1ZIF
 
 | | GAAA RNA TETRALOOP, NMR, 10 STRUCTURES | | Descriptor: | RNA (5'-R(*GP*GP*GP*CP*GP*AP*AP*AP*GP*CP*CP*U)-3') | | Authors: | Jucker, F.M, Heus, H.A, Yip, P.F, Moors, E, Pardi, A. | | Deposit date: | 1996-07-27 | | Release date: | 1997-03-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A network of heterogeneous hydrogen bonds in GNRA tetraloops.
J.Mol.Biol., 264, 1996
|
|
1ZIG
 
 | | GAGA RNA TETRALOOP, NMR, 10 STRUCTURES | | Descriptor: | RNA (5'-R(*GP*GP*GP*CP*GP*AP*GP*AP*GP*CP*CP*U)-3') | | Authors: | Jucker, F.M, Heus, H.A, Yip, P.F, Moors, E, Pardi, A. | | Deposit date: | 1996-07-27 | | Release date: | 1997-03-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A network of heterogeneous hydrogen bonds in GNRA tetraloops.
J.Mol.Biol., 264, 1996
|
|
1ZIH
 
 | | GCAA RNA TETRALOOP, NMR, 10 STRUCTURES | | Descriptor: | RNA (5'-R(*GP*GP*GP*CP*GP*CP*AP*AP*GP*CP*CP*U)-3') | | Authors: | Jucker, F.M, Heus, H.A, Yip, P.F, Moors, E, Pardi, A. | | Deposit date: | 1996-07-27 | | Release date: | 1997-03-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A network of heterogeneous hydrogen bonds in GNRA tetraloops.
J.Mol.Biol., 264, 1996
|
|
1CNT
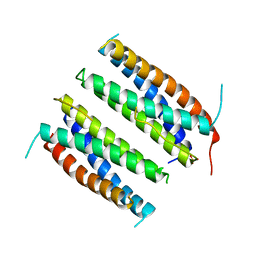 
 | | CILIARY NEUROTROPHIC FACTOR | | Descriptor: | CILIARY NEUROTROPHIC FACTOR, SULFATE ION, YTTERBIUM (III) ION | | Authors: | Mcdonald, N.Q, Panayotatos, N, Hendrickson, W.A. | | Deposit date: | 1996-06-06 | | Release date: | 1997-03-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of dimeric human ciliary neurotrophic factor determined by MAD phasing.
EMBO J., 14, 1995
|
|
1BTO
 | |
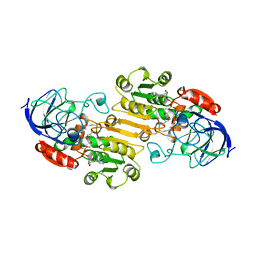
4DPV
 
 | | PARVOVIRUS/DNA COMPLEX | | Descriptor: | DNA (5'-D(*AP*TP*AP*CP*CP*TP*CP*TP*TP*GP*C)-3'), MAGNESIUM ION, PROTEIN (PARVOVIRUS COAT PROTEIN) | | Authors: | Chapman, M.S, Rossmann, M.G. | | Deposit date: | 1996-02-01 | | Release date: | 1997-04-01 | | Last modified: | 2023-04-19 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Canine parvovirus capsid structure, analyzed at 2.9 A resolution.
J.Mol.Biol., 264, 1996
|
|
1ACW
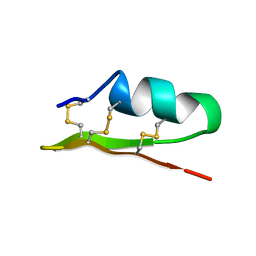 
 | | SOLUTION NMR STRUCTURE OF P01, A NATURAL SCORPION PEPTIDE STRUCTURALLY ANALOGOUS TO SCORPION TOXINS SPECIFIC FOR APAMIN-SENSITIVE POTASSIUM CHANNEL, 25 STRUCTURES | | Descriptor: | NATURAL SCORPION PEPTIDE P01 | | Authors: | Blanc, E, Fremont, V, Sizun, P, Meunier, S, Van Rietschoten, J, Thevand, A, Bernassau, J.M, Darbon, H. | | Deposit date: | 1997-02-10 | | Release date: | 1997-04-01 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of P01, a natural scorpion peptide structurally analogous to scorpion toxins specific for apamin-sensitive potassium channel.
Proteins, 24, 1996
|
|
1COF
 
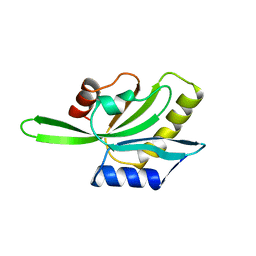 | | YEAST COFILIN, ORTHORHOMBIC CRYSTAL FORM | | Descriptor: | COFILIN | | Authors: | Fedorov, A.A, Lappalainen, P, Fedorov, E.V, Drubin, D.G, Almo, S.C. | | Deposit date: | 1996-11-26 | | Release date: | 1997-04-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure determination of yeast cofilin.
Nat.Struct.Biol., 4, 1997
|
|
1CZI
 
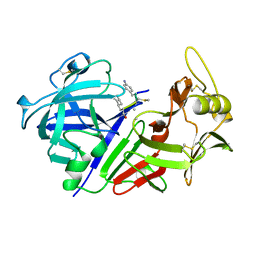 | | CHYMOSIN COMPLEX WITH THE INHIBITOR CP-113972 | | Descriptor: | CHYMOSIN, CP-113972 (NORSTATINE-S-METHYL CYSTEINE-IODO-PHENYLALANINE-PROLINE) | | Authors: | Groves, M.R, Dhanaraj, V, Pitts, J.E, Badasso, M, Hoover, D, Nugent, P, Blundell, T.L. | | Deposit date: | 1997-01-15 | | Release date: | 1997-04-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A 2.3 A resolution structure of chymosin complexed with a reduced bond inhibitor shows that the active site beta-hairpin flap is rearranged when compared with the native crystal structure.
Protein Eng., 11, 1998
|
|
1AB3
 
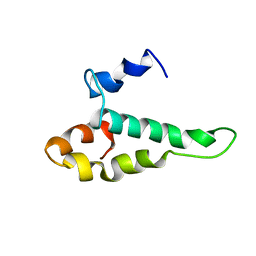 | | RIBOSOMAL PROTEIN S15 FROM THERMUS THERMOPHILUS, NMR, 26 STRUCTURES | | Descriptor: | RIBOSOMAL RNA BINDING PROTEIN S15 | | Authors: | Berglund, H, Rak, A, Serganov, A, Garber, M, Hard, T. | | Deposit date: | 1997-02-03 | | Release date: | 1997-04-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the ribosomal RNA binding protein S15 from Thermus thermophilus.
Nat.Struct.Biol., 4, 1997
|
|
1SRO
 
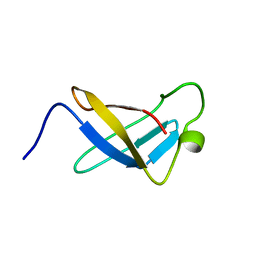 | | S1 RNA BINDING DOMAIN, NMR, 20 STRUCTURES | | Descriptor: | PNPASE | | Authors: | Bycroft, M. | | Deposit date: | 1996-11-27 | | Release date: | 1997-04-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the S1 RNA binding domain: a member of an ancient nucleic acid-binding fold.
Cell(Cambridge,Mass.), 88, 1997
|
|
1LYY
 
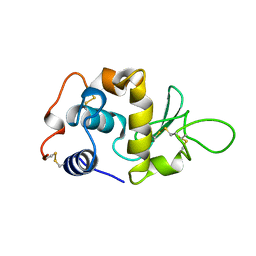 | |
1AEL
 
 | |
2HIU
 
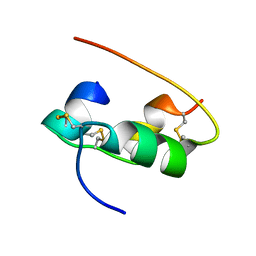 | | NMR STRUCTURE OF HUMAN INSULIN IN 20% ACETIC ACID, ZINC-FREE, 10 STRUCTURES | | Descriptor: | INSULIN | | Authors: | Hua, Q.X, Gozani, S.N, Chance, R.E, Hoffmann, J.A, Frank, B.H, Weiss, M.A. | | Deposit date: | 1996-10-08 | | Release date: | 1997-04-01 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Structure of a protein in a kinetic trap.
Nat.Struct.Biol., 2, 1995
|
|
2MBR
 
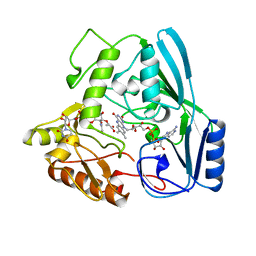 | | MURB WILD TYPE, COMPLEX WITH ENOLPYRUVYL-UDP-N-ACETYLGLUCOSAMINE | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, URIDINE DIPHOSPHO-N-ACETYLENOLPYRUVYLGLUCOSAMINE REDUCTASE, URIDINE-DIPHOSPHATE-2(N-ACETYLGLUCOSAMINYL) BUTYRIC ACID | | Authors: | Benson, T.E, Walsh, C.T, Hogle, J.M. | | Deposit date: | 1996-11-08 | | Release date: | 1997-04-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of the S229A mutant and wild-type MurB in the presence of the substrate enolpyruvyl-UDP-N-acetylglucosamine at 1.8-A resolution.
Biochemistry, 36, 1997
|
|
1IKD
 
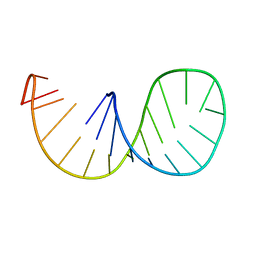 | | ACCEPTOR STEM, NMR, 30 STRUCTURES | | Descriptor: | TRNA ALA ACCEPTOR STEM | | Authors: | Ramos, A, Varani, G. | | Deposit date: | 1996-11-15 | | Release date: | 1997-04-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the acceptor stem of Escherichia coli tRNA Ala: role of the G3.U70 base pair in synthetase recognition.
Nucleic Acids Res., 25, 1997
|
|
1KZU
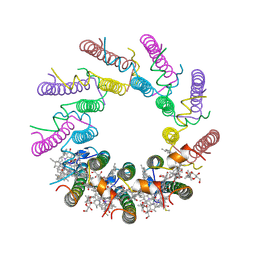 
 | | INTEGRAL MEMBRANE PERIPHERAL LIGHT HARVESTING COMPLEX FROM RHODOPSEUDOMONAS ACIDOPHILA STRAIN 10050 | | Descriptor: | BACTERIOCHLOROPHYLL A, LIGHT HARVESTING PROTEIN B-800/850, Rhodopin b-D-glucoside | | Authors: | Cogdell, R.J, Freer, A.A, Isaacs, N.W, Hawthornthwaite-Lawless, A.M, Mcdermott, G, Papiz, M.Z, Prince, S.M. | | Deposit date: | 1996-08-31 | | Release date: | 1997-04-01 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Apoprotein structure in the LH2 complex from Rhodopseudomonas acidophila strain 10050: modular assembly and protein pigment interactions.
J.Mol.Biol., 268, 1997
|
|
2TPL
 
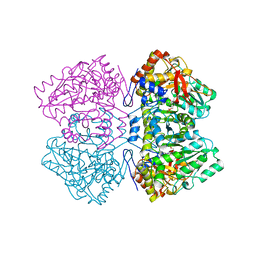 | | TYROSINE PHENOL-LYASE FROM CITROBACTER INTERMEDIUS COMPLEX WITH 3-(4'-HYDROXYPHENYL)PROPIONIC ACID, PYRIDOXAL-5'-PHOSPHATE AND CS+ ION | | Descriptor: | CESIUM ION, HYDROXYPHENYL PROPIONIC ACID, TYROSINE PHENOL-LYASE | | Authors: | Antson, A.A, Demidkina, T.V, Wilson, K.S. | | Deposit date: | 1997-01-23 | | Release date: | 1997-04-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of Citrobacter freundii tyrosine phenol-lyase complexed with 3-(4'-hydroxyphenyl)propionic acid, together with site-directed mutagenesis and kinetic analysis, demonstrates that arginine 381 is required for substrate specificity.
Biochemistry, 36, 1997
|
|
1WSA
 
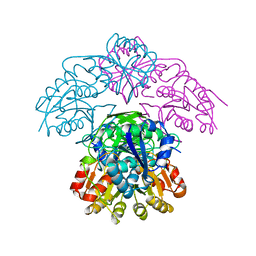 | | STRUCTURE OF L-ASPARAGINASE II PRECURSOR | | Descriptor: | ASPARAGINE AMIDOHYDROLASE | | Authors: | Lubkowski, J, Palm, G.J, Gilliland, G.L, Derst, C, Rohm, K.-H, Wlodawer, A. | | Deposit date: | 1996-08-15 | | Release date: | 1997-04-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure and amino acid sequence of Wolinella succinogenes L-asparaginase.
Eur.J.Biochem., 241, 1996
|
|
1CVL
 
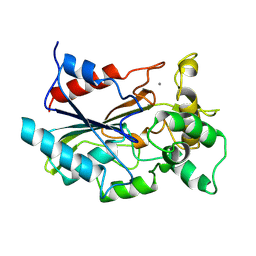 | | CRYSTAL STRUCTURE OF BACTERIAL LIPASE FROM CHROMOBACTERIUM VISCOSUM ATCC 6918 | | Descriptor: | CALCIUM ION, TRIACYLGLYCEROL HYDROLASE | | Authors: | Lang, D.A, Hofmann, B, Haalck, L, Hecht, H.-J, Spener, F, Schmid, R.D, Schomburg, D. | | Deposit date: | 1997-01-09 | | Release date: | 1997-04-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of a bacterial lipase from Chromobacterium viscosum ATCC 6918 refined at 1.6 angstroms resolution.
J.Mol.Biol., 259, 1996
|
|
1TAL
 
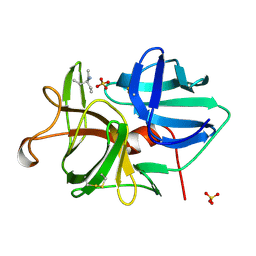 | | ALPHA-LYTIC PROTEASE AT 120 K (SINGLE STRUCTURE MODEL) | | Descriptor: | ALPHA-LYTIC PROTEASE, SULFATE ION, TRIS(HYDROXYETHYL)AMINOMETHANE | | Authors: | Rader, S.D, Agard, D.A. | | Deposit date: | 1996-10-30 | | Release date: | 1997-04-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Conformational substates in enzyme mechanism: the 120 K structure of alpha-lytic protease at 1.5 A resolution.
Protein Sci., 6, 1997
|
|
1TBG
 
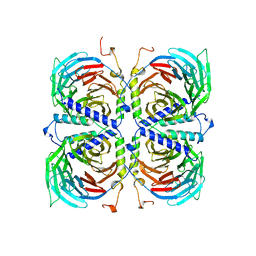 | | BETA-GAMMA DIMER OF THE HETEROTRIMERIC G-PROTEIN TRANSDUCIN | | Descriptor: | TRANSDUCIN | | Authors: | Sondek, J.S, Bohm, A, Lambright, D.G, Hamm, H.E, Sigler, P.B. | | Deposit date: | 1996-06-15 | | Release date: | 1997-04-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of a G-protein beta gamma dimer at 2.1A resolution.
Nature, 379, 1996
|
|
1CDJ
 
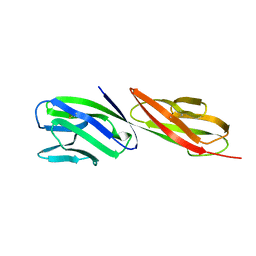 | | STRUCTURE OF T-CELL SURFACE GLYCOPROTEIN CD4 | | Descriptor: | T-CELL SURFACE GLYCOPROTEIN CD4 | | Authors: | Wu, H, Myszka, D, Tendian, S.W, Brouillette, C.G, Sweet, R.W, Chaiken, I.M, Hendrickson, W.A. | | Deposit date: | 1996-11-11 | | Release date: | 1997-04-01 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Kinetic and structural analysis of mutant CD4 receptors that are defective in HIV gp120 binding.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
1CDU
 
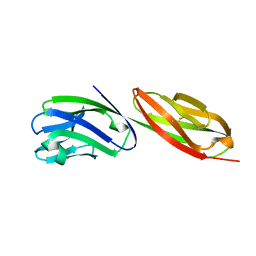 | | STRUCTURE OF T-CELL SURFACE GLYCOPROTEIN CD4 MUTANT WITH PHE 43 REPLACED BY VAL | | Descriptor: | T-CELL SURFACE GLYCOPROTEIN CD4 | | Authors: | Wu, H, Myszka, D, Tendian, S.W, Brouillette, C.G, Sweet, R.W, Chaiken, I.M, Hendrickson, W.A. | | Deposit date: | 1996-11-11 | | Release date: | 1997-04-01 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Kinetic and structural analysis of mutant CD4 receptors that are defective in HIV gp120 binding.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
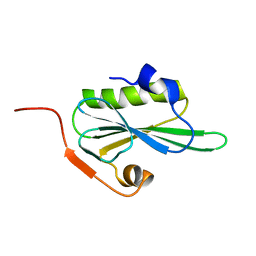
2VIK
 
 | |
