2DCQ
 
 | |
2DIB
 
 | | Solution structure of the 11th filamin domain from human Filamin-B | | Descriptor: | Filamin-B | | Authors: | Tomizawa, T, Koshiba, S, Watanabe, S, Harada, T, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-03-29 | | Release date: | 2006-09-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the 11th filamin domain from human Filamin-B
To be Published
|
|
2DMX
 
 | | Solution structure of the J domain of DnaJ homolog subfamily B member 8 | | Descriptor: | DnaJ homolog subfamily B member 8 | | Authors: | Ohnishi, S, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-04-24 | | Release date: | 2006-10-24 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the J domain of DnaJ homolog subfamily B member 8
To be Published
|
|
1J3N
 
 | | Crystal Structure of 3-oxoacyl-(acyl-carrier protein) Synthase II from Thermus thermophilus HB8 | | Descriptor: | 3-oxoacyl-(acyl-carrier protein) synthase II, CITRIC ACID, MAGNESIUM ION | | Authors: | Bagautdinov, B, Miyano, M, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-02-10 | | Release date: | 2003-03-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of 3-oxoacyl-(acyl-carrier protein) synthase II from Thermus thermophilus HB8.
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2M1T
 
 | |
2KE5
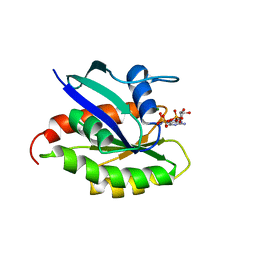 
 | | Solution structure and dynamics of the small GTPase Ralb in its active conformation: significance for effector protein binding | | Descriptor: | MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, Ras-related protein Ral-B | | Authors: | Fenwick, R, Prasannan, S, Campbell, L.J, Nietlispach, D, Evetts, K.A, Camonis, J, Mott, H.R, Owen, D. | | Deposit date: | 2009-01-23 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the small GTPase RalB in its active conformation: significance for effector protein binding
Biochemistry, 48, 2009
|
|
1SLY
 
 | | COMPLEX OF THE 70-KDA SOLUBLE LYTIC TRANSGLYCOSYLASE WITH BULGECIN A | | Descriptor: | 4-O-(4-O-SULFONYL-N-ACETYLGLUCOSAMININYL)-5-METHYLHYDROXY-L-PROLINE-TAURINE, 70-KDA SOLUBLE LYTIC TRANSGLYCOSYLASE | | Authors: | Thunnissen, A.M.W.H, Kalk, K.H, Rozeboom, H.J, Dijkstra, B.W. | | Deposit date: | 1995-08-02 | | Release date: | 1996-08-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of the 70-kDa soluble lytic transglycosylase complexed with bulgecin A. Implications for the enzymatic mechanism.
Biochemistry, 34, 1995
|
|
4OKO
 
 | | Crystal structure of Francisella tularensis REP34 (Rapid Encystment Phenotype Protein 34 KDa) | | Descriptor: | ACETATE ION, Rapid Encystment Phenotype Protein 34 KDa, ZINC ION | | Authors: | Feld, G.K, Segelke, B.W, Corzett, M.H, Hunter, M.S, Frank, M, Rasley, A. | | Deposit date: | 2014-01-22 | | Release date: | 2014-09-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.053 Å) | | Cite: | Structure and Function of REP34 Implicates Carboxypeptidase Activity in Francisella tularensis Host Cell Invasion.
J.Biol.Chem., 289, 2014
|
|
7FAZ
 
 | | Crystal structure of the SARS-CoV-2 main protease in complex with Y180 | | Descriptor: | (2~{R})-~{N}-dibenzofuran-3-yl-~{N}-[(1~{R})-2-[[(1~{S})-1-(4-fluorophenyl)ethyl]amino]-2-oxidanylidene-1-pyridin-3-yl-ethyl]-2-oxidanyl-propanamide, 3C-like proteinase, SODIUM ION | | Authors: | Zeng, R, Quan, B.X, Liu, X.L, Lei, J. | | Deposit date: | 2021-07-08 | | Release date: | 2021-07-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | An orally available M pro inhibitor is effective against wild-type SARS-CoV-2 and variants including Omicron.
Nat Microbiol, 7, 2022
|
|
5SR4
 
 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with Z2479779298 - (R,S) and (S,R) isomers | | Descriptor: | (1R,3S)-3-[(7-fluoro-9H-pyrimido[4,5-b]indol-4-yl)amino]cyclopentan-1-ol, (1S,3R)-3-[(7-fluoro-9H-pyrimido[4,5-b]indol-4-yl)amino]cyclopentan-1-ol, Non-structural protein 3 | | Authors: | Correy, G.J, Fraser, J.S. | | Deposit date: | 2022-06-09 | | Release date: | 2022-07-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Iterative computational design and crystallographic screening identifies potent inhibitors targeting the Nsp3 macrodomain of SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5SR7
 
 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with Z4914649782 - (R,R,S) and (S,S,R) isomers | | Descriptor: | (1R,6S,7R)-3-(7-fluoro-9H-pyrimido[4,5-b]indol-4-yl)-3-azabicyclo[4.1.0]heptane-7-carboxylic acid, (1S,6R,7S)-3-(7-fluoro-9H-pyrimido[4,5-b]indol-4-yl)-3-azabicyclo[4.1.0]heptane-7-carboxylic acid, Non-structural protein 3 | | Authors: | Correy, G.J, Fraser, J.S. | | Deposit date: | 2022-06-09 | | Release date: | 2022-07-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Iterative computational design and crystallographic screening identifies potent inhibitors targeting the Nsp3 macrodomain of SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5SRB
 
 | |
1SRN
 
 | |
9BFB
 
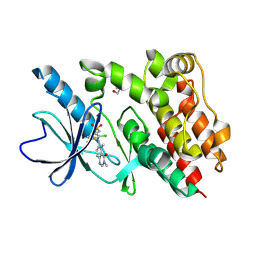 | | Crystal structure of BRAF kinase domain with PF-07284890 | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, N-{2-chloro-3-[(3,5-dimethyl-4-oxo-3,4-dihydroquinazolin-6-yl)amino]-4-fluorophenyl}-3-fluoropropane-1-sulfonamide, ... | | Authors: | Mou, T.-C. | | Deposit date: | 2024-04-17 | | Release date: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Identification of the Clinical Candidate PF-07284890 ( ARRY-461 ), a Highly Potent and Brain Penetrant BRAF Inhibitor for the Treatment of Cancer.
J.Med.Chem., 67, 2024
|
|
6D4O
 
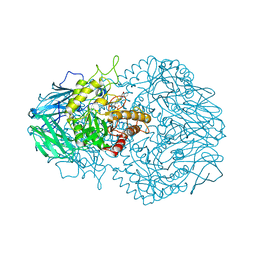 | | Eubacterium eligens beta-glucuronidase bound to an amoxapine-glucuronide conjugate | | Descriptor: | (5aR,9aR)-2-chloro-11-(4-beta-D-glucopyranuronosylpiperazin-1-yl)-5a,6,9,9a-tetrahydrodibenzo[b,f][1,4]oxazepine, Beta-glucuronidase, CHLORIDE ION, ... | | Authors: | Pellock, S.J, Walton, W.G, Redinbo, M.R. | | Deposit date: | 2018-04-18 | | Release date: | 2018-07-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Gut Microbial beta-Glucuronidase Inhibition via Catalytic Cycle Interception.
ACS Cent Sci, 4, 2018
|
|
3QRI
 
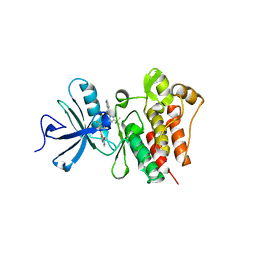 | | The crystal structure of human abl1 kinase domain in complex with DCC-2036 | | Descriptor: | 4-[4-({[3-tert-butyl-1-(quinolin-6-yl)-1H-pyrazol-5-yl]carbamoyl}amino)-3-fluorophenoxy]-N-methylpyridine-2-carboxamide, SODIUM ION, Tyrosine-protein kinase ABL1 | | Authors: | Chan, W.W, Wise, S.C, Kaufman, M.D, Ahn, Y.M, Ensinger, C.L, Haack, T, Hood, M.M, Jones, J, Lord, J.W, Lu, W.P, Miller, D, Patt, W.C, Smith, B.D, Petillo, P.A, Rutkoski, T.J, Telikepalli, H, Vogeti, L, Yao, T, Chun, L, Clark, R, Evangelista, P, Gavrilescu, L.C, Lazarides, K, Zaleskas, V.M, Stewart, L.J, Van Etten, R.A, Flynn, D.L. | | Deposit date: | 2011-02-18 | | Release date: | 2011-06-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conformational Control Inhibition of the BCR-ABL1 Tyrosine Kinase, Including the Gatekeeper T315I Mutant, by the Switch-Control Inhibitor DCC-2036.
Cancer Cell, 19, 2011
|
|
4OOS
 
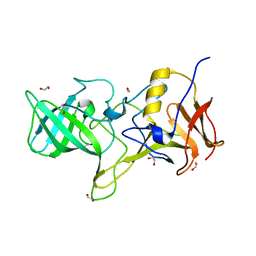 | |
2ZN7
 
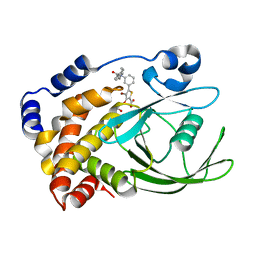 | | CRYSTAL STRUCTURES OF PTP1B-Inhibitor Complexes | | Descriptor: | 4-bromo-3-(carboxymethoxy)-5-{3-[cyclohexyl(phenylcarbonyl)amino]phenyl}thiophene-2-carboxylic acid, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Xu, W, Wu, J. | | Deposit date: | 2008-04-22 | | Release date: | 2008-10-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-based optimization of protein tyrosine phosphatase-1 B inhibitors: capturing interactions with arginine 24
Chemmedchem, 3, 2008
|
|
4XG7
 
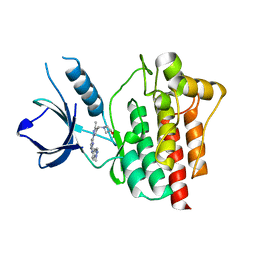 | | Crystal structure of an inhibitor-bound Syk | | Descriptor: | 1-[(3-methyl-1-{2-[(1-methyl-1H-indazol-5-yl)amino]pyrimidin-4-yl}-1H-pyrazol-4-yl)methyl]azetidin-3-ol, Tyrosine-protein kinase SYK | | Authors: | Lee, S.J, Choi, J, Han, B.G, Song, H, Koh, J.S, Lee, B.I. | | Deposit date: | 2014-12-30 | | Release date: | 2015-12-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structures of spleen tyrosine kinase in complex with novel inhibitors: structural insights for design of anticancer drugs
Febs J., 283, 2016
|
|
6TI2
 
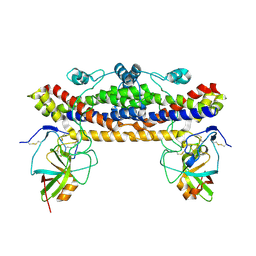 | |
3NCA
 
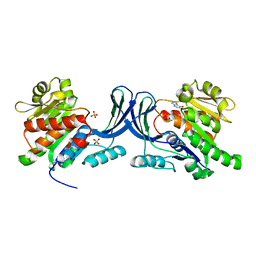 | | X-ray structure of ketohexokinase in complex with a thieno pyridinol compound | | Descriptor: | Ketohexokinase, SULFATE ION, thieno[3,2-b]pyridin-7-ol | | Authors: | Abad, M.C, Gibbs, A.C, Spurlino, J.C. | | Deposit date: | 2010-06-04 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Electron density guided fragment-based lead discovery of ketohexokinase inhibitors.
J.Med.Chem., 53, 2010
|
|
1TFD
 
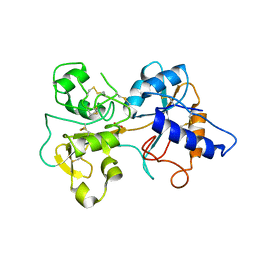 | |
4D6C
 
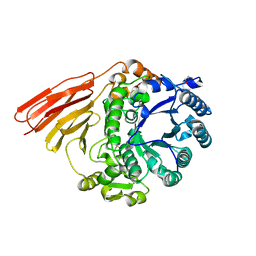 | | Crystal structure of a family 98 glycoside hydrolase catalytic module (Sp3GH98)(L19 mutant) | | Descriptor: | 1,2-ETHANEDIOL, GLYCOSIDE HYDROLASE | | Authors: | Kwan, D.H, Constantinescu, I, Chapanian, R, Higgins, M.A, Samain, E, Boraston, A.B, Kizhakkedathu, J.N, Withers, S.G. | | Deposit date: | 2014-11-11 | | Release date: | 2014-11-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Towards Efficient Enzymes for the Generation of Universal Blood Through Structure-Guided Directed Evolution.
J.Am.Chem.Soc., 137, 2015
|
|
2MYZ
 
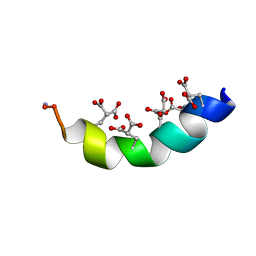 | | The Solution Structure of the Magnesium-bound Conantokin-R1B Mutant | | Descriptor: | Conantokin-R1-B | | Authors: | Kunda, S, Yuan, Y, Balsara, R.D, Zajicek, J, Castellino, F.J. | | Deposit date: | 2015-02-04 | | Release date: | 2015-06-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Hydroxyproline-induced Helical Disruption in Conantokin Rl-B Affects Subunit-selective Antagonistic Activities toward Ion Channels of N-Methyl-d-aspartate Receptors.
J.Biol.Chem., 290, 2015
|
|
2MZL
 
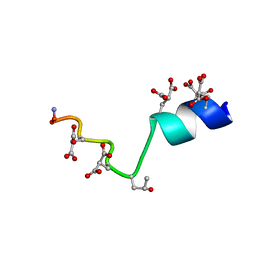 | | The Solution Structure of the Magnesium-bound Conantokin-G Mutant | | Descriptor: | Conantokin-R1-B | | Authors: | Kunda, S, Yuan, Y, Balsara, R.D, Zajicek, J, Castellino, F.J. | | Deposit date: | 2015-02-14 | | Release date: | 2015-06-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Hydroxyproline-induced Helical Disruption in Conantokin Rl-B Affects Subunit-selective Antagonistic Activities toward Ion Channels of N-Methyl-d-aspartate Receptors.
J.Biol.Chem., 290, 2015
|
|
