3TM1
 
 | |
3R0O
 
 | |
3RQI
 
 | |
3RSI
 
 | |
3SES
 
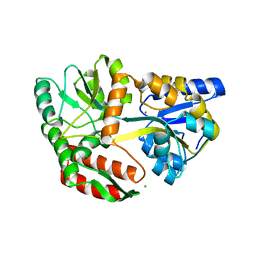 | | Cu-mediated Dimer of Maltose-binding Protein A216H/K220H by Synthetic Symmetrization | | Descriptor: | CHLORIDE ION, COPPER (II) ION, Maltose-binding periplasmic protein, ... | | Authors: | Zhao, M, Soriaga, A.B, Laganowsky, A, Sawaya, M.R, Cascio, D, Yeates, T.O. | | Deposit date: | 2011-06-11 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | An approach to crystallizing proteins by metal-mediated synthetic symmetrization.
Protein Sci., 20, 2011
|
|
3S7Q
 
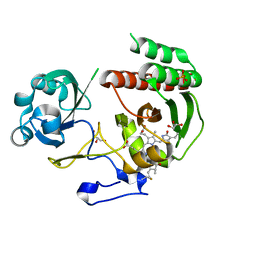 | | Crystal Structure of a Monomeric Infrared Fluorescent Deinococcus radiodurans Bacteriophytochrome chromophore binding domain | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, 3-[2-[(Z)-[5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-pyrrol-1-ium-2-ylidene]methyl]-5-[(Z)-[(3E,4R)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3-yl]propanoic acid, Bacteriophytochrome, ... | | Authors: | Auldridge, M.E, Satyshur, K.A, Forest, K.T. | | Deposit date: | 2011-05-26 | | Release date: | 2012-01-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.748 Å) | | Cite: | Structure-guided engineering enhances a phytochrome-based infrared fluorescent protein.
J.Biol.Chem., 287, 2012
|
|
3SB5
 
 | | Zn-mediated Trimer of T4 Lysozyme R125C/E128C by Synthetic Symmetrization | | Descriptor: | CHLORIDE ION, GLYCEROL, Lysozyme, ... | | Authors: | Laganowsky, A, Soriaga, A.B, Zhao, M, Sawaya, M.R, Cascio, D, Yeates, T.O. | | Deposit date: | 2011-06-03 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | An approach to crystallizing proteins by metal-mediated synthetic symmetrization.
Protein Sci., 20, 2011
|
|
3U8T
 
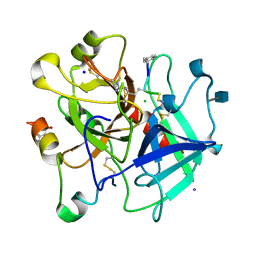 | | Human thrombin complexed with D-Phe-Pro-D-Arg-Cys | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Figueiredo, A.C, Clement, C.C, Philipp, M, Pereira, P.J.B. | | Deposit date: | 2011-10-17 | | Release date: | 2012-04-11 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.861 Å) | | Cite: | Rational design and characterization of d-phe-pro-d-arg-derived direct thrombin inhibitors.
Plos One, 7, 2012
|
|
3TU5
 
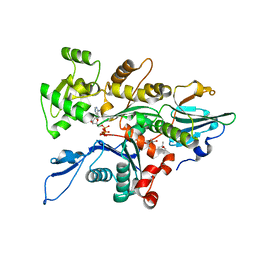 | | Actin complex with Gelsolin Segment 1 fused to Cobl segment | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, Actin, ... | | Authors: | Kudryashov, D.S, Sawaya, M.R, Durer, Z.A.O. | | Deposit date: | 2011-09-15 | | Release date: | 2012-09-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural States and dynamics of the d-loop in actin.
Biophys.J., 103, 2012
|
|
3T4U
 
 | |
3SVG
 
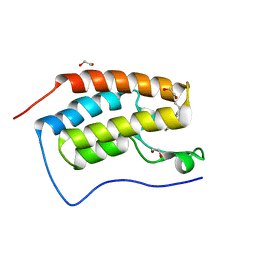 | | Crystal Structure of the first bromodomain of human BRD4 in complex with a 3,5-dimethylisoxazol ligand | | Descriptor: | (1R)-1-[3-(3,5-dimethyl-1,2-oxazol-4-yl)-5-ethoxyphenyl]ethanol, 1,2-ETHANEDIOL, Bromodomain-containing protein 4 | | Authors: | Filippakopoulos, P, Picaud, S, Felletar, I, Hewings, S.D, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Conway, S.J, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-07-12 | | Release date: | 2011-08-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Crystal Structure of the first bromodomain of human BRD4 in complex with a 3,5-dimethylisoxazol ligand
TO BE PUBLISHED
|
|
3SVT
 
 | |
3R9Q
 
 | |
3RR6
 
 | |
3RHD
 
 | | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase GapN from Methanocaldococcus jannaschii DSM 2661 complexed with NADP | | Descriptor: | Lactaldehyde dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Malashkevich, V.N, Toro, R, Seidel, R, Garrett, S, Foti, R, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase GapN from Methanocaldococcus jannaschii DSM 2661 complexed with NADP
To be Published
|
|
3U0Y
 
 | | Crystal structure of the Fucosylgalactoside alpha N-acetylgalactosaminyltransferase (GTA, cisAB mutant L266G, G268A) in complex with compound 382 and UDP | | Descriptor: | 1-(3-phenyl-1,2,4-thiadiazol-5-yl)piperazine, GLYCEROL, Histo-blood group ABO system transferase, ... | | Authors: | Palcic, M.M, Jorgensen, R. | | Deposit date: | 2011-09-29 | | Release date: | 2012-10-24 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A novel compound from a molecular fragment library screen inhibits glycosyltransferases by displacing the metal ion and interfering with substrate binding
To be Published
|
|
3R2N
 
 | |
3R9T
 
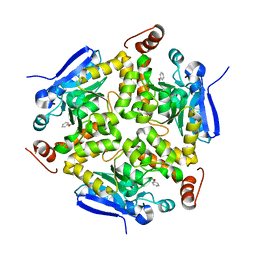 | |
3RCW
 
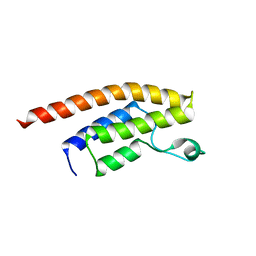 | | Crystal Structure of the bromodomain of human BRD1 | | Descriptor: | 1,2-ETHANEDIOL, 1-methylpyrrolidin-2-one, ACETATE ION, ... | | Authors: | Filippakopoulos, P, Keates, T, Picaud, S, Felletar, I, Pike, A.C.W, Krojer, T, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2011-03-31 | | Release date: | 2011-06-01 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Histone recognition and large-scale structural analysis of the human bromodomain family.
Cell(Cambridge,Mass.), 149, 2012
|
|
3RDE
 
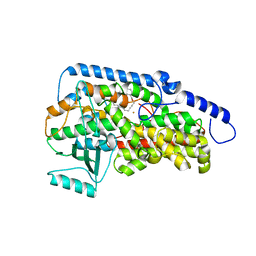 | | Crystal structure of the catalytic domain of porcine leukocyte 12-lipoxygenase | | Descriptor: | 3-{4-[(tridec-2-yn-1-yloxy)methyl]phenyl}propanoic acid, Arachidonate 12-lipoxygenase, 12S-type, ... | | Authors: | Funk, M.O, Xu, S, Marnett, L.J, Mueser, T.C. | | Deposit date: | 2011-04-01 | | Release date: | 2012-04-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.892 Å) | | Cite: | Crystal structure of 12-lipoxygenase catalytic-domain-inhibitor complex identifies a substrate-binding channel for catalysis.
Structure, 20, 2012
|
|
3RGQ
 
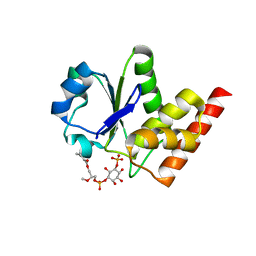 | | Crystal Structure of PTPMT1 in complex with PI(5)P | | Descriptor: | (2R)-3-{[(S)-hydroxy{[(1R,2R,3R,4R,5S,6R)-2,3,4,6-tetrahydroxy-5-(phosphonooxy)cyclohexyl]oxy}phosphoryl]oxy}propane-1,2-diyl dibutanoate, Protein-tyrosine phosphatase mitochondrial 1 | | Authors: | Xiao, J, Engel, J.L. | | Deposit date: | 2011-04-08 | | Release date: | 2011-07-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural and functional analysis of PTPMT1, a phosphatase required for cardiolipin synthesis.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3RFQ
 
 | |
3SWT
 
 | |
3T52
 
 | |
3SR1
 
 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS A69G bound to Ca2+ and thymidine-5',3'-diphosphate at cryogenic temperature | | Descriptor: | CALCIUM ION, THYMIDINE-3',5'-DIPHOSPHATE, thermonuclease | | Authors: | Doctrow, B.M, Schlessman, J.L, Garcia-Moreno E, B, Heroux, A. | | Deposit date: | 2011-07-06 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal structure of Staphylococcal nuclease variant Delta+PHS A69G bound to Ca2+ and thymidine-5',3'-diphosphate at cryogenic temperature
To be Published
|
|
