4Y7R
 
 | | Crystal structure of WDR5 in complex with MYC MbIIIb peptide | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MYC MbIIIb peptide, ... | | Authors: | Sun, Q, Phan, J, Olejniczak, E.T, Thomas, L.R, Fesik, S.W, Tansey, W.P. | | Deposit date: | 2015-02-16 | | Release date: | 2015-04-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Interaction with WDR5 Promotes Target Gene Recognition and Tumorigenesis by MYC.
Mol.Cell, 58, 2015
|
|
4U4Y
 
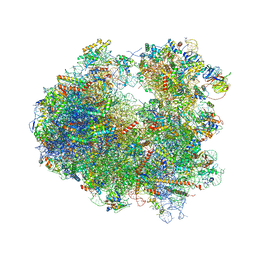 | | Crystal structure of Pactamycin bound to the yeast 80S ribosome | | Descriptor: | 18S ribosomal RNA, 25S ribosomal RNA, 40S ribosomal protein S0-A, ... | | Authors: | Garreau de Loubresse, N, Prokhorova, I, Yusupova, G, Yusupov, M. | | Deposit date: | 2014-07-24 | | Release date: | 2014-10-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural basis for the inhibition of the eukaryotic ribosome.
Nature, 513, 2014
|
|
1MV3
 | |
1MUZ
 
 | |
2VFT
 
 | |
2VFR
 
 | |
2VFU
 
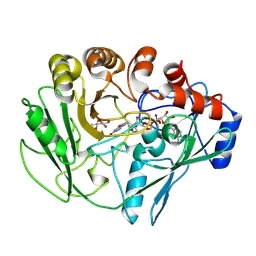 | |
2VFS
 
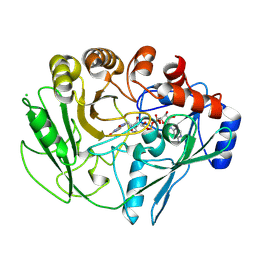 | |
2VFV
 
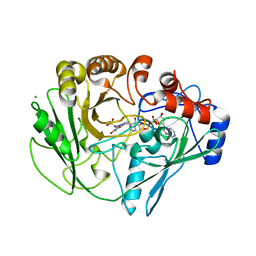 | | Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Sulphite | | Descriptor: | (S)-10-((2S,3S,4R)-5-((S)-((S)-(((2R,3S,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-3,4-DIHYDROXY-TETRAHYDROFURAN-2-YL)METHOXY)(HYDROXY)PHOSPHORYLOXY)(HYDROXY)PHOSPHORYLOXY)-2,3,4-TRIHYDROXYPENTYL)-7,8-DIMETHYL-2,4-DIOXO-2,3,4,4A-TETRAHYDROBENZO[G]PTERIDINE-5(10H)-SULFONIC ACID, CHLORIDE ION, XYLITOL OXIDASE | | Authors: | Forneris, F, Mattevi, A. | | Deposit date: | 2007-11-05 | | Release date: | 2008-01-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Structural Analysis of the Catalytic Mechanism and Stereoselectivity in Streptomyces Coelicolor Alditol Oxidase.
Biochemistry, 47, 2008
|
|
8WX0
 
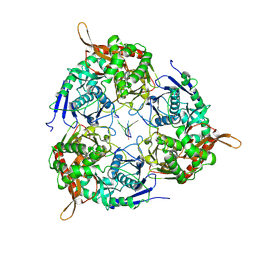 | | PNPase of M.tuberculosis with its RNA substrate | | Descriptor: | Bifunctional guanosine pentaphosphate synthetase/polyribonucleotide nucleotidyltransferase, RNA (24-mer) | | Authors: | Wang, N, Sheng, Y.N. | | Deposit date: | 2023-10-27 | | Release date: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Cryo-EM structures of Mycobacterium tuberculosis polynucleotide phosphorylase suggest a potential mechanism for its RNA substrate degradation.
Arch.Biochem.Biophys., 754, 2024
|
|
8TJG
 
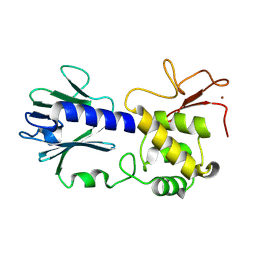 | |
8UFD
 
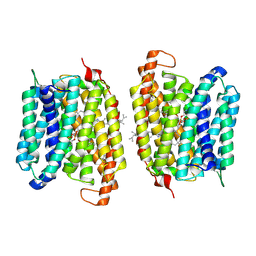 | | Multidrug efflux pump MtEfpA bound with inhibitor BRD8000.3 | | Descriptor: | (1S,3S)-N-[6-bromo-5-(pyrimidin-2-yl)pyridin-2-yl]-2,2-dimethyl-3-(2-methylpropyl)cyclopropane-1-carboxamide, Integral membrane efflux protein EFPA, PHOSPHATIDYLETHANOLAMINE | | Authors: | Wang, S, Liao, M. | | Deposit date: | 2023-10-04 | | Release date: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Structures of the Mycobacterium tuberculosis efflux pump EfpA reveal the mechanisms of transport and inhibition.
Nat Commun, 15, 2024
|
|
8T41
 
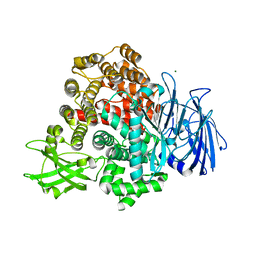 | |
8TA0
 | |
8TBR
 
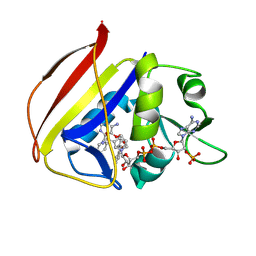 | |
8TA1
 
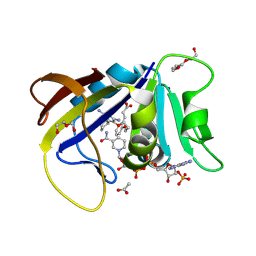 | |
8U12
 
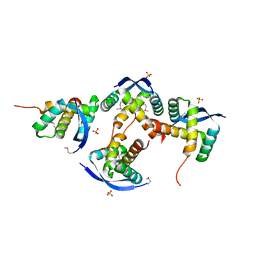 | |
8TXA
 
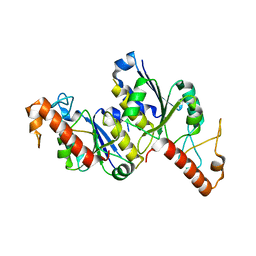 | | Apo Structure of (N1G37) tRNA Methyltransferase from Mycobacterium marinum | | Descriptor: | tRNA (guanine-N(1)-)-methyltransferase | | Authors: | Balsamo, A, Bruno, C, Edele, C, Fabian, T, Jannotta, R, Lee, R, Schryver, D, Warsaw, J, Warsaw, L, Stojanoff, V, Battaile, K, Perez, A, Bolen, R. | | Deposit date: | 2023-08-23 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.591 Å) | | Cite: | Apo Structure of (N1G37) tRNA Methyltransferase from Mycobacterium marinum
To Be Published
|
|
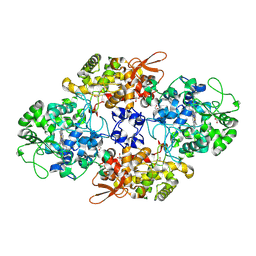
8U3P
 
 | |
7A10
 
 | | LppS with covalent adduct derived from 1g | | Descriptor: | 4-methoxycyclohexa-2,5-diene-1-thione, L,D-transpeptidase 2 | | Authors: | Schnell, R, Steiner, E.M. | | Deposit date: | 2020-08-11 | | Release date: | 2021-04-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | N-Thio-beta-lactams targeting L,D-transpeptidase-2, with activity against drug-resistant strains of Mycobacterium tuberculosis.
Cell Chem Biol, 28, 2021
|
|
7A1E
 
 | | LppS with covalent adduct derived from 1c | | Descriptor: | ACETATE ION, L,D-transpeptidase 2, phenylmethanethiol | | Authors: | Schnell, R, Steiner, E.M. | | Deposit date: | 2020-08-12 | | Release date: | 2021-04-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | N-Thio-beta-lactams targeting L,D-transpeptidase-2, with activity against drug-resistant strains of Mycobacterium tuberculosis.
Cell Chem Biol, 28, 2021
|
|
7A11
 
 | | LppS with covalent adduct derived from 1E | | Descriptor: | ACETATE ION, L,D-transpeptidase 2, propane-1-thiol | | Authors: | Schnell, R, Steiner, E.M. | | Deposit date: | 2020-08-11 | | Release date: | 2021-04-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | N-Thio-beta-lactams targeting L,D-transpeptidase-2, with activity against drug-resistant strains of Mycobacterium tuberculosis.
Cell Chem Biol, 28, 2021
|
|
7A0Z
 
 | | LppS with covalent adduct derived from 1b | | Descriptor: | L,D-transpeptidase 2, TRIS(HYDROXYETHYL)AMINOMETHANE, benzenethiol | | Authors: | Schnell, R, Steiner, E.M. | | Deposit date: | 2020-08-11 | | Release date: | 2021-04-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | N-Thio-beta-lactams targeting L,D-transpeptidase-2, with activity against drug-resistant strains of Mycobacterium tuberculosis.
Cell Chem Biol, 28, 2021
|
|
7A1C
 
 | |
8U2Q
 
 | |
