
 | |
4EOZ
 
 | |
8KHP
 
 | | CULLIN3-KLHL22-RBX1 E3 ligase | | Descriptor: | Cullin-3, E3 ubiquitin-protein ligase RBX1, Kelch-like protein 22 | | Authors: | Su, M.-Y, Su, M.-Y. | | Deposit date: | 2023-08-22 | | Release date: | 2023-12-13 | | Method: | ELECTRON MICROSCOPY (3.67 Å) | | Cite: | Cryo-EM structure of the KLHL22 E3 ligase bound to an oligomeric metabolic enzyme.
Structure, 31, 2023
|
|
8K8T
 
 | | Structure of CUL3-RBX1-KLHL22 complex | | Descriptor: | Cullin-3, Kelch-like protein 22 | | Authors: | Wang, W, Ling, L, Dai, Z, Zuo, P, Yin, Y. | | Deposit date: | 2023-07-31 | | Release date: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | A conserved N-terminal motif of CUL3 contributes to assembly and E3 ligase activity of CRL3 KLHL22.
Nat Commun, 15, 2024
|
|
8K9I
 
 | | Structure of CUL3-RBX1-KLHL22 complex without CUL3 NA motif | | Descriptor: | Cullin-3, E3 ubiquitin-protein ligase RBX1, N-terminally processed, ... | | Authors: | Wang, W, Ling, L, Dai, Z, Zuo, P, Yin, Y. | | Deposit date: | 2023-08-01 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | A conserved N-terminal motif of CUL3 contributes to assembly and E3 ligase activity of CRL3 KLHL22.
Nat Commun, 15, 2024
|
|
