3CEY
 
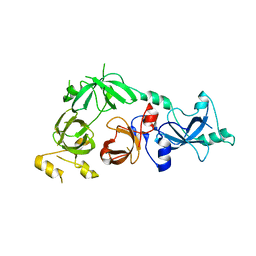 | | Crystal structure of L3MBTL2 | | Descriptor: | Lethal(3)malignant brain tumor-like 2 protein | | Authors: | Nady, N, Guo, Y, Pan, P, Allali-Hassani, A, Qi, C, Zhu, H, Dong, A, Mackenzie, F, Crombet, L, Loppnau, P, Kozieradzki, I, Vedadi, M, Edwards, A.M, Weigelt, J, Bountra, C, Arrowsmith, C.H, Bochkarev, A, Read, R, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-02-29 | | Release date: | 2008-05-06 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Methylation-state-specific recognition of histones by the MBT repeat protein L3MBTL2.
Nucleic Acids Res., 37, 2009
|
|
4GKC
 | |
4FTI
 
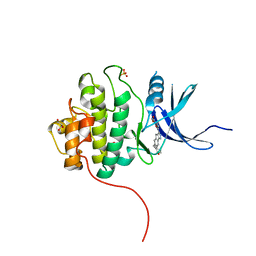 | | Crystal Structure of the CHK1 | | Descriptor: | GLYCEROL, ISOPROPYL ALCOHOL, SULFATE ION, ... | | Authors: | Kang, Y.N, Stuckey, J.A, Chang, P, Russell, A.J. | | Deposit date: | 2012-06-27 | | Release date: | 2012-08-22 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the CHK1
To be Published
|
|
4FTR
 
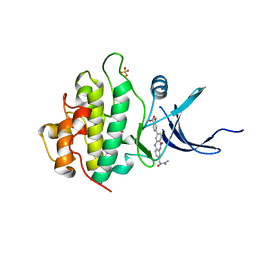 | | Crystal Structure of the CHK1 | | Descriptor: | 2-[3-(3-methoxy-4-nitrophenyl)-11-oxo-10,11-dihydro-5H-dibenzo[b,e][1,4]diazepin-8-yl]-N,N-dimethylacetamide, GLYCEROL, ISOPROPYL ALCOHOL, ... | | Authors: | Kang, Y.N, Stuckey, J.A, Chang, P, Russell, A.J. | | Deposit date: | 2012-06-27 | | Release date: | 2012-08-22 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal Structure of the CHK1
To be Published
|
|
5L90
 
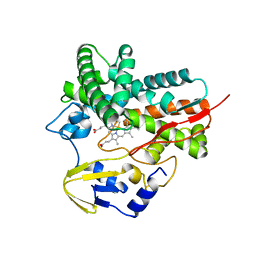 | |
3CTR
 
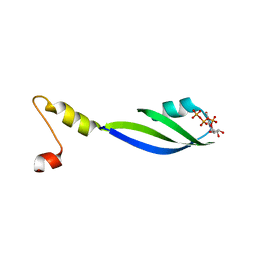 | | Crystal structure of the RRM-domain of the poly(A)-specific ribonuclease PARN bound to m7GTP | | Descriptor: | 7-METHYL-GUANOSINE-5'-TRIPHOSPHATE, Poly(A)-specific ribonuclease PARN | | Authors: | Monecke, T, Schell, S, Dickmanns, A, Ficner, R. | | Deposit date: | 2008-04-14 | | Release date: | 2008-07-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of the RRM domain of poly(A)-specific ribonuclease reveals a novel m(7)G-cap-binding mode.
J.Mol.Biol., 382, 2008
|
|
2RCQ
 | | Crystal structure of human apo Cellular Retinol Binding Protein II (CRBP-II) | | Descriptor: | L(+)-TARTARIC ACID, Retinol-binding protein II, cellular, ... | | Authors: | Monaco, H.L, Capaldi, S, Perduca, M. | | Deposit date: | 2007-09-20 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal structure of human cellular retinol-binding protein II to 1.2 A resolution.
Proteins, 70, 2007
|
|
4G1Y
 
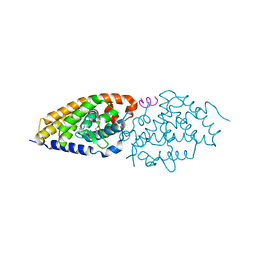 | | Structural basis for the accommodation of bis- and tris-aromatic derivatives in Vitamin D Nuclear Receptor | | Descriptor: | (4E,6Z)-7-(3-{[3,4-bis(hydroxymethyl)benzyl]oxy}phenyl)-3-ethylnona-4,6-dien-3-ol, Nuclear receptor coactivator 1, Vitamin D3 receptor A | | Authors: | Ciesielski, F, Sato, Y, Moras, D, Rochel, N. | | Deposit date: | 2012-07-11 | | Release date: | 2012-09-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural basis for the accommodation of bis- and tris-aromatic derivatives in vitamin d nuclear receptor.
J.Med.Chem., 55, 2012
|
|
3D3W
 
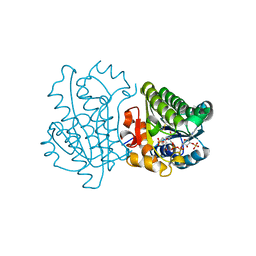 | |
3D4U
 
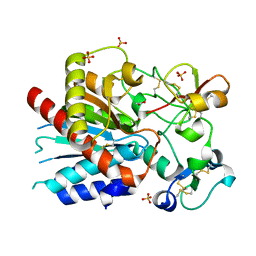 | | Bovine thrombin-activatable fibrinolysis inhibitor (TAFIa) in complex with tick-derived carboxypeptidase inhibitor. | | Descriptor: | ACETATE ION, Carboxypeptidase B2, Carboxypeptidase inhibitor, ... | | Authors: | Sanglas, L, Valnickova, Z, Arolas, J.L, Pallares, I, Guevara, T, Sola, M, Kristensen, T, Enghild, J.J, Aviles, F.X, Gomis-Ruth, F.X. | | Deposit date: | 2008-05-15 | | Release date: | 2008-08-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of activated thrombin-activatable fibrinolysis inhibitor, a molecular link between coagulation and fibrinolysis.
Mol.Cell, 31, 2008
|
|
3CWD
 
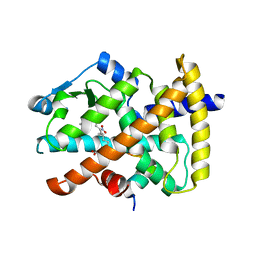 | | Molecular recognition of nitro-fatty acids by PPAR gamma | | Descriptor: | (9E,12Z)-10-nitrooctadeca-9,12-dienoic acid, (9Z,12E)-12-nitrooctadeca-9,12-dienoic acid, Peroxisome proliferator-activated receptor gamma, ... | | Authors: | Martynowski, D, Li, Y. | | Deposit date: | 2008-04-21 | | Release date: | 2008-07-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Molecular recognition of nitrated fatty acids by PPAR gamma.
Nat.Struct.Mol.Biol., 15, 2008
|
|
2RLQ
 | | NMR structure of CCP modules 2-3 of complement factor H | | Descriptor: | Complement factor H | | Authors: | Hocking, H.G, Herbert, A.P, Pangburn, M.K, Kavanagh, D, Barlow, P.N, Uhrin, D. | | Deposit date: | 2007-07-29 | | Release date: | 2008-02-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Structure of the N-terminal region of complement factor H and conformational implications of disease-linked sequence variations.
J.Biol.Chem., 283, 2008
|
|
3D6F
 
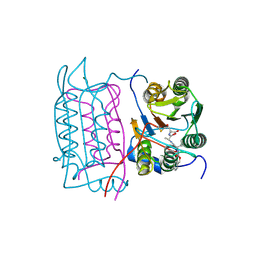 | |
4FXL
 | | Crystal structure of the D76N Beta-2 Microglobulin mutant | | Descriptor: | ACETATE ION, Beta-2-microglobulin, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Ricagno, S, Bellotti, V, Pepys, M.B, Stoppini, M, Bolognesi, M. | | Deposit date: | 2012-07-03 | | Release date: | 2012-08-15 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Hereditary systemic amyloidosis due to Asp76Asn variant beta-2-microglobulin.
N.Engl.J.Med., 366, 2012
|
|
4GD7
 
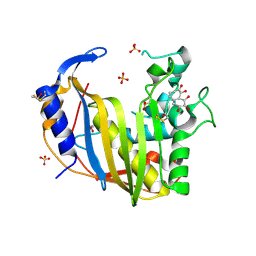 | | Wild-Type Human Thymidylate Synthase with bound Purpurogallin | | Descriptor: | 1,2-ETHANEDIOL, 2,3,4,6-tetrahydroxy-5H-benzo[7]annulen-5-one, SULFATE ION, ... | | Authors: | Celeste, L.R, Lebioda, L. | | Deposit date: | 2012-07-31 | | Release date: | 2013-07-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Oxidation of Cysteine 195 of Huyman Thymidylate Synthase by Purpurogallin
To be published, 2012
|
|
3D2W
 
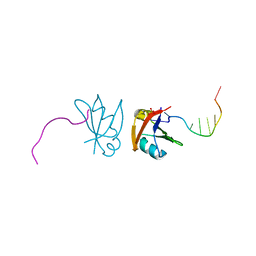 | | Crystal structure of mouse TDP-43 RRM2 domain in complex with DNA | | Descriptor: | DNA (5'-D(*DGP*DTP*DTP*DGP*DAP*DGP*DCP*DGP*DTP*DT)-3'), PHOSPHATE ION, TAR DNA-binding protein 43 | | Authors: | Kuo, P.H, Yuan, H.S. | | Deposit date: | 2008-05-09 | | Release date: | 2009-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural insights into TDP-43 in nucleic-acid binding and domain interactions
Nucleic Acids Res., 37, 2009
|
|
3D48
 
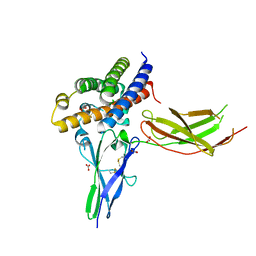 | |
3D57
 
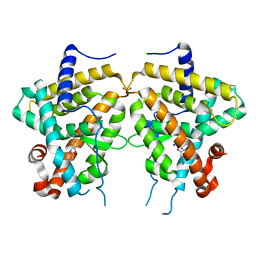 | | TR Variant D355R | | Descriptor: | SULFATE ION, Thyroid hormone receptor beta, [4-(4-HYDROXY-3-IODO-PHENOXY)-3,5-DIIODO-PHENYL]-ACETIC ACID | | Authors: | Jouravel, N. | | Deposit date: | 2008-05-15 | | Release date: | 2008-10-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular basis for dimer formation of TRbeta variant D355R.
Proteins, 75, 2008
|
|
3D5Q
 
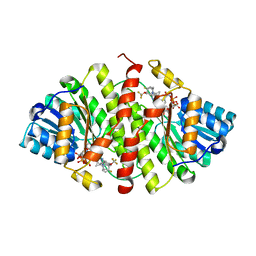 | | Crystal Structure of 11b-HSD1 in Complex with Triazole Inhibitor | | Descriptor: | 3-[1-(4-fluorophenyl)cyclopropyl]-4-(1-methylethyl)-5-[4-(trifluoromethoxy)phenyl]-4H-1,2,4-triazole, Corticosteroid 11-beta-dehydrogenase isozyme 1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Wang, Z, Liu, J, Sudom, A, Walker, N.P.C. | | Deposit date: | 2008-05-16 | | Release date: | 2008-10-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Distinctive molecular inhibition mechanisms for selective inhibitors of human 11beta-hydroxysteroid dehydrogenase type 1.
Bioorg.Med.Chem., 16, 2008
|
|
4G7W
 
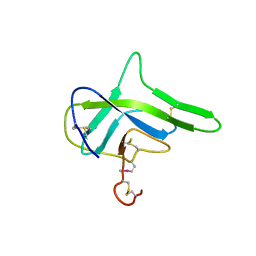 | |
4FSR
 
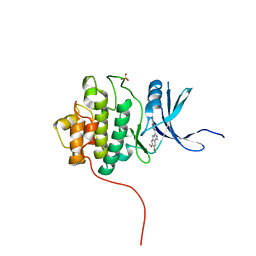 | | Crystal Structure of the CHK1 | | Descriptor: | 6,7-dimethoxy-3-[4-(1H-tetrazol-5-yl)phenyl]-1,4-dihydroindeno[1,2-c]pyrazole, SULFATE ION, Serine/threonine-protein kinase Chk1 | | Authors: | Kang, Y.N, Stuckey, J.A, Chang, P, Russell, A.J. | | Deposit date: | 2012-06-27 | | Release date: | 2012-08-22 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of the CHK1
To be Published
|
|
2RFE
 
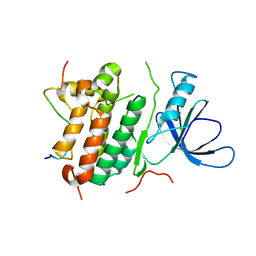 | | Crystal structure of the complex between the EGFR kinase domain and a Mig6 peptide | | Descriptor: | ERBB receptor feedback inhibitor 1, Epidermal growth factor receptor | | Authors: | Zhang, X, Pickin, K.A, Bose, R, Jura, N, Cole, P.A, Kuriyan, J. | | Deposit date: | 2007-09-28 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Inhibition of the EGF receptor by binding of MIG6 to an activating kinase domain interface.
Nature, 450, 2007
|
|
4G9C
 
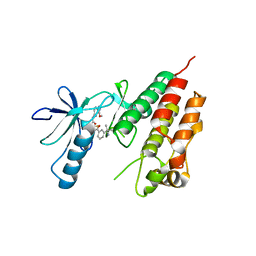 | | Human B-Raf Kinase Domain bound to a Type II Pyrazolopyridine Inhibitor | | Descriptor: | 3-{[3-(2-cyanopropan-2-yl)benzoyl]amino}-2,6-difluoro-N-(3-methoxy-2H-pyrazolo[3,4-b]pyridin-5-yl)benzamide, Serine/threonine-protein kinase B-raf | | Authors: | Voegtli, W.C, Sturgis, H.L. | | Deposit date: | 2012-07-23 | | Release date: | 2012-11-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Pyrazolopyridine inhibitors of B-Raf(V600E). Part 4: Rational design and kinase selectivity profile of cell potent type II inhibitors.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
2RHP
 
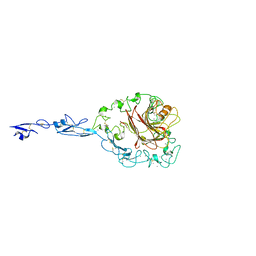 | | The Thrombospondin-1 Polymorphism Asn700Ser Associated with Cornoary Artery Disease Causes Local and Long-Ranging Changes in Protein Structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Carlson, C.B, Keck, J.L, Mosher, D.F. | | Deposit date: | 2007-10-09 | | Release date: | 2008-05-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Influences of the N700S Thrombospondin-1 Polymorphism on Protein Structure and Stability.
J.Biol.Chem., 283, 2008
|
|
3CQG
 
 | | Nucleoporin Nup107/Nup133 interaction complex, delta finger mutant | | Descriptor: | Nuclear pore complex protein Nup107, Nuclear pore complex protein Nup133 | | Authors: | Jeudy, S, Boehmer, T, Berke, I, Schwartz, T.U. | | Deposit date: | 2008-04-02 | | Release date: | 2008-07-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and functional studies of Nup107/Nup133 interaction and its implications for the architecture of the nuclear pore complex.
Mol.Cell, 30, 2008
|
|
