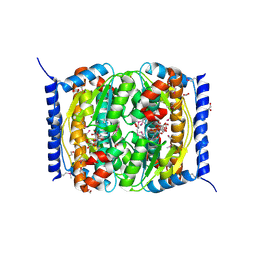
3ETN
 
 | |
3EU8
 
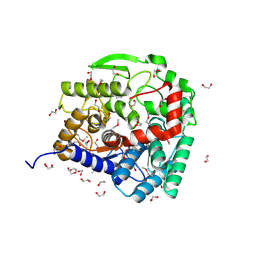 | |
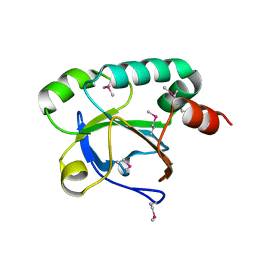
3EWL
 
 | |
3GKX
 
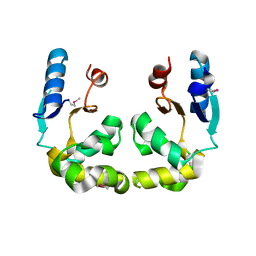 | |
3GZS
 
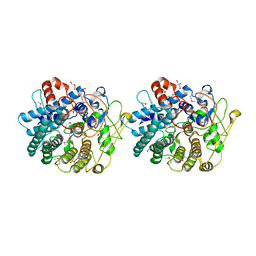 | |
3IG2
 
 | | The Crystal Structure of a Putative Phenylalanyl-tRNA synthetase (PheRS) beta chain domain from Bacteroides fragilis to 2.1A | | Descriptor: | MAGNESIUM ION, Phenylalanyl-tRNA synthetase beta chain | | Authors: | Stein, A.J, Sather, A, Hendricks, R, Keigher, L, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-07-27 | | Release date: | 2009-09-01 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | The Crystal Structure of a Putative Phenylalanyl-tRNA synthetase (PheRS) beta chain domain from Bacteroides fragilis to 2.1A
To be Published
|
|
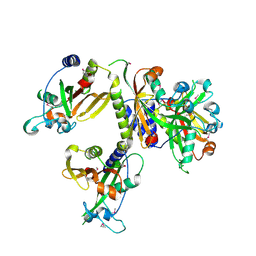
3EVY
 
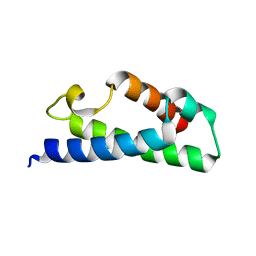 | | Crystal structure of a fragment of a putative type I restriction enzyme R protein from Bacteroides fragilis | | Descriptor: | Putative type I restriction enzyme R protein | | Authors: | Bonanno, J.B, Gilmore, M, Bain, K.T, Miller, S, Sampathkumar, P, Wasserman, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-10-13 | | Release date: | 2008-10-21 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of a fragment of a putative type I restriction enzyme R protein from Bacteroides fragilis
To be Published
|
|
3PET
 
 | |
3FHL
 
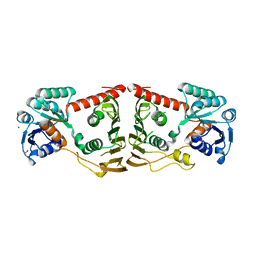 | | Crystal structure of a putative oxidoreductase from bacteroides fragilis nctc 9343 | | Descriptor: | GLYCEROL, MAGNESIUM ION, Putative oxidoreductase | | Authors: | Patskovsky, Y, Ramagopal, U, Toro, R, Gilmore, M, Miller, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-12-09 | | Release date: | 2008-12-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Crystal Structure of a Putative Oxidoreductase from Bacteroides Fragilis Nctc 9343
To be Published
|
|
3NQP
 
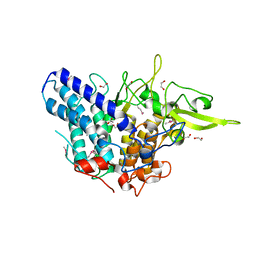 | |
3RWX
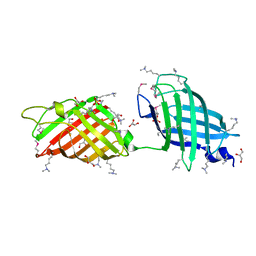 
 | |
3K50
 
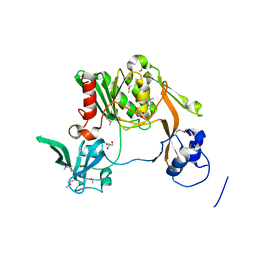 | |
3SLR
 
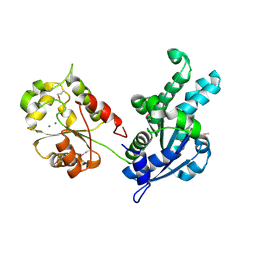 | | Crystal structure of N-terminal part of the protein BF1531 from Bacteroides fragilis containing phosphatase domain complexed with Mg. | | Descriptor: | MAGNESIUM ION, uncharacterized protein BF1531 | | Authors: | Fedorov, A.A, Fedorov, E.V, Toro, R, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2011-06-24 | | Release date: | 2011-07-20 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.712 Å) | | Cite: | Crystal structure of N-terminal part of the protein BF1531 from Bacteroides fragilis
containing phosphatase domain complexed with Mg.
TO BE PUBLISHED
|
|
7POO
 
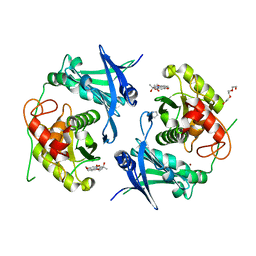 | | Crystal structure of profragilysin-3 (proBFT-3) from Bacteroides fragilis in complex with foliosidine in P212121. | | Descriptor: | ACETATE ION, BFT-3, PROLINE, ... | | Authors: | Eckhard, U, Guevara, T, Gomis-Ruth, F.X. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Repositioning small molecule drugs as allosteric inhibitors of the BFT-3 toxin from enterotoxigenic Bacteroides fragilis.
Protein Sci., 31, 2022
|
|
7POQ
 
 | | Crystal structure of profragilysin-3 (proBFT-3) from Bacteroides fragilis in complex with foliosidine in P41212. | | Descriptor: | 1,2-ETHANEDIOL, BFT-3, CHLORIDE ION, ... | | Authors: | Eckhard, U, Guevara, T, Gomis-Ruth, F.X. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Repositioning small molecule drugs as allosteric inhibitors of the BFT-3 toxin from enterotoxigenic Bacteroides fragilis.
Protein Sci., 31, 2022
|
|
7PND
 
 | | Crystal structure of profragilysin-3 (proBFT-3) from Bacteroides fragilis at 1.85 A resolution. | | Descriptor: | BFT-3, DIMETHYL SULFOXIDE, FORMIC ACID, ... | | Authors: | Eckhard, U, Guevara, T, Gomis-Ruth, F.X. | | Deposit date: | 2021-09-06 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Repositioning small molecule drugs as allosteric inhibitors of the BFT-3 toxin from enterotoxigenic Bacteroides fragilis.
Protein Sci., 31, 2022
|
|
7POL
 
 | | Crystal structure of profragilysin-3 (proBFT-3) from Bacteroides fragilis in complex with flumequine | | Descriptor: | (12~{R})-7-fluoranyl-12-methyl-4-oxidanylidene-1-azatricyclo[7.3.1.0^{5,13}]trideca-2,5(13),6,8-tetraene-3-carboxylic acid, BFT-3, CHLORIDE ION, ... | | Authors: | Eckhard, U, Guevara, T, Gomis-Ruth, F.X. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Repositioning small molecule drugs as allosteric inhibitors of the BFT-3 toxin from enterotoxigenic Bacteroides fragilis.
Protein Sci., 31, 2022
|
|
7POU
 
 | | Crystal structure of profragilysin-3 (proBFT-3) from Bacteroides fragilis in complex with hesperetin. | | Descriptor: | (2S)-5,7-dihydroxy-2-(3-hydroxy-4-methoxyphenyl)-2,3-dihydro-4H-1-benzopyran-4-one, BFT-3, DIMETHYL SULFOXIDE, ... | | Authors: | Eckhard, U, Guevara, T, Gomis-Ruth, F.X. | | Deposit date: | 2021-09-09 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Repositioning small molecule drugs as allosteric inhibitors of the BFT-3 toxin from enterotoxigenic Bacteroides fragilis.
Protein Sci., 31, 2022
|
|
7QNB
 
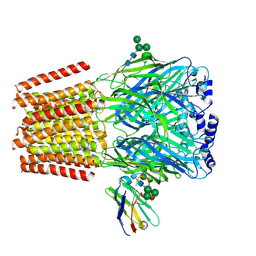 | | Cryo-EM structure of human full-length beta3gamma2 GABA(A)R in complex with GABA and nanobody Nb25 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GAMMA-AMINO-BUTANOIC ACID, ... | | Authors: | Sente, A, Desai, R, Naydenova, K, Malinauskas, T, Jounaidi, Y, Miehling, J, Zhou, X, Masiulis, S, Hardwick, S.W, Chirgadze, D.Y, Miller, K.W, Aricescu, A.R. | | Deposit date: | 2021-12-20 | | Release date: | 2022-04-13 | | Last modified: | 2023-11-15 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Differential assembly diversifies GABA A receptor structures and signalling.
Nature, 604, 2022
|
|
7QVP
 
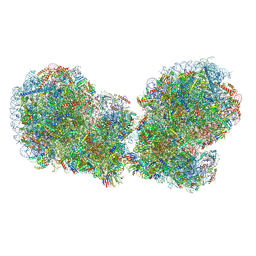 | | Human collided disome (di-ribosome) stalled on XBP1 mRNA | | Descriptor: | 18S ribosomal RNA, 28S ribosomal RNA, 40S ribosomal protein S10, ... | | Authors: | Denk, T.G, Tesina, P, Beckmann, R. | | Deposit date: | 2022-01-22 | | Release date: | 2022-10-12 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation.
Nat Commun, 13, 2022
|
|
7QGG
 
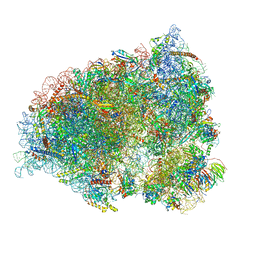 | | Neuronal RNA granules are ribosome complexes stalled at the pre-translocation state | | Descriptor: | 40S ribosomal protein S10, 40S ribosomal protein S11, 40S ribosomal protein S12, ... | | Authors: | Pulk, A, Kipper, K, Mansour, A. | | Deposit date: | 2021-12-08 | | Release date: | 2022-10-26 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.86 Å) | | Cite: | Neuronal RNA granules are ribosome complexes stalled at the pre-translocation state.
J.Mol.Biol., 434, 2022
|
|
7R7B
 
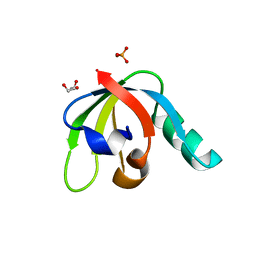 | |
7REW
 
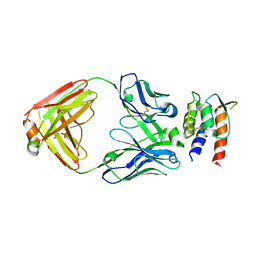 | | Crystal Structure of IL-13 in complex with MMAb3 Fab | | Descriptor: | IL13, anti-cyno interleukin 13 Fab heavy chain, anti-cyno interleukin 13 Fab light chain | | Authors: | Sudom, A, Min, X. | | Deposit date: | 2021-07-13 | | Release date: | 2022-05-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Development of a potent high-affinity human therapeutic antibody via novel application of recombination signal sequence-based affinity maturation.
J.Biol.Chem., 298, 2022
|
|
3GJO
 
 | |
4YEX
 
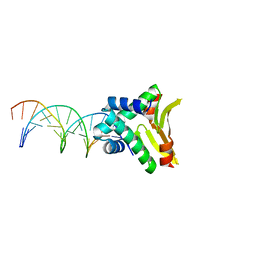 | | HUaa-19bp | | Descriptor: | DNA-binding protein HU-alpha, synthetic DNA strand | | Authors: | Hammel, M, Reyes, F.E, Parpana, R, Tainer, J.A, Adhya, S, Amlanjyoti, D. | | Deposit date: | 2015-02-24 | | Release date: | 2016-06-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | HU multimerization shift controls nucleoid compaction.
Sci Adv, 2, 2016
|
|
