6W0B
 
 | | Open-gate KcsA soaked in 2 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.604 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
5VDK
 
 | | Crystal structure of human WEE2 kinase domain in complex with MK1775 | | Descriptor: | 1-[6-(2-hydroxypropan-2-yl)pyridin-2-yl]-6-{[4-(4-methylpiperazin-1-yl)phenyl]amino}-2-(prop-2-en-1-yl)-1,2-dihydro-3H-pyrazolo[3,4-d]pyrimidin-3-one, PHOSPHATE ION, Wee1-like protein kinase 2 | | Authors: | Zhu, J.-Y, Schonbrunn, E. | | Deposit date: | 2017-04-03 | | Release date: | 2017-08-23 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural Basis of Wee Kinases Functionality and Inactivation by Diverse Small Molecule Inhibitors.
J. Med. Chem., 60, 2017
|
|
5IFZ
 
 | |
8VUK
 | |
7M3H
 
 | |
6W0M
 
 | | Human 8-oxoguanine glycosylase crosslinked with oxoG lesion containing DNA | | Descriptor: | 2-(2-ethoxyethoxy)ethanethiol, DNA (5'-D(P*CP*GP*TP*CP*CP*AP*(8OG)P*GP*TP*CP*TP*AP*C)-3'), DNA (5'-D(P*GP*GP*TP*AP*GP*AP*CP*CP*TP*GP*GP*AP*CP*GP*C)-3'), ... | | Authors: | Shigdel, U, Verdine, G. | | Deposit date: | 2020-03-02 | | Release date: | 2020-09-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | The trajectory of intrahelical lesion recognition and extrusion by the human 8-oxoguanine DNA glycosylase.
Nat Commun, 11, 2020
|
|
5VDW
 
 | | Human cyclic GMP-AMP synthase (cGAS) in complex with Compound F1 | | Descriptor: | Cyclic GMP-AMP synthase, ZINC ION, [2-(1,3-thiazol-4-yl)-1H-benzimidazol-1-yl]acetic acid | | Authors: | Byrnes, L.J, Hall, J.D. | | Deposit date: | 2017-04-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.711 Å) | | Cite: | The catalytic mechanism of cyclic GMP-AMP synthase (cGAS) and implications for innate immunity and inhibition.
Protein Sci., 26, 2017
|
|
6W0R
 
 | |
5IKD
 
 | |
7M4E
 
 | | DNA Polymerase Lambda, dCTP:At Mg2+ Reaction State Ternary Complex, 120 min | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYCYTIDINE-5'-TRIPHOSPHATE, DNA (5'-D(*CP*AP*GP*TP*AP*CP*C)-3'), ... | | Authors: | Jamsen, J.A, Wilson, S.H. | | Deposit date: | 2021-03-21 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.903 Å) | | Cite: | Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints.
Nat Commun, 13, 2022
|
|
8VVX
 
 | |
5VJ0
 
 | |
5VJS
 
 | | De Novo Photosynthetic Reaction Center Protein Equipped with Heme B, a synthetic Zn porphyrin, and Zn(II) cations | | Descriptor: | CHLORIDE ION, PROTOPORPHYRIN IX CONTAINING FE, Reaction Center Maquette, ... | | Authors: | Ennist, N.M, Dutton, P.L, Stayrook, S.E, Moser, C.C. | | Deposit date: | 2017-04-19 | | Release date: | 2018-04-25 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | De novo protein design of photochemical reaction centers.
Nat Commun, 13, 2022
|
|
7M45
 
 | | DNA Polymerase Lambda, TTP:At Mg2+ Reaction State Ternary Complex, 120 sec | | Descriptor: | 1,2-ETHANEDIOL, DNA (5'-D(*CP*AP*GP*TP*AP*CP*T)-3'), DNA (5'-D(*CP*GP*GP*CP*AP*GP*TP*AP*CP*TP*G)-3'), ... | | Authors: | Jamsen, J.A, Wilson, S.H. | | Deposit date: | 2021-03-21 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.889 Å) | | Cite: | Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints.
Nat Commun, 13, 2022
|
|
6W15
 
 | |
8VHV
 
 | |
7M4G
 
 | | DNA Polymerase Lambda, dCTP:At Mg2+ Product State Ternary Complex, 960 min | | Descriptor: | 1,2-ETHANEDIOL, DNA (5'-D(*CP*AP*GP*TP*AP*CP*C)-3'), DNA (5'-D(*CP*GP*GP*CP*AP*GP*TP*AP*CP*TP*G)-3'), ... | | Authors: | Jamsen, J.A, Wilson, S.H. | | Deposit date: | 2021-03-21 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints.
Nat Commun, 13, 2022
|
|
8VWA
 
 | | CryoEM Structure of a FtsH Helical Assembly in the Presence of ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, ATP-dependent zinc metalloprotease FtsH, MAGNESIUM ION, ... | | Authors: | Li, Y, Zhu, J, Zhang, Z, Wang, F, Egelman, E.H, Tezcan, F.A. | | Deposit date: | 2024-01-31 | | Release date: | 2025-01-15 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Transforming an ATP-dependent enzyme into a dissipative, self-assembling system.
Nat.Chem.Biol., 21, 2025
|
|
6W1F
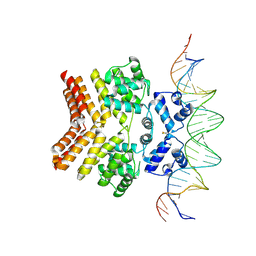 
 | |
7M4D
 
 | | DNA Polymerase Lambda, dCTP:At Ca2+ Ground State Ternary Complex | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYCYTIDINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Jamsen, J.A, Wilson, S.H. | | Deposit date: | 2021-03-21 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.817 Å) | | Cite: | Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints.
Nat Commun, 13, 2022
|
|
5IGF
 
 | |
6W2V
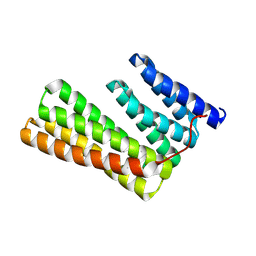 
 | | Junction 23, DHR14-DHR18 | | Descriptor: | Junction 23 DHR14-DHR18 | | Authors: | Bick, M.J, Brunette, T.J, Baker, D. | | Deposit date: | 2020-03-08 | | Release date: | 2020-04-15 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Modular repeat protein sculpting using rigid helical junctions.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
7M4K
 
 | | DNA Polymerase Lambda, TTPaS:At Ca2+ Ground State Ternary Complex | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, DNA (5'-D(*CP*AP*GP*TP*AP*C)-3'), ... | | Authors: | Jamsen, J.A, Wilson, S.H. | | Deposit date: | 2021-03-21 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.719 Å) | | Cite: | Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints.
Nat Commun, 13, 2022
|
|
8W3M
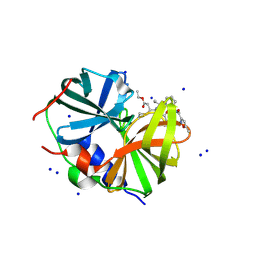 
 | | Crystal Structure of Enterovirus 68 3C Protease with AG7404 at 1.97 Angstroms | | Descriptor: | Peptidase C3, SODIUM ION, ethyl (4R)-4-({(2S)-2-[3-{[(5-methyl-1,2-oxazol-3-yl)carbonyl]amino}-2-oxopyridin-1(2H)-yl]pent-4-ynoyl}amino)-5-[(3S)-2-oxopyrrolidin-3-yl]pentanoate | | Authors: | Azzolino, V.N, Shaqra, A.M, Schiffer, C.A. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-15 | | Last modified: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural Analysis of Inhibitor Binding to Enterovirus-D68 3C Protease.
Viruses, 17, 2025
|
|
6W37
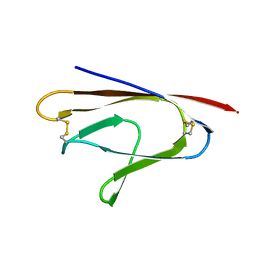 
 | |
