2LEK
 
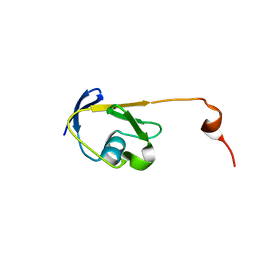 | | Solution NMR structure of a Thiamine Biosynthesis (ThiS) Protein RPA3574 from Rhodopseudomonas palustris refined with NH RDCs. Northeast Structural Genomics Consortium target RpR325 | | Descriptor: | Putative thiamin biosynthesis ThiS | | Authors: | Ramelot, T.A, Cort, J.R, Lee, H, Wang, H, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Acton, T.B, Xiao, R, Swapna, G, Everett, J.K, Prestegard, J.H, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2011-06-16 | | Release date: | 2011-06-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a Thiamine Biosynthesis (ThiS) Protein RPA3574 from Rhodopseudomonas palustris. Northeast Structural Genomics Consortium target RpR325
To be Published
|
|
2L01
 
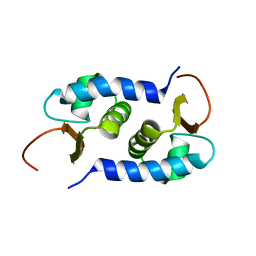 | | Solution NMR Structure of protein BVU3908 from Bacteroides vulgatus, Northeast Structural Genomics Consortium Target BvR153 | | Descriptor: | Uncharacterized protein | | Authors: | Eletsky, A, Lee, C, Wang, K, Ciccosanti, T.B, Hamilton, R, Acton, J.B, Xiao, G.B, Everett, J.K, Prestegard, J.H, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-06-29 | | Release date: | 2010-08-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of protein BVU3908 from Bacteroides vulgatus
To be Published
|
|
2L1N
 
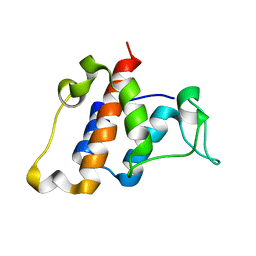 | | Solution NMR structure of the protein YP_399305.1 | | Descriptor: | Uncharacterized protein | | Authors: | Mohanty, B, Serrano, P, Geralt, M, Horst, R, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2010-07-30 | | Release date: | 2010-08-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the protein YP_399305.1
To be Published
|
|
2LV7
 
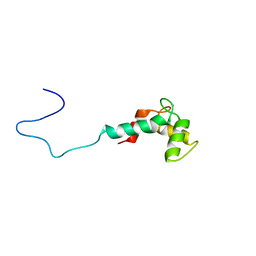 | | Solution structure of Ca2+-bound CaBP7 N-terminal doman | | Descriptor: | CALCIUM ION, Calcium-binding protein 7 | | Authors: | Mccue, H.V, Patel, P, Herbert, A.P, Lian, L, Burgoyne, R.D, Haynes, L.P. | | Deposit date: | 2012-06-29 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the Ca2+-bound N-terminal Domain of CaBP7: A REGULATOR OF GOLGI TRAFFICKING.
J.Biol.Chem., 287, 2012
|
|
2MK9
 
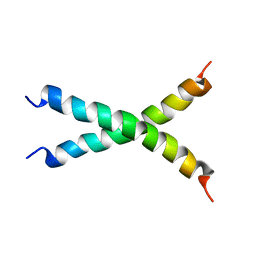 | |
2MKA
 
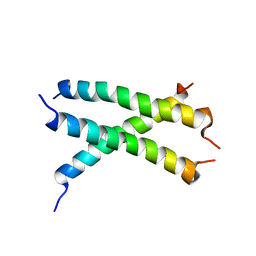 | |
2JW8
 | | Solution structure of stereo-array isotope labelled (SAIL) C-terminal dimerization domain of SARS coronavirus nucleocapsid protein | | Descriptor: | Nucleocapsid protein | | Authors: | Takeda, M, Chang, C, Ikeya, T, Guntert, P, Chang, Y, Hsu, Y, Huang, T, Kainosho, M. | | Deposit date: | 2007-10-06 | | Release date: | 2008-08-26 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the c-terminal dimerization domain of SARS coronavirus nucleocapsid protein solved by the SAIL-NMR method
J.Mol.Biol., 380, 2008
|
|
2MM2
 | |
2MM0
 | | Solution Structures of active Ptr ToxB and its inactive homolog highlight protein dynamics as a modulator of toxin activity | | Descriptor: | Host-selective toxin protein | | Authors: | Nyarko, A, Singarapu, K, Figueroa, M, Manning, V, Pandelova, I, Wolpert, T, Ciuffetti, L, Barbar, E. | | Deposit date: | 2014-03-06 | | Release date: | 2014-08-06 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structures of Pyrenophora tritici-repentis ToxB and Its Inactive Homolog Reveal Potential Determinants of Toxin Activity.
J.Biol.Chem., 289, 2014
|
|
2MXW
 
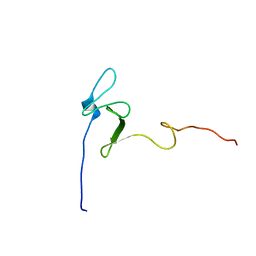 | |
2N03
 
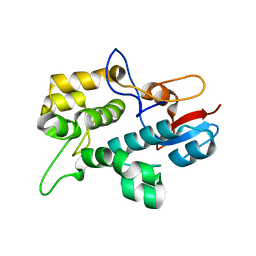 | | Solution NMR Structure plectin repeat domain 6 (4403-4606) of Plectin from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR6354E | | Descriptor: | Plectin | | Authors: | Pulavarti, S, Eletsky, A, Huang, Y, Janjua, H, Xiao, R, Acton, T, Everett, J, Montelione, G, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2015-03-04 | | Release date: | 2015-05-13 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure plectin repeat domain 6 (4403-4606) of Plectin from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR6354E
To be Published
|
|
2MS3
 | |
2MUU
 
 | |
2O6V
 
 | |
2NBD
 
 | |
2NBE
 
 | |
2JVZ
 
 | | Solution NMR Structure of the Second and Third KH Domains of KSRP | | Descriptor: | Far upstream element-binding protein 2 | | Authors: | Diaz-Moreno, I, Hollingworth, D, Garcia-Mayoral, M.F, Kelly, G, Cukier, C.D, Ramos, A. | | Deposit date: | 2007-09-28 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the Second and Third KH Domains of KSRP
To be Published, 2007
|
|
2KMM
 
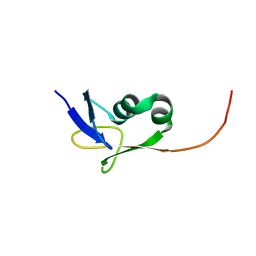 | | Solution NMR structure of the TGS domain of PG1808 from Porphyromonas gingivalis. Northeast Structural Genomics Consortium Target PgR122A (418-481) | | Descriptor: | Guanosine-3',5'-bis(Diphosphate) 3'-pyrophosphohydrolase | | Authors: | Yang, Y, Ramelot, T.A, Cort, J.R, Lee, D, Ciccosanti, C, Hamilton, K, Nair, R, Rost, B, Acton, T.B, Xiao, R, Swapna, G, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-07-31 | | Release date: | 2009-11-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the TGS domain of PG1808 from
Porphyromonas gingivalis. Northeast Structural Genomics Consortium Target
PgR122A (418-481)
To be Published
|
|
2KH2
 
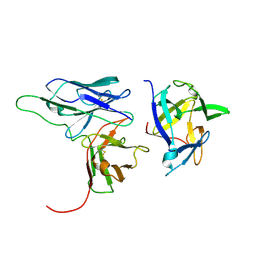 | | Solution structure of a scFv-IL-1B complex | | Descriptor: | Interleukin-1 beta, scFv | | Authors: | Wilkinson, I.C, Hall, C.J, Veverka, V, Muskett, F.W, Stephens, P.E, Taylor, R.J, Henry, A.J, Carr, M.D. | | Deposit date: | 2009-03-23 | | Release date: | 2009-09-08 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | High resolution NMR-based model for the structure of a scFv-IL-1beta complex: potential for NMR as a key tool in therapeutic antibody design and development.
J.Biol.Chem., 284, 2009
|
|
2KKO
 
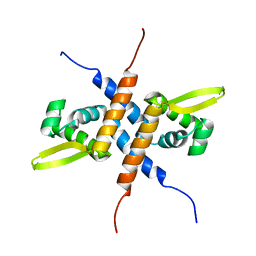 | | Solution NMR structure of the homodimeric winged helix-turn-helix DNA-binding domain (fragment 1-100) Mb0332 from Mycobacterium bovis, a possible ArsR-family transcriptional regulator. Northeast Structural Genomics Consortium Target MbR242E. | | Descriptor: | POSSIBLE TRANSCRIPTIONAL REGULATORY PROTEIN (POSSIBLY ARSR-FAMILY) | | Authors: | Ramelot, T.A, Cort, J.R, Wang, D, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Swapna, G, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-26 | | Release date: | 2009-08-11 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the homodimeric winged helix-turn-helix DNA-binding domain (fragment 1-100) Mb0332 from Mycobacterium bovis, a possible ArsR-family transcriptional regulator. Northeast Structural Genomics Consortium Target MbR242E.
To be Published
|
|
2KCD
 
 | | Solution NMR structure of SSP0047 from Staphylococcus saprophyticus. Northeast Structural Genomics Consortium Target SyR6. | | Descriptor: | Uncharacterized protein SSP0047 | | Authors: | Ramelot, T.A, Ding, K, Chen, C.X, Jiang, M, Ciccosanti, C, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T.B, Rost, B, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-19 | | Release date: | 2009-02-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of SSP0047 from Staphylococcus saprophyticus. Northeast Structural Genomics Consortium Target SyR6.
To be Published
|
|
2KJR
 
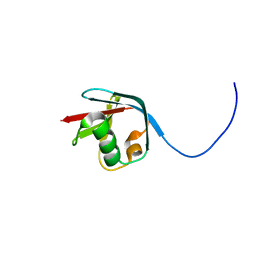 | | Solution NMR structure of the N-terminal Ubiquitin-like Domain from Tubulin-binding Cofactor B, CG11242, from Drosophila melanogaster. Northeast Structural Genomics Consortium Target FR629A (residues 8-92) | | Descriptor: | CG11242 | | Authors: | Ramelot, T.A, Cort, J.R, Shastry, R, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Swapna, G, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-08 | | Release date: | 2009-06-23 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the N-terminal Ubiquitin-like Domain from
Tubulin-binding Cofactor B, CG11242, from Drosophila melanogaster. Northeast
Structural Genomics Consortium Target FR629A (residues 8-92)
To be Published
|
|
2KPI
 
 | | Solution NMR structure of Streptomyces coelicolor SCO3027 modeled with Zn+2 bound, Northeast Structural Genomics Consortium Target RR58 | | Descriptor: | Uncharacterized protein SCO3027, ZINC ION | | Authors: | Ramelot, T.A, Cort, J.R, Garcia, M, Yee, A, Arrowmith, C.H, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-10-15 | | Release date: | 2010-02-02 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of Streptomyces coelicolor SCO3027 modeled with Zn+2 bound, Northeast Structural Genomics Consortium Target RR58
To be Published
|
|
2KKL
 
 | | Solution NMR structure of FHA domain of Mb1858 from Mycobacterium bovis. Northeast Structural Genomics Consortium Target MbR243C (24-155). | | Descriptor: | Uncharacterized protein Mb1858 | | Authors: | Yang, Y, Ramelot, T.A, Wang, D, Foote, E.L, Jiang, M, Nair, R, Rost, B, Swapna, G, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-25 | | Release date: | 2009-07-07 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of FHA domain of Mb1858 from Mycobacterium bovis. Northeast Structural Genomics Consortium Target MbR243C (24-155).
To be Published
|
|
2KKP
 
 | | Solution NMR structure of the phage integrase SAM-like Domain from Moth 1796 from Moorella thermoacetica. Northeast Structural Genomics Consortium Target MtR39K (residues 64-171). | | Descriptor: | Phage integrase | | Authors: | Ramelot, T.A, Wang, H, Ciccosanti, C, Jang, M, Nair, R, Rost, B, Swapna, G, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-26 | | Release date: | 2009-09-08 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the phage integrase SAM-like Domain from Moth 1796
from Moorella thermoacetica. Northeast
Structural Genomics Consortium Target MtR39K (residues 64-171).
To be Published
|
|
