2N13
 
 | |
2MSG
 
 | | Solid-state NMR structure of ubiquitin | | Descriptor: | Ubiquitin | | Authors: | Lakomek, N, Habenstein, B, Loquet, A, Shi, C, Giller, K, Wolff, S, Becker, S, Fasshuber, H, Lange, A. | | Deposit date: | 2014-08-04 | | Release date: | 2015-02-18 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Structural heterogeneity in microcrystalline ubiquitin studied by solid-state NMR.
Protein Sci., 24, 2015
|
|
2MQJ
 
 | |
2N7D
 
 | |
7P08
 
 | |
7P0V
 
 | | Crystal structure of human SF3A1 ubiquitin-like domain in complex with U1 snRNA stem-loop 4 | | Descriptor: | Isoform 2 of Splicing factor 3A subunit 1, PENTAETHYLENE GLYCOL, RNA (5'-R(P*GP*GP*GP*GP*AP*CP*UP*GP*CP*GP*UP*UP*CP*GP*CP*GP*CP*UP*UP*UP*CP*CP*CP*C)-3'), ... | | Authors: | Sabath, K, de Vries, T, Jonas, S. | | Deposit date: | 2021-06-30 | | Release date: | 2022-04-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Sequence-specific RNA recognition by an RGG motif connects U1 and U2 snRNP for spliceosome assembly.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7OOJ
 
 | | Structure of D-Thr53 Ubiquitin | | Descriptor: | CADMIUM ION, Ubiquitin | | Authors: | Becker, S. | | Deposit date: | 2021-05-27 | | Release date: | 2022-05-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A litmus test for classifying recognition mechanisms of transiently binding proteins.
Nat Commun, 13, 2022
|
|
2OOB
 
 | |
6V5D
 
 | | EROS3 RDC and NOE Derived Ubiquitin Ensemble | | Descriptor: | Ubiquitin | | Authors: | Lange, O.F, Lakomek, N.A, Smith, C.A, Griesinger, C, de Groot, B.L. | | Deposit date: | 2019-12-04 | | Release date: | 2020-01-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Enhancing NMR derived ensembles with kinetics on multiple timescales.
J.Biomol.Nmr, 74, 2020
|
|
2OJR
 
 | |
2NR2
 
 | | The MUMO (minimal under-restraining minimal over-restraining) method for the determination of native states ensembles of proteins | | Descriptor: | Ubiquitin | | Authors: | Richter, B, Gsponer, J, Varnai, P, Salvatella, X, Vendruscolo, M. | | Deposit date: | 2006-11-01 | | Release date: | 2007-05-08 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The MUMO (minimal under-restraining minimal over-restraining) method for the determination of native state ensembles of proteins
J.Biomol.Nmr, 37, 2007
|
|
6W9R
 
 | |
2O6V
 
 | |
2PEA
 
 | |
2PE9
 
 | |
3VFK
 
 | | The structure of monodechloro-teicoplanin in complex with its ligand, using ubiquitin as a ligand carrier | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-amino-2-deoxy-beta-D-glucopyranose, 8-METHYLNONANOIC ACID, ... | | Authors: | Economou, N.J, Weeks, S.D, Grasty, K.C, Loll, P.J. | | Deposit date: | 2012-01-09 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of the complex between teicoplanin and a bacterial cell-wall peptide: use of a carrier-protein approach.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3VDZ
 
 | | Tailoring Encodable Lanthanide-Binding Tags as MRI Contrast Agents: xq-dSE3-Ubiquitin at 2.4 Angstroms | | Descriptor: | GADOLINIUM ATOM, SULFATE ION, Ubiquitin-40S ribosomal protein S27a | | Authors: | Daughtry, K.D, Martin, L.J, Surraju, A, Imperiali, B, Allen, K.N. | | Deposit date: | 2012-01-06 | | Release date: | 2012-11-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Tailoring encodable lanthanide-binding tags as MRI contrast agents.
Chembiochem, 13, 2012
|
|
3ZLZ
 
 | |
4A20
 
 | | Crystal structure of the Ubl domain of Mdy2 (Get5) at 1.78A | | Descriptor: | SULFATE ION, UBIQUITIN-LIKE PROTEIN MDY2 | | Authors: | Simon, A.C, Simpson, P.J, Murray, J.W, Isaacson, R.L. | | Deposit date: | 2011-09-21 | | Release date: | 2012-11-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structure of the Sgt2/Get5 Complex Provides Insights Into Get-Mediated Targeting of Tail-Anchored Membrane Proteins
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
8CMR
 
 | | Linear specific OTU-type DUB SnOTU from the pathogen S. negenvensis in complex with linear di-ubiquitin | | Descriptor: | OTU domain-containing protein, Polyubiquitin-B | | Authors: | Uthoff, M, Hermanns, T, Boll, V, Hofmann, K, Baumann, U. | | Deposit date: | 2023-02-21 | | Release date: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Functional and structural diversity in deubiquitinases of the Chlamydia-like bacterium Simkania negevensis.
Nat Commun, 14, 2023
|
|
8DMB
 
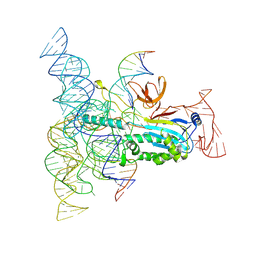 | | Structure of Desulfovirgula thermocuniculi IsrB (DtIsrB) in complex with omega RNA and target DNA | | Descriptor: | MAGNESIUM ION, Ubiquitin-like protein SMT3,IsrB protein,monomeric superfolder Green Fluorescent Protein, non-target DNA, ... | | Authors: | Seiichi, H, Kappel, K, Zhang, F. | | Deposit date: | 2022-07-08 | | Release date: | 2022-10-19 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structure of the OMEGA nickase IsrB in complex with omega RNA and target DNA.
Nature, 610, 2022
|
|
8E7O
 
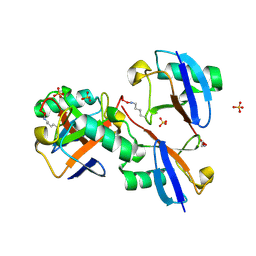 | | CRYSTAL STRUCTURE OF LYS48-LINKED TETRAUBIQUITIN | | Descriptor: | SULFATE ION, Ubiquitin | | Authors: | Lemma, B.E, Fushman, D. | | Deposit date: | 2022-08-24 | | Release date: | 2022-11-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mechanism of selective recognition of Lys48-linked polyubiquitin by macrocyclic peptide inhibitors of proteasomal degradation.
Nat Commun, 14, 2023
|
|
8DMS
 
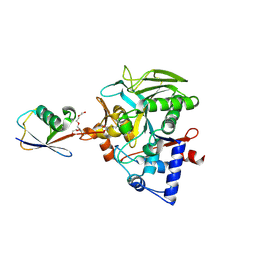 | |
8DMQ
 
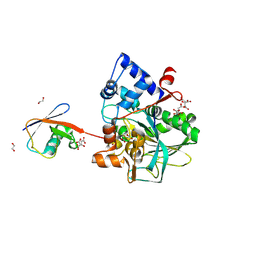 | |
6GF2
 
 | | The structure of the ubiquitin-like modifier FAT10 reveals a novel targeting mechanism for degradation by the 26S proteasome | | Descriptor: | Ubiquitin D | | Authors: | Aichem, A, Anders, S, Catone, N, Roessler, P, Stotz, S, Berg, A, Schwab, R, Scheuermann, S, Bialas, J, Schmidtke, G, Peter, C, Groettrup, M, Wiesner, S. | | Deposit date: | 2018-04-29 | | Release date: | 2018-08-08 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation.
Nat Commun, 9, 2018
|
|
