5D0G
 
 | |
5D0H
 
 | |
5D0R
 
 | | Crystal structure of human soluble Adenylyl Cyclase with the inhibitor bithionol | | Descriptor: | 1,2-ETHANEDIOL, 2,2'-sulfanediylbis(4,6-dichlorophenol), ACETATE ION, ... | | Authors: | Kleinboelting, S, Steegborn, C. | | Deposit date: | 2015-08-03 | | Release date: | 2016-03-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Bithionol Potently Inhibits Human Soluble Adenylyl Cyclase through Binding to the Allosteric Activator Site.
J.Biol.Chem., 291, 2016
|
|
5D15
 
 | | Crystal structure of an adenylyl cyclase Ma1120 from Mycobacterium avium in complex with ATP and calcium ion | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Bharambe, N.G, Barathy, D.V, Suguna, K. | | Deposit date: | 2015-08-03 | | Release date: | 2016-08-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Substrate specificity determinants of class III nucleotidyl cyclases
Febs J., 283, 2016
|
|
5IV3
 
 | | Crystal structure of human soluble adenylyl cyclase in complex with alpha,beta-methyleneadenosine-5'-triphosphate and the allosteric inhibitor LRE1 | | Descriptor: | 1,2-ETHANEDIOL, 6-chloro-N~4~-cyclopropyl-N~4~-[(thiophen-2-yl)methyl]pyrimidine-2,4-diamine, ACETATE ION, ... | | Authors: | Kleinboelting, S, Steegborn, C. | | Deposit date: | 2016-03-18 | | Release date: | 2016-08-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Discovery of LRE1 as a specific and allosteric inhibitor of soluble adenylyl cyclase.
Nat.Chem.Biol., 12, 2016
|
|
5IV4
 
 | | Crystal structure of the human soluble adenylyl cyclase in complex with the allosteric inhibitor LRE1 | | Descriptor: | 1,2-ETHANEDIOL, 6-chloro-N~4~-cyclopropyl-N~4~-[(thiophen-2-yl)methyl]pyrimidine-2,4-diamine, ACETATE ION, ... | | Authors: | Kleinboelting, S, Steegborn, C. | | Deposit date: | 2016-03-18 | | Release date: | 2016-08-17 | | Last modified: | 2022-11-30 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Discovery of LRE1 as a specific and allosteric inhibitor of soluble adenylyl cyclase.
Nat.Chem.Biol., 12, 2016
|
|
5O5K
 
 | | X-ray structure of a bacterial adenylyl cyclase soluble domain | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, Adenylate cyclase, MANGANESE (II) ION, ... | | Authors: | Vercellino, I, Korkhov, V.M. | | Deposit date: | 2017-06-02 | | Release date: | 2017-11-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Role of the nucleotidyl cyclase helical domain in catalytically active dimer formation.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5O5L
 | |
5OYH
 
 | | crystal structure of the catalytic core of a rhodopsin-guanylyl cyclase with converted specificity in complex with ATPalphaS | | Descriptor: | ADENOSINE-5'-SP-ALPHA-THIO-TRIPHOSPHATE, CALCIUM ION, GLYCEROL, ... | | Authors: | Broser, M, Scheib, U, Hegemann, P. | | Deposit date: | 2017-09-09 | | Release date: | 2018-06-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.249 Å) | | Cite: | Rhodopsin-cyclases for photocontrol of cGMP/cAMP and 2.3 angstrom structure of the adenylyl cyclase domain.
Nat Commun, 9, 2018
|
|
5X4T
 
 | |
5X4U
 
 | |
5X4V
 
 | | Roseoflavin substituted OaPAC | | Descriptor: | 1-deoxy-1-[8-(dimethylamino)-7-methyl-2,4-dioxo-3,4-dihydrobenzo[g]pteridin-10(2H)-yl]-D-ribitol, Photoactivated adenylyl cyclase | | Authors: | Ohki, M, Park, S.-Y. | | Deposit date: | 2017-02-14 | | Release date: | 2017-07-26 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular mechanism of photoactivation of a light-regulated adenylate cyclase.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
6AO9
 
 | |
6AOA
 
 | |
6AOB
 
 | |
6FHT
 
 | | Crystal structure of an artificial phytochrome regulated adenylate/guanylate cyclase in its dark adapted Pr form | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome,Adenylate cyclase, DODECAETHYLENE GLYCOL, ... | | Authors: | Etzl, S, Winkler, A. | | Deposit date: | 2018-01-15 | | Release date: | 2018-04-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure-guided design and functional characterization of an artificial red light-regulated guanylate/adenylate cyclase for optogenetic applications.
J. Biol. Chem., 293, 2018
|
|
6JT0
 
 | | Structure of human soluble guanylate cyclase in the unliganded state | | Descriptor: | Guanylate cyclase soluble subunit alpha-1, Guanylate cyclase soluble subunit beta-1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Chen, L, Kang, Y, Liu, R, Wu, J.-X. | | Deposit date: | 2019-04-08 | | Release date: | 2019-08-28 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Structural insights into the mechanism of human soluble guanylate cyclase.
Nature, 574, 2019
|
|
6JT1
 
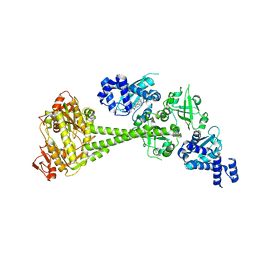 | | Structure of human soluble guanylate cyclase in the heme oxidised state | | Descriptor: | Guanylate cyclase soluble subunit alpha-1, Guanylate cyclase soluble subunit beta-1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Chen, L, Kang, Y, Liu, R, Wu, J.-X. | | Deposit date: | 2019-04-08 | | Release date: | 2019-08-28 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Structural insights into the mechanism of human soluble guanylate cyclase.
Nature, 574, 2019
|
|
6JT2
 
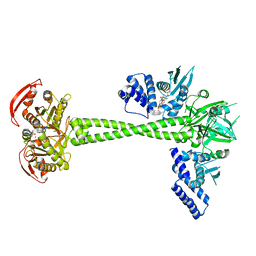 | | Structure of human soluble guanylate cyclase in the NO activated state | | Descriptor: | Guanylate cyclase soluble subunit alpha-1, Guanylate cyclase soluble subunit beta-1, MAGNESIUM ION, ... | | Authors: | Chen, L, Kang, Y, Liu, R, Wu, J.-X. | | Deposit date: | 2019-04-08 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural insights into the mechanism of human soluble guanylate cyclase.
Nature, 574, 2019
|
|
6PAS
 
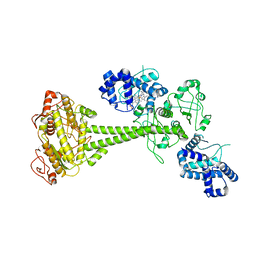 | | Inactive State of Manduca sexta soluble guanylate cyclase | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, Soluble guanylyl cyclase alpha-1 subunit, Soluble guanylyl cyclase beta-1 subunit | | Authors: | Yokom, A.L, Horst, B.G, Marletta, M.A, Hurley, J.H. | | Deposit date: | 2019-06-11 | | Release date: | 2019-10-23 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (5.1 Å) | | Cite: | Allosteric activation of the nitric oxide receptor soluble guanylate cyclase mapped by cryo-electron microscopy.
Elife, 8, 2019
|
|
6PAT
 
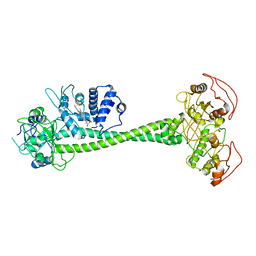 | | Active State of Manduca sexta soluble Guanylate Cyclase | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, Soluble guanylyl cyclase alpha-1 subunit, Soluble guanylyl cyclase beta-1 subunit | | Authors: | Yokom, A.L, Horst, B.G, Marletta, M.A, Hurley, J.H. | | Deposit date: | 2019-06-11 | | Release date: | 2019-10-23 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (5.8 Å) | | Cite: | Allosteric activation of the nitric oxide receptor soluble guanylate cyclase mapped by cryo-electron microscopy.
Elife, 8, 2019
|
|
6R3Q
 
 | |
6R4O
 
 | | Structure of a truncated adenylyl cyclase bound to MANT-GTP, forskolin and an activated stimulatory Galphas protein | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Adenylate cyclase 9, ... | | Authors: | Qi, C, Sorrentino, S, Medalia, O, Korkhov, V.M. | | Deposit date: | 2019-03-22 | | Release date: | 2019-05-08 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | The structure of a membrane adenylyl cyclase bound to an activated stimulatory G protein.
Science, 364, 2019
|
|
6R4P
 
 | |
6SIR
 
 | |
