5MNW
 
 | |
5MTI
 
 | | Bamb_5917 Acyl-Carrier Protein | | Descriptor: | Phosphopantetheine-binding protein | | Authors: | Gallo, A, Kosol, S, Griffiths, D, Masschelein, J, Alkhalaf, L, Smith, H, Valentic, T, Tsai, S, Challis, G, Lewandowski, J.R. | | Deposit date: | 2017-01-09 | | Release date: | 2018-08-01 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structural basis for chain release from the enacyloxin polyketide synthase
Nat.Chem., 2019
|
|
5YOD
 
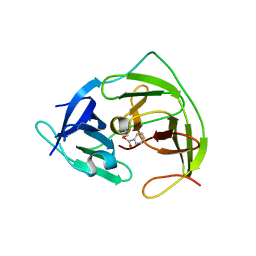 | |
5ZEW
 | |
5LVY
 | |
5MWQ
 
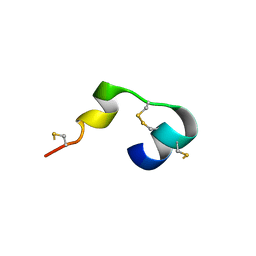 | |
5N8M
 
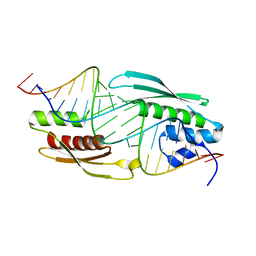 | |
3AME
 
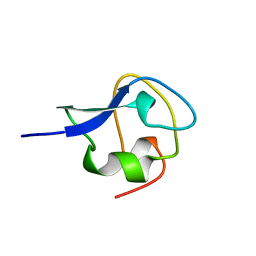 | | TYPE III ANTIFREEZE PROTEIN ISOFORM HPLC 12 Q9TQ44T | | Descriptor: | PROTEIN (ANTIFREEZE PROTEIN TYPE III) | | Authors: | Graether, S.P, Deluca, C.I, Baardsnes, J, Hill, G.A, Davies, P.L, Jia, Z. | | Deposit date: | 1999-01-24 | | Release date: | 1999-04-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Quantitative and qualitative analysis of type III antifreeze protein structure and function.
J.Biol.Chem., 274, 1999
|
|
3AIH
 
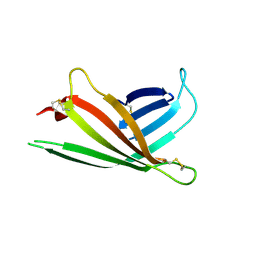 | | Human OS-9 MRH domain complexed with alpha3,alpha6-Man5 | | Descriptor: | Protein OS-9, alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)-beta-D-mannopyranose | | Authors: | Satoh, T, Chen, Y, Hu, D, Hanashima, S, Yamamoto, K, Yamaguchi, Y. | | Deposit date: | 2010-05-14 | | Release date: | 2010-12-22 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Basis for Oligosaccharide Recognition of Misfolded Glycoproteins by OS-9 in ER-Associated Degradation
Mol.Cell, 40, 2010
|
|
5N8L
 
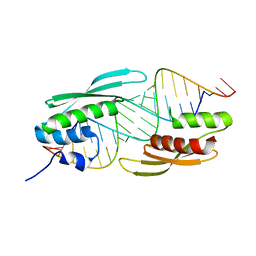 | |
5M9Z
 
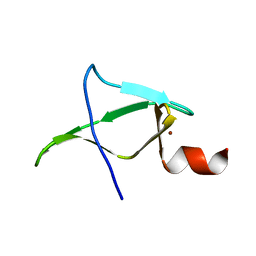 | | Second zinc-binding domain from yeast Pcf11 | | Descriptor: | Protein PCF11, ZINC ION | | Authors: | Mackereth, C. | | Deposit date: | 2016-11-02 | | Release date: | 2017-08-02 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Distinct roles of Pcf11 zinc-binding domains in pre-mRNA 3'-end processing.
Nucleic Acids Res., 45, 2017
|
|
1RNK
 | |
5M0A
 | |
5WPY
 
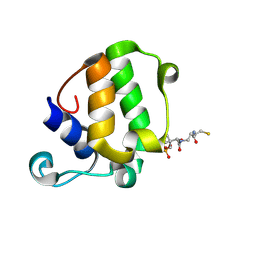 | |
5Z1Y
 
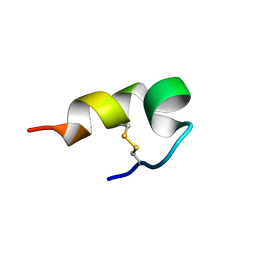 | | mBjAMP1 structure | | Descriptor: | mBjAMP1 peptide | | Authors: | Nam, J.Y, Lee, C.W. | | Deposit date: | 2017-12-28 | | Release date: | 2019-01-02 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | mBjAMP1 structure
To Be Published
|
|
5M1S
 
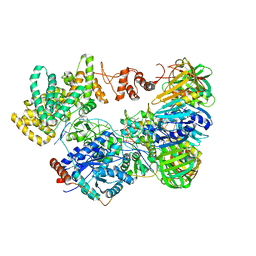 | | Cryo-EM structure of the E. coli replicative DNA polymerase-clamp-exonuclase-theta complex bound to DNA in the editing mode | | Descriptor: | DNA Primer Strand, DNA Template Strand, DNA polymerase III subunit alpha, ... | | Authors: | Fernandez-Leiro, R, Conrad, J, Scheres, S.H.W, Lamers, M.H. | | Deposit date: | 2016-10-10 | | Release date: | 2017-01-18 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (6.7 Å) | | Cite: | Self-correcting mismatches during high-fidelity DNA replication.
Nat. Struct. Mol. Biol., 24, 2017
|
|
5NQH
 
 | | Structure of the human Fe65-PTB2 homodimer | | Descriptor: | Amyloid beta A4 precursor protein-binding family B member 1, GLYCEROL, SULFATE ION | | Authors: | Feilen, L.P, Haubrich, K, Sinning, I, Konietzko, U, Kins, S, Simon, B, Wild, K. | | Deposit date: | 2017-04-20 | | Release date: | 2017-05-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Fe65-PTB2 Dimerization Mimics Fe65-APP Interaction.
Front Mol Neurosci, 10, 2017
|
|
5NCA
 
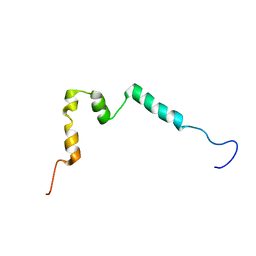 | | Solution structure of ComGC from Streptococcus pneumoniae | | Descriptor: | Competence protein ComGC | | Authors: | Erlendsson, S, Schmeider, P, Lichtenberg, C, Teilum, K, Akbey, U. | | Deposit date: | 2017-03-03 | | Release date: | 2017-07-05 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structure of the competence pilus major pilin ComGC in Streptococcus pneumoniae.
J. Biol. Chem., 292, 2017
|
|
5NBB
 
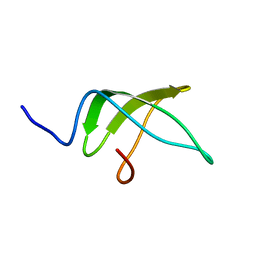 | | Structure of the C-terminal domain of the Escherichia Coli ProQ RNA binding protein | | Descriptor: | RNA chaperone ProQ | | Authors: | Gonzales, G, Hardwick, S, Maslen, S, Skehel, M, Holmqvist, E, Vogel, J, Bateman, A, Luisi, B, Broadhurst, R. | | Deposit date: | 2017-03-01 | | Release date: | 2017-05-03 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structure of the Escherichia coli ProQ RNA-binding protein.
RNA, 23, 2017
|
|
5NIQ
 
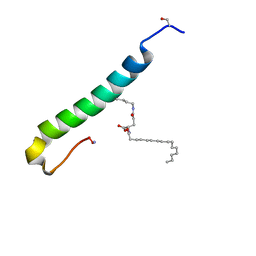 | |
5N2O
 
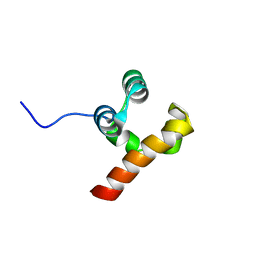 | | Structure Of P63 SAM Domain L514F Mutant Causative Of AEC Syndrome | | Descriptor: | Tumor protein 63 | | Authors: | Rinnenthal, J, Wuerz, J.M, Osterburg, C, Guentert, P, Doetsch, V. | | Deposit date: | 2017-02-08 | | Release date: | 2018-02-07 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Protein aggregation of the p63 transcription factor underlies severe skin fragility in AEC syndrome.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
5ZUX
 
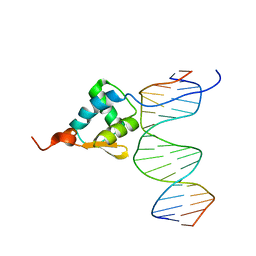 | |
6A4C
 
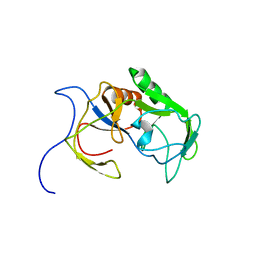 | | Solution structure of MXAN_0049 | | Descriptor: | Uncharacterized protein MXAN_0049 | | Authors: | Chen, C, Feng, Y. | | Deposit date: | 2018-06-19 | | Release date: | 2019-06-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of MXAN_0049
To Be Published
|
|
5WUZ
 
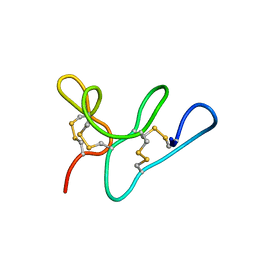 | | Morintides mO1 | | Descriptor: | Morintide mO1 | | Authors: | Xiao, T, Tam, J.P. | | Deposit date: | 2016-12-21 | | Release date: | 2017-01-25 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Morintides: 8C-Hevein-like peptides without a protein cargo from the drumstick tree Moringa oleifera
To Be Published
|
|
5WYO
 
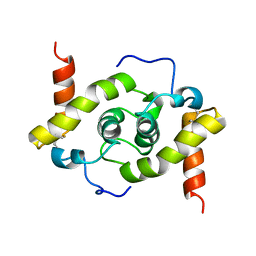 | | Solution structure of E.coli HdeA | | Descriptor: | Acid stress chaperone HdeA | | Authors: | Yang, C, Hu, Y, Jin, C. | | Deposit date: | 2017-01-14 | | Release date: | 2017-11-22 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Characterizations of the Interactions between Escherichia coli Periplasmic Chaperone HdeA and Its Native Substrates during Acid Stress
Biochemistry, 56, 2017
|
|
