8CH7
 
 | | RDC-refined Interleukin-4 (wild type) pH 5.6 | | 分子名称: | Interleukin-4 | | 著者 | Vaz, D.C, Rodrigues, J.R, Loureiro-Ferreira, N, Mueller, T, Sebald, W, Redfield, C, Brito, R.M.M. | | 登録日 | 2023-02-07 | | 公開日 | 2023-10-18 | | 最終更新日 | 2024-01-17 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Lessons on protein structure from interleukin-4: All disulfides are not created equal.
Proteins, 92, 2024
|
|
8CGF
 
 | | Interleukin-4 (wild type) pH 2.4 | | 分子名称: | Interleukin-4 | | 著者 | Vaz, D.C, Rodrigues, J.R, Loureiro-Ferreira, N, Mueller, T, Sebald, W, Redfield, C, Brito, R.M.M. | | 登録日 | 2023-02-04 | | 公開日 | 2023-10-18 | | 最終更新日 | 2024-01-17 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Lessons on protein structure from interleukin-4: All disulfides are not created equal.
Proteins, 92, 2024
|
|
5M8I
 
 | |
5GJJ
 | |
5GO0
 | |
5GWM
 
 | |
5GPH
 
 | |
6DST
 
 | |
1OO3
 
 | |
7RWR
 
 | | An RNA aptamer that decreases flavin redox potential | | 分子名称: | FLAVIN MONONUCLEOTIDE, RNA (38-MER) | | 著者 | Gremminger, T, Li, J, Chen, S, Heng, X. | | 登録日 | 2021-08-20 | | 公開日 | 2022-07-20 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | An RNA aptamer that shifts the reduction potential of metabolic cofactors.
Nat.Chem.Biol., 18, 2022
|
|
1N66
 
 | | Structure of the pyrimidine-rich internal loop in the Y-domain of poliovirus 3'UTR | | 分子名称: | internal loop in the Y-domain of poliovirus 3'UTR | | 著者 | Lescrinier, E.M, Tessari, M, van Kuppeveld, F.J, Melchers, W.J, Hilbers, C.W, Heus, H.A. | | 登録日 | 2002-11-08 | | 公開日 | 2003-08-19 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure of the Pyrimidine-rich Internal Loop in the Poliovirus 3'-UTR: The Importance of Maintaining Pseudo-2-fold Symmetry in RNA Helices Containing Two Adjacent Non-canonical Base-pairs.
J.Mol.Biol., 331, 2003
|
|
1OO4
 
 | |
6EWV
 
 | | Solution Structure of Docking Domain Complex of RXP NRPS: Kj12C NDD - Kj12B CDD | | 分子名称: | NRPS Kj12C-NDD, NRPS Kj12B-CDD | | 著者 | Hacker, C, Cai, X, Kegler, C, Zhao, L, Weickhmann, A.K, Bode, H.B, Woehnert, J. | | 登録日 | 2017-11-06 | | 公開日 | 2018-10-31 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure-based redesign of docking domain interactions modulates the product spectrum of a rhabdopeptide-synthesizing NRPS.
Nat Commun, 9, 2018
|
|
6F46
 
 | |
6EWU
 
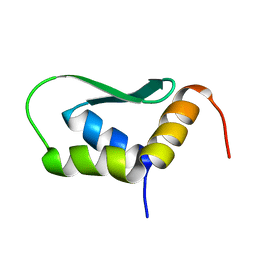 | | Solution Structure of Rhabdopeptide NRPS Docking Domain Kj12C-NDD | | 分子名称: | NRPS Kj12C-NDD | | 著者 | Hacker, C, Cai, X, Kegler, C, Zhao, L, Weickhmann, A.K, Bode, H.B, Woehnert, J. | | 登録日 | 2017-11-06 | | 公開日 | 2018-10-31 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLID-STATE NMR | | 主引用文献 | Structure-based redesign of docking domain interactions modulates the product spectrum of a rhabdopeptide-synthesizing NRPS.
Nat Commun, 9, 2018
|
|
6G4A
 
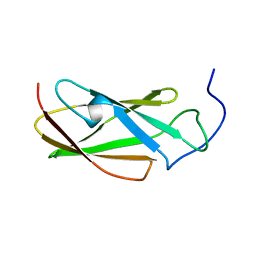 | | FLN5 (full length) | | 分子名称: | Gelation factor | | 著者 | Waudby, C.A, Wlodarski, T, Karyadi, M.-E, Cassaignau, A.M.E, Chan, S.H.S, Wentink, A.S, Schmidt-Engler, J.M, Camilloni, C, Vendruscolo, M, Cabrita, L.D, Christodoulou, J. | | 登録日 | 2018-03-27 | | 公開日 | 2019-04-10 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Mapping energy landscapes of a growing filamin domain reveals an intermediate associated with proline isomerization during biosynthesis
To Be Published
|
|
7U67
 
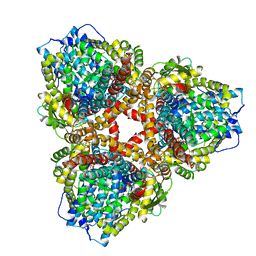 | | Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and GTP | | 分子名称: | Deoxyguanosinetriphosphate triphosphohydrolase, GUANOSINE-5'-TRIPHOSPHATE, Inhibitor of dGTPase, ... | | 著者 | Klemm, B.P, Hsu, A.L, Borgnia, M.J, Schaaper, R.M. | | 登録日 | 2022-03-03 | | 公開日 | 2022-08-31 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (2.5 Å) | | 主引用文献 | Mechanism by which T7 bacteriophage protein Gp1.2 inhibits Escherichia coli dGTPase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7U66
 
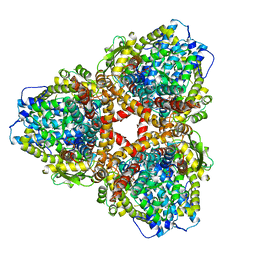 | | Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and dGTP | | 分子名称: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, Deoxyguanosinetriphosphate triphosphohydrolase, Inhibitor of dGTPase, ... | | 著者 | Klemm, B.P, Dillard, L.B, Borgnia, M.J, Schaaper, R.M. | | 登録日 | 2022-03-03 | | 公開日 | 2022-08-31 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Mechanism by which T7 bacteriophage protein Gp1.2 inhibits Escherichia coli dGTPase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
6GD5
 
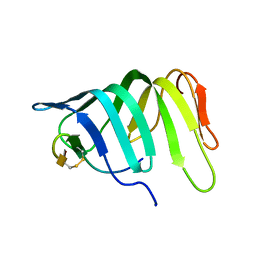 | |
1OSX
 | | Solution Structure of the Extracellular Domain of BLyS Receptor 3 (BR3) | | 分子名称: | Tumor necrosis factor receptor superfamily member 13C | | 著者 | Gordon, N.C, Pan, B, Hymowitz, S.G, Yin, J.P, Kelley, R.F, Cochran, A.G, Yan, M, Dixit, V.M, Fairbrother, W.J, Starovasnik, M.A. | | 登録日 | 2003-03-20 | | 公開日 | 2003-05-27 | | 最終更新日 | 2022-02-23 | | 実験手法 | SOLUTION NMR | | 主引用文献 | BAFF/BLyS receptor 3 comprises a minimal TNF receptor-like module that encodes a highly focused ligand-binding site
Biochemistry, 42, 2003
|
|
1P7M
 
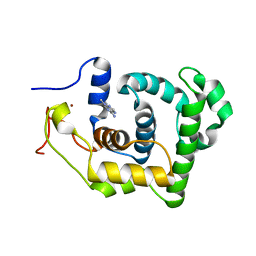 | | SOLUTION STRUCTURE AND BASE PERTURBATION STUDIES REVEAL A NOVEL MODE OF ALKYLATED BASE RECOGNITION BY 3-METHYLADENINE DNA GLYCOSYLASE I | | 分子名称: | 3-METHYL-3H-PURIN-6-YLAMINE, DNA-3-methyladenine glycosylase I, ZINC ION | | 著者 | Cao, C, Kwon, K, Jiang, Y.L, Drohat, A.C, Stivers, J.T. | | 登録日 | 2003-05-02 | | 公開日 | 2003-11-25 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure and base perturbation studies reveal a novel mode of alkylated base recognition by 3-methyladenine DNA glycosylase I
J.Biol.Chem., 278, 2003
|
|
1MYU
 | |
7X5C
 
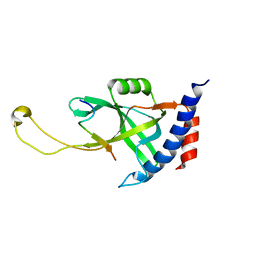 | | Solution structure of Tetrahymena p75OB1-p50PBM | | 分子名称: | Telomerase associated protein p50PBM, Telomerase-associated protein p75OB1 | | 著者 | Wu, B, Tang, T, Xue, H.J, Wu, J, Lei, M. | | 登録日 | 2022-03-04 | | 公開日 | 2022-10-19 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Association of the CST complex and p50 in Tetrahymena is crucial for telomere maintenance
Structure, 2022
|
|
1QBH
 
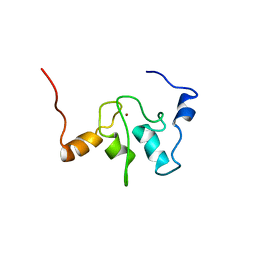 | | SOLUTION STRUCTURE OF A BACULOVIRAL INHIBITOR OF APOPTOSIS (IAP) REPEAT | | 分子名称: | INHIBITOR OF APOPTOSIS PROTEIN (2MIHB/C-IAP-1), ZINC ION | | 著者 | Hinds, M.G, Norton, R.S, Vaux, D.L, Day, C.L. | | 登録日 | 1999-04-20 | | 公開日 | 1999-10-20 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of a baculoviral inhibitor of apoptosis (IAP) repeat.
Nat.Struct.Biol., 6, 1999
|
|
5ZKV
 
 | | Solution structure of molten globule state of L94G mutant of horse cytochrome-c | | 分子名称: | Cytochrome c, HEME C | | 著者 | Naiyer, A, Islam, A, Hassan, M.I, Sundd, M, Ahmad, F. | | 登録日 | 2018-03-26 | | 公開日 | 2019-05-22 | | 最終更新日 | 2023-06-14 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of molten globule state of L94G mutant of horse cytochrome-c
To Be Published
|
|
