7CSY
 
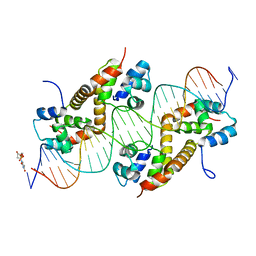 | | Pseudomonas aeruginosa antitoxin HigA with higBA promoter | | Descriptor: | DNA (28-MER), DNA (29-MER), HTH cro/C1-type domain-containing protein, ... | | Authors: | Song, Y.J, Luo, G.H, Bao, R. | | Deposit date: | 2020-08-17 | | Release date: | 2021-01-13 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Pseudomonas aeruginosa antitoxin HigA functions as a diverse regulatory factor by recognizing specific pseudopalindromic DNA motifs.
Environ.Microbiol., 23, 2021
|
|
7CWC
 
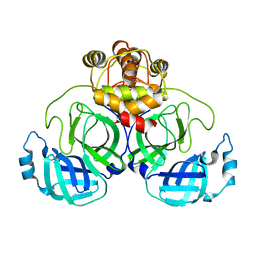 | |
7CWB
 
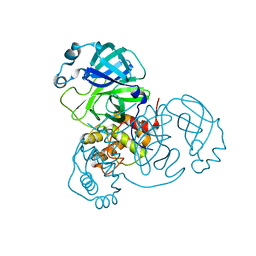 | |
4LWG
 
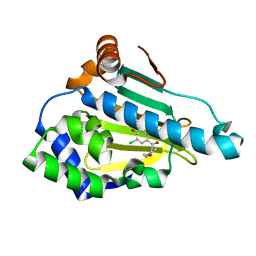 | | Crystal Structure of the human Hsp90-alpha N-domain bound to the hsp90 inhibitor FJ4 | | Descriptor: | 1-(5-chloro-2,4-dihydroxyphenyl)-2-(4-methoxyphenyl)ethanone, Heat shock protein HSP 90-alpha | | Authors: | Li, J, Shi, F, Xiong, B, He, J. | | Deposit date: | 2013-07-27 | | Release date: | 2014-07-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.599 Å) | | Cite: | Discovery of potent N-(isoxazol-5-yl)amides as HSP90 inhibitors.
Eur.J.Med.Chem., 87, 2014
|
|
4LWF
 
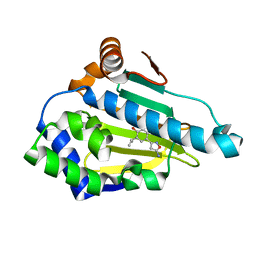 | | Crystal Structure of the human Hsp90-alpha N-domain bound to the hsp90 inhibitor FJ3 | | Descriptor: | 4-(5-amino-1,2-oxazol-3-yl)-6-(propan-2-yl)benzene-1,3-diol, Heat shock protein HSP 90-alpha | | Authors: | Li, J, Shi, F, Xiong, B, He, J. | | Deposit date: | 2013-07-27 | | Release date: | 2014-07-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Discovery of potent N-(isoxazol-5-yl)amides as HSP90 inhibitors.
Eur.J.Med.Chem., 87, 2014
|
|
5F52
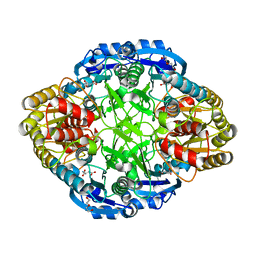 
 | | Erwinia chrysanthemi L-asparaginase + Aspartic acid | | Descriptor: | ASPARTIC ACID, DI(HYDROXYETHYL)ETHER, L-asparaginase | | Authors: | Nguyen, H.A, Lavie, A. | | Deposit date: | 2015-12-04 | | Release date: | 2016-04-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structural Insight into Substrate Selectivity of Erwinia chrysanthemi l-Asparaginase.
Biochemistry, 55, 2016
|
|
2RFV
 
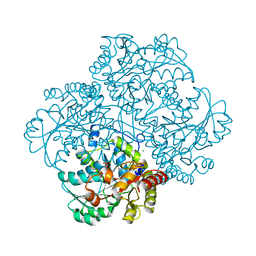 | | High resolution structure of L-methionine gamma-lyase from Citrobacter freundii | | Descriptor: | CHLORIDE ION, Methionine gamma-lyase | | Authors: | Nikulin, A.D, Revtovich, S.V, Morozova, E.A, Nevskaya, N.A, Nikonov, S.V, Garber, M.B, Demidkina, T.V. | | Deposit date: | 2007-10-02 | | Release date: | 2008-08-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.355 Å) | | Cite: | High-resolution structure of methionine gamma-lyase from Citrobacter freundii.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
4LWH
 
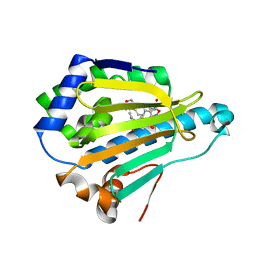 | | Crystal Structure of the human Hsp90-alpha N-domain bound to the hsp90 inhibitor FJ5 | | Descriptor: | Heat shock protein HSP 90-alpha, N-{3-[2,4-dihydroxy-5-(propan-2-yl)phenyl]-4-[4-(morpholin-4-ylmethyl)phenyl]-1,2-oxazol-5-yl}cyclopropanecarboxamide | | Authors: | Li, J, Shi, F, Xiong, B, He, J. | | Deposit date: | 2013-07-27 | | Release date: | 2014-07-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Discovery of potent N-(isoxazol-5-yl)amides as HSP90 inhibitors.
Eur.J.Med.Chem., 87, 2014
|
|
6LJ9
 
 | |
7C4R
 
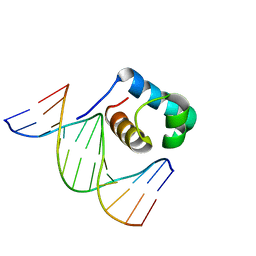 | | Crystal structure of hydrogen peroxide treated zebrafish TRF2 complexed with DNA | | Descriptor: | DNA (5'-D(*D*CP*DP*CP*DP*CP*DP*TP*DP*AP*DP*AP*DP*CP*DP*CP*DP*CP*DP*TP*DP*AP*DP*A)-3'), DNA (5'-D(*D*TP*DP*TP*DP*AP*DP*GP*DP*GP*DP*GP*DP*TP*DP*TP*DP*AP*DP*G)-3'), DNA (5'-D(*D*TP*DP*TP*DP*AP*DP*GP*DP*GP*DP*GP*DP*TP*DP*TP*DP*AP*DP*GP*DP*GP*DP*G)-3'), ... | | Authors: | Jin, Z, Park, J.H, Yun, J.H, Park, S.Y, Lee, W. | | Deposit date: | 2020-05-18 | | Release date: | 2021-05-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Crystal structure of hydrogen peroxide treated zebrafish TRF2 myb-domain complexed with DNA
To Be Published
|
|
7CC2
 
 | |
7EHJ
 
 | | human MTHFD2 in complex with compound 21, cofactor and phosphate. | | Descriptor: | (2S)-2-[[4-[(4-azanyl-6-oxidanyl-pyrimidin-5-yl)carbamoylamino]phenyl]carbonylamino]pentanedioic acid, Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, mitochondrial, ... | | Authors: | Lee, L.C, Peng, Y.H, Wu, S.Y. | | Deposit date: | 2021-03-29 | | Release date: | 2021-08-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Xanthine Derivatives Reveal an Allosteric Binding Site in Methylenetetrahydrofolate Dehydrogenase 2 (MTHFD2).
J.Med.Chem., 64, 2021
|
|
7EHN
 
 | | Human MTHFD2 in complex with compound 21 and 9 | | Descriptor: | (2S)-2-[[4-[(4-azanyl-6-oxidanyl-pyrimidin-5-yl)carbamoylamino]phenyl]carbonylamino]pentanedioic acid, 3-[4-[[1-[(4-chloranyl-1H-indol-2-yl)methyl]-3,7-dimethyl-2,6-bis(oxidanylidene)purin-8-yl]amino]-6-methyl-pyrimidin-2-yl]propanoic acid, Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, ... | | Authors: | Lee, L.C, Peng, Y.H, Wu, S.Y. | | Deposit date: | 2021-03-30 | | Release date: | 2021-08-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Xanthine Derivatives Reveal an Allosteric Binding Site in Methylenetetrahydrofolate Dehydrogenase 2 (MTHFD2).
J.Med.Chem., 64, 2021
|
|
7EHM
 
 | | Human MTHFD2 in complex with compound 21 and 15 | | Descriptor: | (2S)-2-[[4-[(4-azanyl-6-oxidanyl-pyrimidin-5-yl)carbamoylamino]phenyl]carbonylamino]pentanedioic acid, (2S)-2-[[4-[[1-[(3,4-dichlorophenyl)methyl]-3,7-dimethyl-2,6-bis(oxidanylidene)purin-8-yl]amino]phenyl]carbonylamino]pentanedioic acid, Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, ... | | Authors: | Lee, L.C, Peng, Y.H, Wu, S.Y. | | Deposit date: | 2021-03-30 | | Release date: | 2021-08-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Xanthine Derivatives Reveal an Allosteric Binding Site in Methylenetetrahydrofolate Dehydrogenase 2 (MTHFD2).
J.Med.Chem., 64, 2021
|
|
7EHV
 
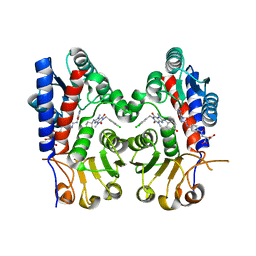 | | Human MTHFD2 in complex with compound 21 and 3 | | Descriptor: | (2S)-2-[[4-[(4-azanyl-6-oxidanyl-pyrimidin-5-yl)carbamoylamino]phenyl]carbonylamino]pentanedioic acid, 1-(3,4-dichlorobenzyl)-8-(((1R,4R)-4-hydroxycyclohexyl)amino)-3,7-dimethyl-3,7-dihydro-1H-purine-2,6-dione, Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, ... | | Authors: | Lee, L.C, Peng, Y.H, Wu, S.Y. | | Deposit date: | 2021-03-30 | | Release date: | 2021-08-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Xanthine Derivatives Reveal an Allosteric Binding Site in Methylenetetrahydrofolate Dehydrogenase 2 (MTHFD2).
J.Med.Chem., 64, 2021
|
|
5F8S
 
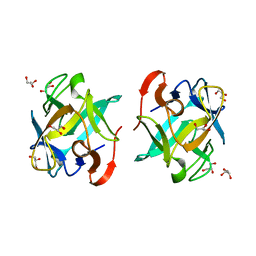 | | Crystal structure of a Crenomytilus grayanus lectin | | Descriptor: | GLYCEROL, GalNAc/Gal-specific lectin | | Authors: | Liao, J.-H, Huang, K.-F, Tu, I.-F, Lee, I.-M, Wu, S.-H. | | Deposit date: | 2015-12-09 | | Release date: | 2016-04-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | A Multivalent Marine Lectin from Crenomytilus grayanus Possesses Anti-cancer Activity through Recognizing Globotriose Gb3
J.Am.Chem.Soc., 138, 2016
|
|
6LJB
 
 | |
8U6G
 
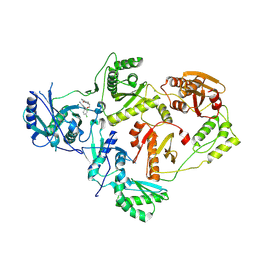 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with 3-(2-(2-(3-acryloyl-2-oxo-2,3-dihydro-1H-benzo[d]imidazol-1-yl)ethoxy)-4-chlorophenoxy)-5-chlorobenzonitrile (JLJ744), a non-nucleoside inhibitor | | Descriptor: | 3-chloro-5-{4-chloro-2-[2-(2-oxo-3-propanoyl-2,3-dihydro-1H-benzimidazol-1-yl)ethoxy]phenoxy}benzonitrile, MAGNESIUM ION, Reverse transcriptase/ribonuclease H, ... | | Authors: | Prucha, G, Carter, Z, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6B
 
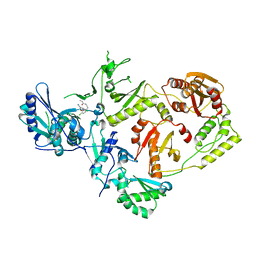 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with N-(4-chloro-3-(3-chloro-5-cyanophenoxy)phenethyl)acrylamide (JLJ731), a non-nucleoside inhibitor | | Descriptor: | N-{2-[4-chloro-3-(3-chloro-5-cyanophenoxy)phenyl]ethyl}prop-2-enamide, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Hollander, K, Henry, S, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6P
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with 3-(2-((2-cyanoindolizin-8-yl)oxy)phenoxy)-N,N-dimethylpropanamide (JLJ754), a non-nucleoside inhibitor | | Descriptor: | 3-(2-{[(4S)-2-cyanoindolizin-8-yl]oxy}phenoxy)-N,N-dimethylpropanamide, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Prucha, G, Henry, S, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6A
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with (JLJ729), a non-nucleoside inhibitor | | Descriptor: | N-(3-{2-[5-chloro-2-(3-chloro-5-cyanophenoxy)phenoxy]ethyl}phenyl)prop-2-enamide, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Hollander, K, Carter, Z, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6Q
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with 8-(2-(3-oxo-3-(pyrrolidin-1-yl)propoxy)phenoxy)indolizine-2-carbonitrile (JLJ755), a non-nucleoside inhibitor | | Descriptor: | (4S)-8-{2-[3-oxo-3-(pyrrolidin-1-yl)propoxy]phenoxy}indolizine-2-carbonitrile, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Prucha, G, Henry, S, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6O
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with5-(2-(3-oxo-3-(pyrrolidin-1-yl)propoxy)phenoxy)-2-naphthonitrile (JLJ753), a non-nucleoside inhibitor | | Descriptor: | 5-{2-[3-oxo-3-(pyrrolidin-1-yl)propoxy]phenoxy}naphthalene-2-carbonitrile, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Prucha, G, Henry, S, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U69
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with 3-chloro-5-(4-chloro-2-(2-(5-chloro-2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)ethoxy)phenoxy)benzonitrile (JLJ334), a non-nucleoside inhibitor | | Descriptor: | 3-chloro-5-{4-chloro-2-[2-(5-chloro-2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)ethoxy]phenoxy}benzonitrile, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Hollander, K, Carter, Z, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
8U6E
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with N-(4-chloro-3-(3-chloro-5-cyanophenoxy)phenethyl)-N-methylacrylamide (JLJ738), a non-nucleoside inhibitor | | Descriptor: | MAGNESIUM ION, N-{2-[4-chloro-3-(3-chloro-5-cyanophenoxy)phenyl]ethyl}-N-methylprop-2-enamide, Reverse transcriptase/ribonuclease H, ... | | Authors: | Prucha, G, Henry, S, Jorgensen, W.L, Anderson, K.S. | | Deposit date: | 2023-09-13 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Covalent and noncovalent strategies for targeting Lys102 in HIV-1 reverse transcriptase.
Eur.J.Med.Chem., 262, 2023
|
|
