4A1O
 
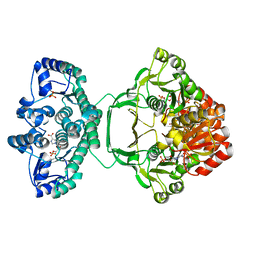 | | Crystal structure of Mycobacterium tuberculosis PurH complexed with AICAR and a novel nucleotide CFAIR, at 2.48 A resolution. | | Descriptor: | 5-(FORMYLAMINO)-1-(5-O-PHOSPHONO-BETA-D-RIBOFURANOSYL)-1H-IMIDAZOLE-4-CARBOXYLIC ACID, AMINOIMIDAZOLE 4-CARBOXAMIDE RIBONUCLEOTIDE, BIFUNCTIONAL PURINE BIOSYNTHESIS PROTEIN PURH, ... | | Authors: | Le Nours, J, Bulloch, E.M.M, Zhang, Z, Greenwood, D.R, Middleditch, M.J, Dickson, J.M.J, Baker, E.N. | | Deposit date: | 2011-09-17 | | Release date: | 2011-09-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Structural Analyses of a Purine Biosynthetic Enzyme from Mycobacterium Tuberculosis Reveal a Novel Bound Nucleotide.
J.Biol.Chem., 286, 2011
|
|
6TWF
 
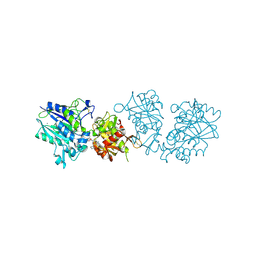 | | Human CD73 (ecto 5'-nucleotidase) in complex with PSB12604 (an AOPCP derivative, compound 21 in publication) in the closed state | | Descriptor: | 5'-nucleotidase, CALCIUM ION, ZINC ION, ... | | Authors: | Pippel, J, Strater, N. | | Deposit date: | 2020-01-13 | | Release date: | 2020-02-19 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | 2-Substituted alpha , beta-Methylene-ADP Derivatives: Potent Competitive Ecto-5'-nucleotidase (CD73) Inhibitors with Variable Binding Modes.
J.Med.Chem., 63, 2020
|
|
8W3H
 
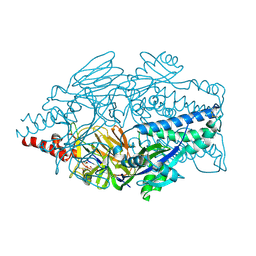 | | Crystal structure of prefusion-stabilized RSV F protein UFCR1-iSS | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Prefusion-stabilized RSV F protein UFCR1-iSS | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
8W3K
 
 | | Crystal structure of prefusion-stabilized RSV F protein UFCR2-iSS-NQ | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Prefusion-stabilized RSV F protein UFCR2-iSS-NQ | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
8W3E
 
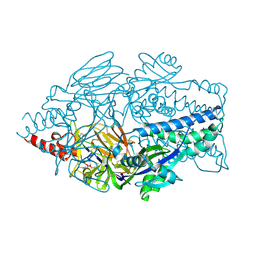 | | Crystal structure of prefusion-stabilized RSV F protein UFCR1 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
8W3G
 
 | | Crystal structure of prefusion-stabilized RSV F protein UFCR1(1TD0) | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Prefusion-stabilized RSV F protein UFCR1(1TD0) | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
3L7I
 
 | |
8W3F
 
 | | Crystal structure of prefusion-stabilized RSV F protein UFCR1-P2-NQ | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Prefusion-stabilized RSV F protein UFCR1-P2-NQ | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
8W3J
 
 | | Crystal structure of prefusion-stabilized RSV F protein UFCR2-iSS | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Prefusion-stabilized RSV F protein UFCR2-iSS | | Authors: | Lee, Y.Z, Stanfield, R.L, Wilson, I.A, Zhu, J. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Nat Commun, 15, 2024
|
|
6U01
 
 | | Dihydrodipicolinate synthase (DHDPS) from C.jejuni, N84D mutant with pyruvate bound in the active site | | Descriptor: | 1,2-ETHANEDIOL, 4-hydroxy-tetrahydrodipicolinate synthase, ACETATE ION, ... | | Authors: | Saran, S, Majdi Yazdi, M, Lehnert, L, Palmer, D.R.J, Sanders, D.A.R. | | Deposit date: | 2019-08-13 | | Release date: | 2019-12-04 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Asparagine-84, a regulatory allosteric site residue, helps maintain the quaternary structure of Campylobacter jejuni dihydrodipicolinate synthase.
J.Struct.Biol., 209, 2020
|
|
1MUX
 
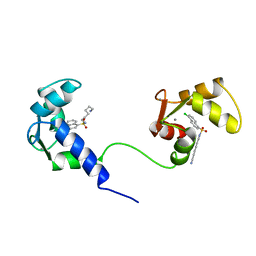 | | SOLUTION NMR STRUCTURE OF CALMODULIN/W-7 COMPLEX: THE BASIS OF DIVERSITY IN MOLECULAR RECOGNITION, 30 STRUCTURES | | Descriptor: | CALCIUM ION, CALMODULIN, N-(6-AMINOHEXYL)-5-CHLORO-1-NAPHTHALENESULFONAMIDE | | Authors: | Osawa, M, Swindells, M.B, Tanikawa, J, Tanaka, T, Mase, T, Furuya, T, Ikura, M. | | Deposit date: | 1997-09-06 | | Release date: | 1998-10-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of calmodulin-W-7 complex: the basis of diversity in molecular recognition.
J.Mol.Biol., 276, 1998
|
|
5DX0
 
 | |
8DUA
 | | SIV E660.CR54 SOS-2P Env Trimer with ITS92.02 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Envelope glycoprotein gp120, Envelope glycoprotein gp41, ... | | Authors: | Gorman, J, Kwong, P.D. | | Deposit date: | 2022-07-27 | | Release date: | 2022-10-26 | | Last modified: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (4.32 Å) | | Cite: | Cryo-EM structures of prefusion SIV envelope trimer.
Nat.Struct.Mol.Biol., 29, 2022
|
|
5DXJ
 
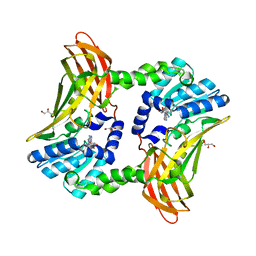 | | Crystal structure of CARM1 and sinefungin | | Descriptor: | GLYCEROL, Histone-arginine methyltransferase CARM1, SINEFUNGIN | | Authors: | Boriack-Sjodin, P.A. | | Deposit date: | 2015-09-23 | | Release date: | 2015-11-25 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural Insights into Ternary Complex Formation of Human CARM1 with Various Substrates.
Acs Chem.Biol., 11, 2016
|
|
4ZZI
 
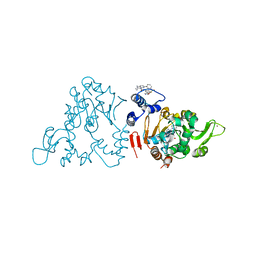 | | SIRT1/Activator/Inhibitor Complex | | Descriptor: | (3S)-1,3-dimethyl-N-[3-(1,3-oxazol-5-yl)phenyl]-6-[3-(trifluoromethyl)phenyl]-2,3-dihydropyrido[2,3-b]pyrazine-4(1H)-carboxamide, 4-(4-{2-[(methylsulfonyl)amino]ethyl}piperidin-1-yl)thieno[3,2-d]pyrimidine-6-carboxamide, NAD-dependent protein deacetylase sirtuin-1, ... | | Authors: | Dai, H. | | Deposit date: | 2015-05-22 | | Release date: | 2015-07-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.7346 Å) | | Cite: | Crystallographic structure of a small molecule SIRT1 activator-enzyme complex.
Nat Commun, 6, 2015
|
|
4AYG
 
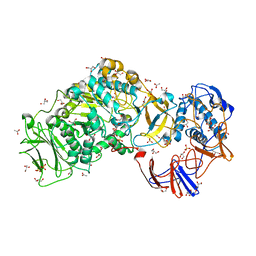 | | Lactobacillus reuteri N-terminally truncated glucansucrase GTF180 in orthorhombic apo-form | | Descriptor: | ACETIC ACID, CALCIUM ION, GLUCANSUCRASE, ... | | Authors: | Pijning, T, Vujicic-Zagar, A, Kralj, S, Dijkhuizen, L, Dijkstra, B.W. | | Deposit date: | 2012-06-21 | | Release date: | 2013-07-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Flexibility of Truncated and Full-Length Glucansucrase Gtf180 Enzymes from Lactobacillus Reuteri 180.
FEBS J., 281, 2014
|
|
5Y9T
 
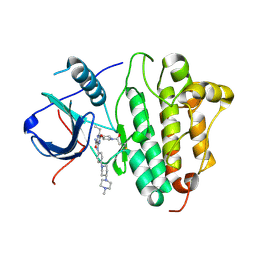 | | Crystal Structure of EGFR T790M mutant in complex with naquotinib | | Descriptor: | 6-ethyl-3-[[4-[4-(4-methylpiperazin-1-yl)piperidin-1-yl]phenyl]amino]-5-[(3R)-1-prop-2-enoylpyrrolidin-3-yl]oxy-pyrazin e-2-carboxamide, Epidermal growth factor receptor | | Authors: | Mimasu, S, Tomimoto, Y, Maiko, I, Yasushi, A, Tatsuya, N. | | Deposit date: | 2017-08-28 | | Release date: | 2018-07-11 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Pharmacological and Structural Characterizations of Naquotinib, a Novel Third-Generation EGFR Tyrosine Kinase Inhibitor, inEGFR-Mutated Non-Small Cell Lung Cancer.
Mol. Cancer Ther., 17, 2018
|
|
5JAU
 
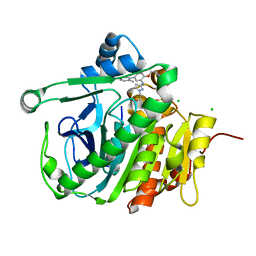 | |
6KVM
 
 | | Crystal structure of Chicken MHC Class II for 1.9 angstrom | | Descriptor: | GLYCEROL, MHC class II alpha chain, MHC class II beta chain 2, ... | | Authors: | Zhang, L, Xia, C. | | Deposit date: | 2019-09-04 | | Release date: | 2019-12-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.897 Å) | | Cite: | A Newly Recognized Pairing Mechanism of the alpha- and beta-Chains of the Chicken Peptide-MHC Class II Complex.
J Immunol., 204, 2020
|
|
1NHK
 
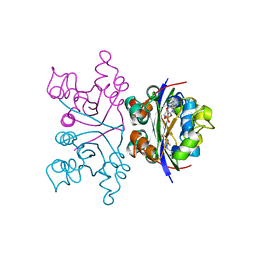 | |
5DR4
 
 | | Endothiapepsin in complex with fragment 231 | | Descriptor: | 7-methyl-3,4-dihydro-2~{H}-pyrido[1,2-a]pyrimidin-3-ol, ACETATE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Schiebel, J, Heine, A, Klebe, G. | | Deposit date: | 2015-09-15 | | Release date: | 2016-09-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystallographic Fragment Screening of an Entire Library
To Be Published
|
|
8W10
 
 | | Plasmodium vivax PMX-MK7602 inhibitor complex | | Descriptor: | (2R,3S)-3-[(2S)-2-amino-4,4-diethyl-6-oxo-1,3-diazinan-1-yl]-N-[(1R,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl]-2-(methoxymethyl)-2-methyl-2,3-dihydro-1-benzofuran-5-carboxamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hodder, A.N, Scally, S.W, Cowman, A.F. | | Deposit date: | 2024-02-14 | | Release date: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Plasmodium vivax PMX-MK7602 inhibitor complex
To Be Published
|
|
3LD9
 
 | |
3LE9
 
 | |
3FOE
 
 | | Structural insight into the quinolone-DNA cleavage complex of type IIA topoisomerases | | Descriptor: | 7-[(3R)-3-aminopyrrolidin-1-yl]-8-chloro-1-cyclopropyl-6-fluoro-4-oxo-1,4-dihydroquinoline-3-carboxylic acid, DNA (5'-D(P*AP*CP*CP*AP*AP*GP*GP*TP*CP*AP*TP*GP*AP*AP*T)-3'), DNA (5'-D(P*AP*GP*TP*CP*AP*TP*TP*CP*AP*TP*GP*AP*CP*CP*TP*TP*GP*GP*T)-3'), ... | | Authors: | Laponogov, I, Sohi, M.K, Veselkov, D.A, Pan, X.-S, Sawhney, R, Thompson, A.W, McAuley, K.E, Fisher, L.M, Sanderson, M.R. | | Deposit date: | 2008-12-30 | | Release date: | 2009-02-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (4.001 Å) | | Cite: | Structural insight into the quinolone-DNA cleavage complex of type IIA topoisomerases
Nat.Struct.Mol.Biol., 16, 2009
|
|
