8DJ0
 
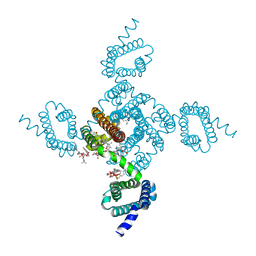 | | Crystal structure of NavAb L123T as a basis for the human Nav1.7 Inherited Erythromelalgia I848T mutation | | Descriptor: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, Ion transport protein | | Authors: | Wisedchaisri, G, Gamal El-Din, T.M, Zheng, N, Catterall, W.A. | | Deposit date: | 2022-06-29 | | Release date: | 2023-04-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis for severe pain caused by mutations in the S4-S5 linkers of voltage-gated sodium channel Na V 1.7.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
9D53
 
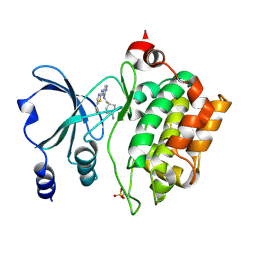 | | PAK4 in complex with compound 7 | | Descriptor: | N~2~-{[(1s,4s)-4-aminocyclohexyl]methyl}-N~4~-(5-cyclopropyl-1,3-thiazol-2-yl)pyrimidine-2,4-diamine, Serine/threonine-protein kinase PAK 4 | | Authors: | Boone, C.D, Olland, A.M, Suto, R.K. | | Deposit date: | 2024-08-13 | | Release date: | 2025-04-02 | | Last modified: | 2025-07-16 | | Method: | X-RAY DIFFRACTION (2.468 Å) | | Cite: | Identification of a p21-activated kinase 1 (PAK1) inhibitor with 10-fold selectivity against PAK2.
Bioorg.Med.Chem.Lett., 127, 2025
|
|
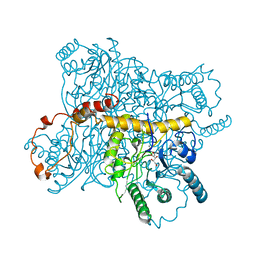
4XIS
 
 | |
6QVO
 
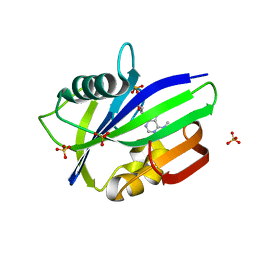 | | Crystal structure of human MTH1 in complex with N6-methyl-dAMP | | Descriptor: | 7,8-dihydro-8-oxoguanine triphosphatase, GLYCEROL, N6-METHYL-DEOXY-ADENOSINE-5'-MONOPHOSPHATE, ... | | Authors: | Scaletti, E, Vallin, K.S, Brautigam, L, Sarno, A, Warpman Berglund, U, Helleday, T, Stenmark, P, Jemth, A.S. | | Deposit date: | 2019-03-04 | | Release date: | 2020-03-18 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | MutT homologue 1 (MTH1) removes N6-methyl-dATP from the dNTP pool.
J.Biol.Chem., 295, 2020
|
|
8Z6H
 
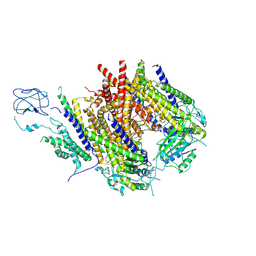 | | Structure of Polycystin-1/Polycystin-2 complex with Phosphatidylglycerol-bound | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, 2-acetamido-2-deoxy-beta-D-glucopyranose, Polycystin-1, ... | | Authors: | Chen, M.Y, Su, Q, Shi, Y.G. | | Deposit date: | 2024-04-19 | | Release date: | 2025-04-23 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structure of Polycystin-1/Polycystin-2 complex with Phosphatidylglycerol-bound
To Be Published
|
|
6YKK
 
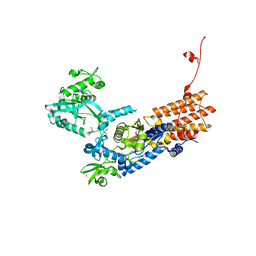 | | Neisseria gonorrhoeae Leucyl-tRNA Synthetase in Complex with Compound 15 | | Descriptor: | 1,2-ETHANEDIOL, Leucine--tRNA ligase, MAGNESIUM ION, ... | | Authors: | Pang, L, Strelkov, S.V, Weeks, S.D. | | Deposit date: | 2020-04-06 | | Release date: | 2020-12-02 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.236 Å) | | Cite: | Synthesis and structure-activity studies of novel anhydrohexitol-based Leucyl-tRNA synthetase inhibitors.
Eur.J.Med.Chem., 211, 2021
|
|
6WSJ
 
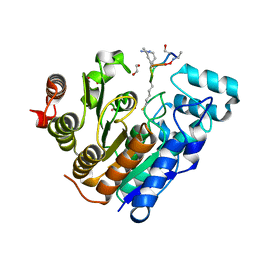 | |
6RB1
 
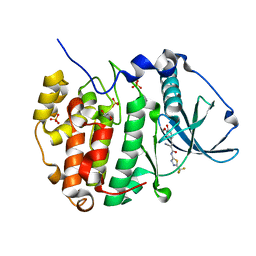 | | Human protein kinase CK2 alpha in complex with 2-cyano-2-propenamide compound 1 | | Descriptor: | (~{E})-2-cyano-3-(3-methoxy-4-oxidanyl-phenyl)-~{N}-[5-(trifluoromethyl)-1,3,4-thiadiazol-2-yl]prop-2-enamide, 1,2-ETHANEDIOL, Casein kinase II subunit alpha, ... | | Authors: | Battistutta, R, Lolli, G. | | Deposit date: | 2019-04-08 | | Release date: | 2020-04-08 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | A novel class of selective CK2 inhibitors targeting its open hinge conformation.
Eur.J.Med.Chem., 195, 2020
|
|
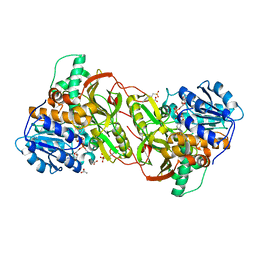
4XQC
 
 | |
8D51
 
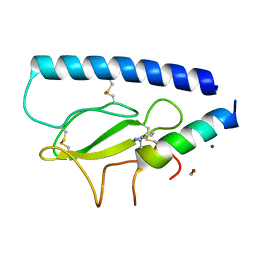 | | Parathyroid hormone 1 receptor extracellular domain complexed with a peptide ligand containing beta-3-homotryptophan | | Descriptor: | 1,2-ETHANEDIOL, PTHrP[1-36], Parathyroid hormone/parathyroid hormone-related peptide receptor, ... | | Authors: | Yu, Z, Kreitler, D.F, Gellman, S.H. | | Deposit date: | 2022-06-03 | | Release date: | 2023-06-14 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Harnessing Aromatic-Histidine Interactions through Synergistic Backbone Extension and Side Chain Modification.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8G92
 
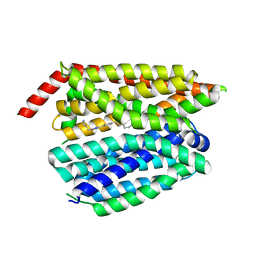 | | Structure of inhibitor 16d-bound SPNS2 | | Descriptor: | 3-[3-(4-decylphenyl)-1,2,4-oxadiazol-5-yl]propan-1-amine, Sphingosine-1-phosphate transporter SPNS2 | | Authors: | Chen, H, Li, X. | | Deposit date: | 2023-02-21 | | Release date: | 2023-05-24 | | Last modified: | 2023-12-13 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structural and functional insights into Spns2-mediated transport of sphingosine-1-phosphate.
Cell, 186, 2023
|
|
8ZKH
 
 | | Structure of Polycystin-1/Polycystin-2 complex with phosphatidylglycerol-bound | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Chen, M.Y, Su, Q, Shi, Y.G. | | Deposit date: | 2024-05-16 | | Release date: | 2025-05-21 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Structure of Polycystin-1/Polycystin-2 complex with phosphatidylglycerol-bound
To Be Published
|
|
5M53
 
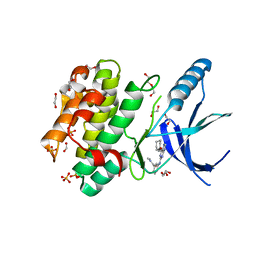 | | Nek2 bound to arylaminopurine inhibitor 11 | | Descriptor: | 1,2-ETHANEDIOL, 3-[[6-(cyclohexylmethoxy)-7~{H}-purin-2-yl]amino]-~{N},~{N}-dimethyl-benzamide, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Bayliss, R, Carr, K.H. | | Deposit date: | 2016-10-20 | | Release date: | 2016-11-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-guided design of purine-based probes for selective Nek2 inhibition.
Oncotarget, 8, 2017
|
|
8VJV
 
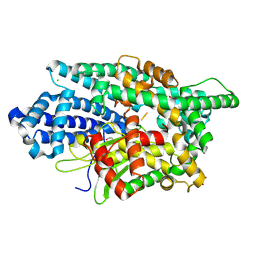 | |
6Z66
 
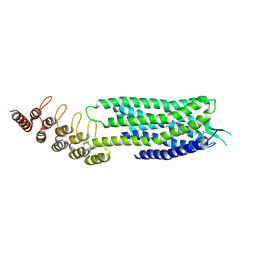 | | Crystal structure of apo-state neurotensin receptor 1 | | Descriptor: | Neurotensin receptor type 1,Neurotensin receptor type 1,DARPin | | Authors: | Deluigi, M, Klipp, A, Hilge, M, Merklinger, L, Klenk, C, Plueckthun, A. | | Deposit date: | 2020-05-27 | | Release date: | 2021-02-10 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3.192 Å) | | Cite: | Complexes of the neurotensin receptor 1 with small-molecule ligands reveal structural determinants of full, partial, and inverse agonism.
Sci Adv, 7, 2021
|
|
8RP3
 
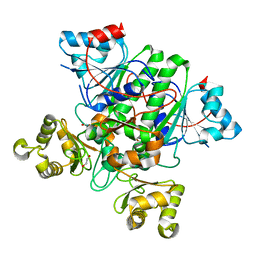 | |
9CM6
 
 | |
4XCZ
 
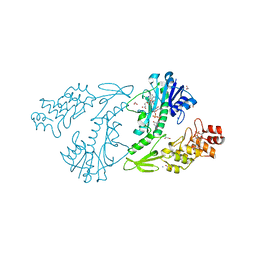 | | X-ray structure of the N-formyltransferase QdtF from Providencia alcalifaciens in complex with TDP-Qui3n and N5-THF | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, N-{[4-({[(6R)-2-amino-5-formyl-4-oxo-1,4,5,6,7,8-hexahydropteridin-6-yl]methyl}amino)phenyl]carbonyl}-L-glutamic acid, ... | | Authors: | Thoden, J.B, Woodford, C.R, Holden, H.M. | | Deposit date: | 2014-12-18 | | Release date: | 2015-01-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | New Role for the Ankyrin Repeat Revealed by a Study of the N-Formyltransferase from Providencia alcalifaciens.
Biochemistry, 54, 2015
|
|
6RC6
 
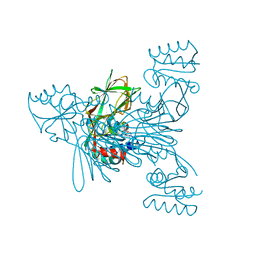 | |
8YXQ
 
 | |
5XXM
 
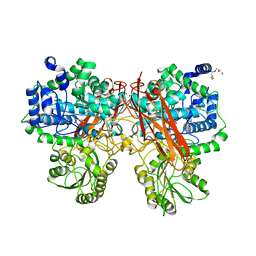 | | Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone | | Descriptor: | D-glucono-1,5-lactone, MAGNESIUM ION, Periplasmic beta-glucosidase, ... | | Authors: | Nakajima, M, Ishiguro, R, Tanaka, N, Abe, K, Maeda, T, Miyanaga, A, Takahash, Y, Sugimoto, N, Nakai, H, Taguchi, H. | | Deposit date: | 2017-07-04 | | Release date: | 2017-12-13 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Function and structure relationships of a beta-1,2-glucooligosaccharide-degrading beta-glucosidase.
FEBS Lett., 591, 2017
|
|
8RPC
 
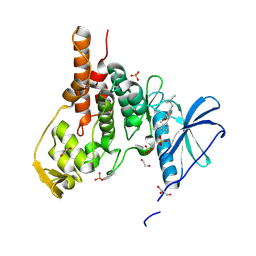 | | Crystal structure of PfCLK3 with TCMDC-135051 | | Descriptor: | 1,2-ETHANEDIOL, 4-[2-[5-(diethylaminomethyl)-2-methoxy-phenyl]-1~{H}-pyrrolo[2,3-b]pyridin-4-yl]-2-propan-2-yl-benzoic acid, GLYCEROL, ... | | Authors: | Yelland, T.S, Benazir, A, Hole, A. | | Deposit date: | 2024-01-15 | | Release date: | 2024-11-06 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (2.079 Å) | | Cite: | Targeting Pf CLK3 with Covalent Inhibitors: A Novel Strategy for Malaria Treatment.
J.Med.Chem., 67, 2024
|
|
9BA0
 
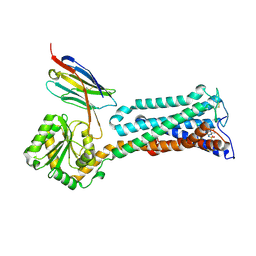 | | Structural mechanism of CB1R binding to peripheral and biased inverse agonists | | Descriptor: | CNb36, Cannabinoid receptor 1,Glycogen synthase, N-(N-{(E)-[(4S)-3-(4-chlorophenyl)-4-phenyl-4,5-dihydro-1H-pyrazol-1-yl][4-(trifluoromethyl)benzene-1-sulfonamido]methylidene}carbamimidoyl)acetamide | | Authors: | Kumari, P, Dvoracsko, S, Enos, M.D, Ramesh, K, Lim, D, Hassan, S.A, Kunos, G, Cinar, R, Iyer, M.R, Rosenbaum, D.M. | | Deposit date: | 2024-04-03 | | Release date: | 2024-11-20 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.13 Å) | | Cite: | Structural mechanism of CB 1 R binding to peripheral and biased inverse agonists.
Nat Commun, 15, 2024
|
|
6R5N
 
 | |
966C
 
 | | CRYSTAL STRUCTURE OF FIBROBLAST COLLAGENASE-1 COMPLEXED TO A DIPHENYL-ETHER SULPHONE BASED HYDROXAMIC ACID | | Descriptor: | CALCIUM ION, MMP-1, N-HYDROXY-2-[4-(4-PHENOXY-BENZENESULFONYL)-TETRAHYDRO-PYRAN-4-YL]-ACETAMIDE, ... | | Authors: | Lovejoy, B, Welch, A, Carr, S, Luong, C, Broka, C, Hendricks, R.T, Campbell, J, Walker, K, Martin, R, Van Wart, H, Browner, M.F. | | Deposit date: | 1998-08-07 | | Release date: | 1999-08-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structures of MMP-1 and -13 reveal the structural basis for selectivity of collagenase inhibitors.
Nat.Struct.Biol., 6, 1999
|
|
