6DXA
 
 | |
6QJK
 
 | |
6QFF
 
 | | Crystal Structure of Human Kallikrein 6 in complex with GSK144 | | Descriptor: | 4-[(5-phenyl-1~{H}-imidazol-2-yl)methylamino]-2-(pyridin-3-ylmethoxy)benzenecarboximidamide, GLYCEROL, Kallikrein-6 | | Authors: | Thorpe, J.H. | | Deposit date: | 2019-01-10 | | Release date: | 2019-05-08 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Evaluation of a crystallographic surrogate for kallikrein 5 in the discovery of novel inhibitors for Netherton syndrome.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
6DPX
 
 | | X-ray crystal structure of AmpC beta-lactamase with inhibitor | | Descriptor: | (3-{[(3-chloro-2-hydroxyphenyl)sulfonyl]amino}phenyl)acetic acid, Beta-lactamase | | Authors: | Singh, I. | | Deposit date: | 2018-06-09 | | Release date: | 2018-07-04 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Ultra-large library docking for discovering new chemotypes.
Nature, 566, 2019
|
|
6QLM
 
 | | Cathepsin-K in complex with MIV-701 | | Descriptor: | (CARBAMOYLMETHYL-CARBOXYMETHYL-AMINO)-ACETIC ACID, Cathepsin K, GLYCEROL, ... | | Authors: | Derbyshire, D.J. | | Deposit date: | 2019-02-01 | | Release date: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Successful development of 3-oxohexahydrofuropyrrole amino acid amides as inhibitors of Cathepsin-K.
To Be Published
|
|
6QM0
 
 | | Cathepsin-K in complex with amino-oxaazabicyclo[3.3.0]octanyl containing inhibitor | | Descriptor: | (CARBAMOYLMETHYL-CARBOXYMETHYL-AMINO)-ACETIC ACID, Cathepsin K, ~{N}-[(2~{S})-1-[(3~{R},3~{a}~{R},6~{R},6~{a}~{R})-6-azanyl-3-oxidanyl-2,3,3~{a},5,6,6~{a}-hexahydrofuro[3,2-b]pyrrol-4-yl]-4-methyl-1-oxidanylidene-pentan-2-yl]-3-fluoranyl-4-[2-(4-methylpiperazin-1-yl)-1,3-thiazol-4-yl]benzamide | | Authors: | Derbyshire, D.J. | | Deposit date: | 2019-02-01 | | Release date: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Successful development of 3-oxohexahydrofuropyrrole amino acid amides as inhibitors of Cathepsin-K.
To Be Published
|
|
6D9B
 
 | |
6DXF
 
 | |
6QV1
 
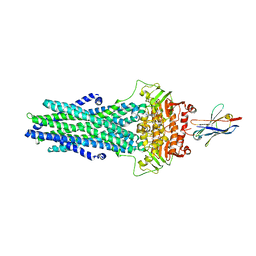 | | Structure of ATPgS-bound outward-facing TM287/288 in complex with nanobody Nb_TM1 | | Descriptor: | ABC transporter, ATP-binding protein, MAGNESIUM ION, ... | | Authors: | Hutter, C.A.J, Huerlimann, L.M, Zimmermann, I, Egloff, P, Seeger, M.A. | | Deposit date: | 2019-03-01 | | Release date: | 2019-05-29 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (3.48 Å) | | Cite: | The extracellular gate shapes the energy profile of an ABC exporter.
Nat Commun, 10, 2019
|
|
6QV2
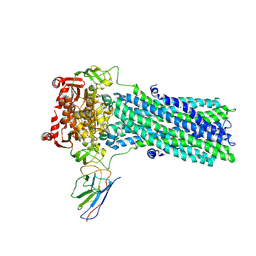 
 | | Structure of ATPgS-bound outward-facing TM287/288 in complex with nanobody Nb_TM#2 | | Descriptor: | ABC transporter, ATP-binding protein, MAGNESIUM ION, ... | | Authors: | Hutter, C.A.J, Huerlimann, L.M, Zimmermann, I, Egloff, P, Seeger, M.A. | | Deposit date: | 2019-03-01 | | Release date: | 2019-05-29 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (4.23 Å) | | Cite: | The extracellular gate shapes the energy profile of an ABC exporter.
Nat Commun, 10, 2019
|
|
6D3Y
 
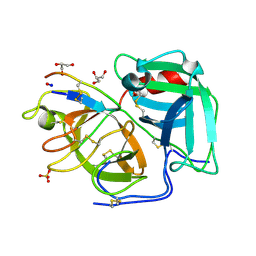 | |
6D46
 
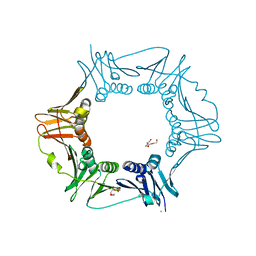 | |
6D5A
 
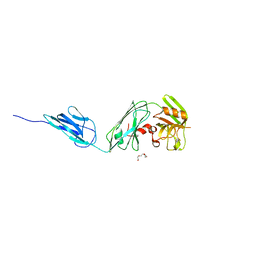 | |
6QLX
 
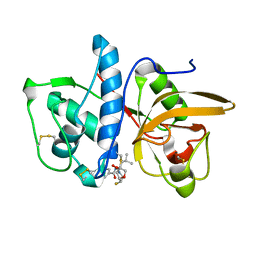 | | Cathepsin-K in complex with fluoro-oxa-azabicyclo[3.3.0]octanyl containing inhibitor | | Descriptor: | Cathepsin K, ~{N}-[(2~{S})-1-[(3~{R},3~{a}~{R},6~{S},6~{a}~{S})-6-fluoranyl-3-oxidanyl-2,3,3~{a},5,6,6~{a}-hexahydrofuro[3,2-b]pyrrol-4-yl]-4-methyl-1-oxidanylidene-pentan-2-yl]-4-thiophen-2-yl-benzamide | | Authors: | Derbyshire, D.J. | | Deposit date: | 2019-02-01 | | Release date: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Successful development of 3-oxohexahydrofuropyrrole amino acid amides as inhibitors of Cathepsin-K.
To Be Published
|
|
6DEG
 
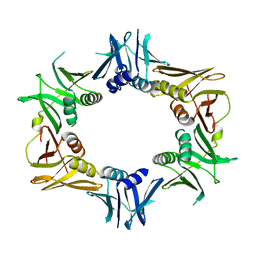 | |
6DEX
 
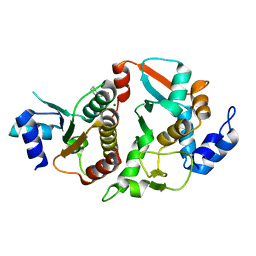 | | Structure of Eremothecium gossypii Shu1:Shu2 complex | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, MAGNESIUM ION, Suppressor of hydroxyurea sensitivity protein 1, ... | | Authors: | Singh, N, Corbett, K.D. | | Deposit date: | 2018-05-13 | | Release date: | 2018-05-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of Eremothecium gossypii Shu1:Shu2 complex
To Be Published
|
|
5DQF
 
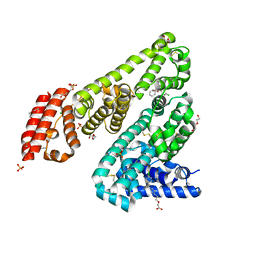 | | Horse Serum Albumin (ESA) in complex with Cetirizine | | Descriptor: | (2-{4-[(R)-(4-chlorophenyl)(phenyl)methyl]piperazin-1-yl}ethoxy)acetic acid, (2-{4-[(S)-(4-chlorophenyl)(phenyl)methyl]piperazin-1-yl}ethoxy)acetic acid, CHLORIDE ION, ... | | Authors: | Handing, K.B, Shabalin, I.G, Majorek, K.A, Chruszcz, M, Almo, S.C, Minor, W, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2015-09-14 | | Release date: | 2015-12-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structure of equine serum albumin in complex with cetirizine reveals a novel drug binding site.
Mol.Immunol., 71, 2016
|
|
6D9C
 
 | |
6DNT
 
 | | UDP-N-acetylglucosamine 4-epimerase from Methanobrevibacter ruminantium M1 in complex with UDP-N-acetylmuramic acid | | Descriptor: | (2R)-2-{[(2R,3R,4R,5S,6R)-3-(acetylamino)-2-{[(S)-{[(R)-{[(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}-5-hydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-4-yl]oxy}propanoic acid, 1,2-ETHANEDIOL, NAD-dependent epimerase/dehydratase, ... | | Authors: | Carbone, V, Schofield, L.R, Sang, C, Sutherland-Smith, A.J, Ronimus, R.S. | | Deposit date: | 2018-06-07 | | Release date: | 2018-10-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Structural determination of archaeal UDP-N-acetylglucosamine 4-epimerase from Methanobrevibacter ruminantium M1 in complex with the bacterial cell wall intermediate UDP-N-acetylmuramic acid.
Proteins, 86, 2018
|
|
6DPZ
 
 | | X-ray crystal structure of AmpC beta-lactamase with inhibitor | | Descriptor: | (1R,2S)-2-{[(pyrrolidin-1-yl)sulfonyl]amino}cyclooctane-1-carboxylic acid, Beta-lactamase | | Authors: | Singh, I. | | Deposit date: | 2018-06-09 | | Release date: | 2018-07-04 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Ultra-large library docking for discovering new chemotypes.
Nature, 566, 2019
|
|
6QZJ
 
 | | Crystal structure of human Aquaporin 7 at 2.2 A resolution | | Descriptor: | Aquaporin-7, GLYCEROL, PHOSPHATE ION | | Authors: | de Mare, S.W.-H, Venskutonyte, R, Eltschkner, S, Lindkvist-Petersson, K. | | Deposit date: | 2019-03-11 | | Release date: | 2019-11-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Basis for Glycerol Efflux and Selectivity of Human Aquaporin 7.
Structure, 28, 2020
|
|
6DRU
 
 | | Xylosidase from Aspergillus niger | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Glycosyl hydrolases family 31 family protein, ... | | Authors: | Cao, H, Xu, W, Betancourt, M, Walton, J.D, Brumm, P, Phillips Jr, G.N, Enzyme Discovery for Natural Product Biosynthesis (NatPro) | | Deposit date: | 2018-06-13 | | Release date: | 2018-08-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of alpha-Xylosidase fromAspergillus nigerin Complex with a Hydrolyzed Xyloglucan Product and New Insights in Accurately Predicting Substrate Specificities of GH31 Family Glycosidases.
Acs Sustain Chem Eng, 8, 2020
|
|
6QLW
 
 | | Cathepsin-K in complex with MIV-710 | | Descriptor: | CHLORIDE ION, Cathepsin K, SODIUM ION, ... | | Authors: | Derbyshire, D.J. | | Deposit date: | 2019-02-01 | | Release date: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Successful development of 3-oxohexahydrofuropyrrole amino acid amides as inhibitors of Cathepsin-K.
To Be Published
|
|
6QPL
 
 | | Crystal structure of Spindlin1 in complex with the inhibitor MS31 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Johansson, C, Krojer, T, Xiong, Y, Jin, J, Arrowsmith, C.H, Bountra, C, Edwards, A, Oppermann, U.C.T. | | Deposit date: | 2019-02-14 | | Release date: | 2019-07-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Discovery of a Potent and Selective Fragment-like Inhibitor of Methyllysine Reader Protein Spindlin 1 (SPIN1).
J.Med.Chem., 62, 2019
|
|
6D3X
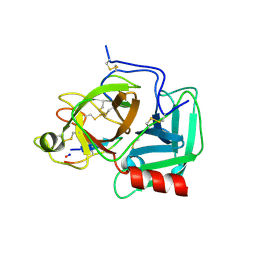 
 | |
