2OEU
 
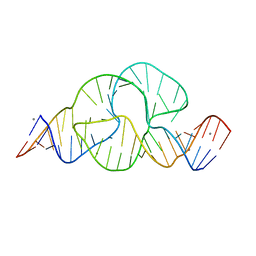 | | Full-length hammerhead ribozyme with Mn(II) bound | | Descriptor: | 5'-R(*GP*GP*CP*GP*UP*(OMC)P*CP*UP*GP*GP*(5BU)P*AP*UP*CP*CP*AP*AP*UP*CP*(DC))-3', Hammerhead Ribozyme, MANGANESE (II) ION | | Authors: | Martick, M, Scott, W.G. | | Deposit date: | 2007-01-01 | | Release date: | 2007-02-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Solvent structure and hammerhead ribozyme catalysis.
Chem.Biol., 15, 2008
|
|
5O7N
 
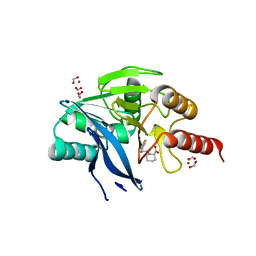 | | Beta-lactamase VIM-2 in complex with (2R)-1-(2-Benzyl-3-mercaptopropanoyl)piperidine-2-carboxylic acid | | Descriptor: | (2~{R})-1-[(2~{S})-2-(phenylmethyl)-3-sulfanyl-propanoyl]piperidine-2-carboxylic acid, 1,2-ETHANEDIOL, Beta-lactamase VIM-2, ... | | Authors: | Buettner, D, Kramer, J.S, Pogoryelov, D, Proschak, E. | | Deposit date: | 2017-06-09 | | Release date: | 2018-06-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Challenges in the Development of a Thiol-Based Broad-Spectrum Inhibitor for Metallo-beta-Lactamases.
Acs Infect Dis., 4, 2018
|
|
1XZN
 
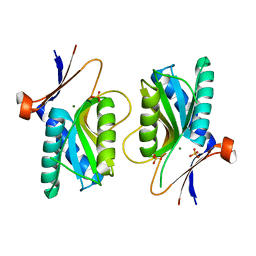 | | PYRR, THE REGULATOR OF THE PYRIMIDINE BIOSYNTHETIC OPERON IN BACILLUS CALDOLYTICUS, sulfate-bound form | | Descriptor: | MAGNESIUM ION, PyrR bifunctional protein, SULFATE ION | | Authors: | Chander, P, Halbig, K.M, Miller, J.K, Fields, C.J, Bonner, H.K, Grabner, G.K, Switzer, R.L, Smith, J.L. | | Deposit date: | 2004-11-12 | | Release date: | 2005-03-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Structure of the Nucleotide Complex of PyrR, the pyr Attenuation Protein from Bacillus caldolyticus, Suggests Dual Regulation by Pyrimidine and Purine Nucleotides.
J.Bacteriol., 187, 2005
|
|
1XNC
 
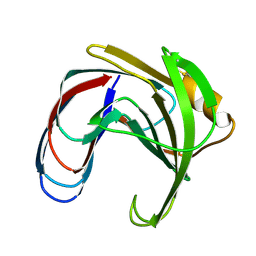 | |
3MWS
 
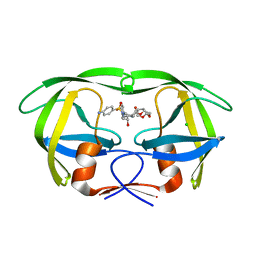 | | Crystal Structure of Group N HIV-1 Protease | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, CHLORIDE ION, HIV-1 Protease | | Authors: | Sayer, J.M, Agniswamy, J, Weber, I.T, Louis, J.M. | | Deposit date: | 2010-05-06 | | Release date: | 2011-03-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.09 Å) | | Cite: | Autocatalytic maturation, physical/chemical properties, and crystal structure of group N HIV-1 protease: relevance to drug resistance.
Protein Sci., 19, 2010
|
|
2O3E
 
 | |
1XZ8
 
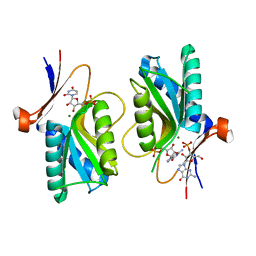 | | Pyrr, The Regulator Of The Pyrimidine Biosynthetic Operon In Bacillus caldolyticus, Nucleotide-bound form | | Descriptor: | GUANOSINE-3'-MONOPHOSPHATE, GUANOSINE-5'-MONOPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Chander, P, Halbig, K.M, Miller, J.K, Fields, C.J, Bonner, H.K, Grabner, G.K, Switzer, R.L, Smith, J.L. | | Deposit date: | 2004-11-11 | | Release date: | 2005-03-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of the Nucleotide Complex of PyrR, the pyr Attenuation Protein from Bacillus caldolyticus, Suggests Dual Regulation by Pyrimidine and Purine Nucleotides.
J.Bacteriol., 187, 2005
|
|
2OB2
 
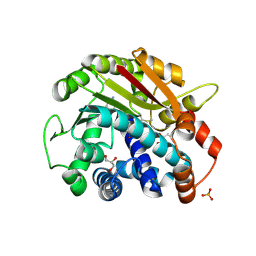 | | ppm1 in the absence of 1,8-ANS (cf 1JD) | | Descriptor: | GLYCEROL, Leucine carboxyl methyltransferase 1, PHOSPHATE ION, ... | | Authors: | Groves, M.R, Mueller, I.B, Kreplin, X, Mueller-Dieckmann, J. | | Deposit date: | 2006-12-18 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | A method for the general identification of protein crystals in crystallization experiments using a noncovalent fluorescent dye.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
3N9R
 
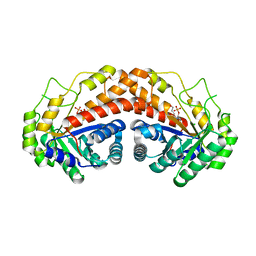 | | Class II fructose-1,6-bisphosphate aldolase from helicobacter pylori in complex with N-(4-hydroxybutyl)-phosphoglycolohydroxamic acid, a competitive inhibitor | | Descriptor: | 2-[hydroxy(4-hydroxybutyl)amino]-2-oxoethyl dihydrogen phosphate, CHLORIDE ION, Fructose-bisphosphate aldolase, ... | | Authors: | Coincon, M, Sygusch, S. | | Deposit date: | 2010-05-31 | | Release date: | 2010-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Rational Design, Synthesis, and Evaluation of New Selective Inhibitors of Microbial Class II (Zinc Dependent) Fructose Bis-phosphate Aldolases.
J.Med.Chem., 53, 2010
|
|
2OK6
 | |
2OB1
 
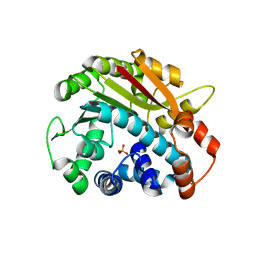 | | ppm1 with 1,8-ANS | | Descriptor: | Leucine carboxyl methyltransferase 1, PHOSPHATE ION | | Authors: | Groves, M.R. | | Deposit date: | 2006-12-18 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A method for the general identification of protein crystals in crystallization experiments using a noncovalent fluorescent dye.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
1ZPK
 
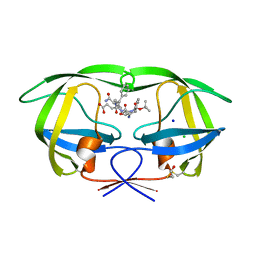 | | Crystal structure of the complex of mutant HIV-1 protease (A71V, V82T, I84V) with a hydroxyethylamine peptidomimetic inhibitor BOC-PHE-PSI[R-CH(OH)CH2NH]-PHE-GLU-PHE-NH2 | | Descriptor: | CHLORIDE ION, N-{(2R,3S)-3-[(tert-butoxycarbonyl)amino]-2-hydroxy-4-phenylbutyl}-L-phenylalanyl-L-alpha-glutamyl-L-phenylalaninamide, PROTEASE RETROPEPSIN, ... | | Authors: | Duskova, J, Skalova, T, Dohnalek, J, Petrokova, H, Hasek, J. | | Deposit date: | 2005-05-17 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Mutational Study of Pseudopeptide Inhibitor Binding to HIV-1 Protease; Analysis of Four X-ray Structures
To be Published
|
|
1ZAQ
 
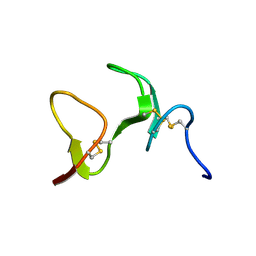 | |
3OG9
 
 | | Structure of YahD with Malic acid | | Descriptor: | D-MALATE, protein yahD a copper inducible hydrolase | | Authors: | Martinez Font, J, Mancini, S, Tauberger, E, Moniot, S. | | Deposit date: | 2010-08-16 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Regulation and structure of YahD, a copper inducible alpha/beta hydrolase of Lactococcus lactis IL1403
Fems Microbiol.Lett., 314, 2011
|
|
2OIZ
 | |
2PUG
 
 | | CRYSTAL STRUCTURE OF THE LACI FAMILY MEMBER, PURR, BOUND TO DNA: MINOR GROOVE BINDING BY ALPHA HELICES | | Descriptor: | DNA (5'-D(*TP*AP*CP*GP*CP*AP*AP*AP*CP*GP*TP*TP*TP*GP*CP*GP*T )-3'), HYPOXANTHINE, PROTEIN (PURINE REPRESSOR) | | Authors: | Lu, F, Schumacher, M.A, Arvidson, D.N, Haldimann, A, Wanner, B.L, Zalkin, H, Brennan, R.G. | | Deposit date: | 1997-10-04 | | Release date: | 1998-05-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-based redesign of corepressor specificity of the Escherichia coli purine repressor by substitution of residue 190.
Biochemistry, 37, 1998
|
|
1Y3G
 
 | | Crystal Structure of a Silanediol Protease Inhibitor Bound to Thermolysin | | Descriptor: | (2S)-2-{[(AMINOMETHYL)(DIHYDROXY)SILYL]METHYL}-4-METHYLPENTANAL, 3-PHENYLPROPANAL, CALCIUM ION, ... | | Authors: | Juers, D.H, Kim, J, Matthews, B.W, Sieburth, S.M. | | Deposit date: | 2004-11-24 | | Release date: | 2006-01-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Analysis of Silanediols as Transition-State-Analogue Inhibitors of the Benchmark Metalloprotease Thermolysin(,).
Biochemistry, 44, 2005
|
|
3OOH
 | | Crystal structure of E. Coli purine nucleoside phosphorylase with PO4 | | Descriptor: | PHOSPHATE ION, Purine nucleoside phosphorylase deoD-type | | Authors: | Mikleusevic, G, Stefanic, Z, Narzyk, M, Wielgus-Kutrowska, B, Bzowska, A, Luic, M. | | Deposit date: | 2010-08-31 | | Release date: | 2011-07-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Validation of the catalytic mechanism of Escherichia coli purine nucleoside phosphorylase by structural and kinetic studies.
Biochimie, 93, 2011
|
|
2PFW
 | |
1Y58
 
 | |
1ZLF
 
 | | Crystal structure of a complex of mutant HIV-1 protease (A71V, V82T, I84V) with a hydroxyethylamine peptidomimetic inhibitor | | Descriptor: | N-{(2R,3S)-3-[(tert-butoxycarbonyl)amino]-2-hydroxy-4-phenylbutyl}-L-phenylalanyl-L-glutaminyl-L-phenylalaninamide, PROTEASE RETROPEPSIN | | Authors: | Skalova, T, Dohnalek, J, Duskova, J, Petrokova, H, Hasek, J. | | Deposit date: | 2005-05-06 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | HIV-1 protease mutations and inhibitor modifications monitored on a series of complexes. Structural basis for the effect of the A71V mutation on the active site
J.Med.Chem., 49, 2006
|
|
3Q48
 
 | | Crystal structure of Pseudomonas aeruginosa CupB2 chaperone | | Descriptor: | Chaperone CupB2 | | Authors: | Cai, X, Wang, R, Filloux, A, Waksman, G, Meng, G. | | Deposit date: | 2010-12-23 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and functional characterization of Pseudomonas aeruginosa CupB chaperones
Plos One, 6, 2011
|
|
2OMA
 
 | |
1ZJD
 
 | | Crystal Structure of the Catalytic Domain of Coagulation Factor XI in Complex with Kunitz Protease Inhibitor Domain of Protease Nexin II | | Descriptor: | Catalytic Domain of Coagulation Factor XI, Kunitz Protease Inhibitory Domain of Protease Nexin II | | Authors: | Jin, L, Navaneetham, D, Pandey, P, Strickler, J.E, Babine, R.E, Walsh, P.N, Abdel-Meguid, S.S. | | Deposit date: | 2005-04-28 | | Release date: | 2005-08-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and Mutational Analyses of the Molecular Interactions between the Catalytic Domain of Factor XIa and the Kunitz Protease Inhibitor Domain of Protease Nexin 2
J.Biol.Chem., 280, 2005
|
|
5O1Q
 
 | | LysF1 sh3b domain structure | | Descriptor: | sh3b domain | | Authors: | Benesik, M, Novacek, J, Janda, L, Dopitova, R, Pernisova, M, Melkova, K, Tisakova, L, Doskar, J, Zidek, L, Hejatko, J, Pantucek, R. | | Deposit date: | 2017-05-19 | | Release date: | 2017-09-20 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Role of SH3b binding domain in a natural deletion mutant of Kayvirus endolysin LysF1 with a broad range of lytic activity.
Virus Genes, 54, 2018
|
|
