2AAR
 
 | | Structure of trigger factor binding domain in biologically homologous complex with eubacterial ribosome. | | Descriptor: | 23S ribosomal RNA, 50S ribosomal protein L23, 50S ribosomal protein L29, ... | | Authors: | Baram, D, Pyetan, E, Sittner, A, Auerbach-Nevo, T, Bashan, A, Yonath, A. | | Deposit date: | 2005-07-14 | | Release date: | 2005-08-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure of trigger factor binding domain in biologically homologous complex with eubacterial ribosome reveals its chaperone action
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
7ASN
 
 | |
7ASM
 
 | |
1XBP
 
 | | Inhibition of peptide bond formation by pleuromutilins: The structure of the 50S ribosomal subunit from Deinococcus radiodurans in complex with Tiamulin | | Descriptor: | 23S RIBOSOMAL RNA, 50S ribosomal protein L11, 50S ribosomal protein L13, ... | | Authors: | Schluenzen, F, Pyetan, E, Fucini, P, Yonath, A, Harms, J.M. | | Deposit date: | 2004-08-31 | | Release date: | 2005-03-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Inhibition of peptide bond formation by pleuromutilins: the structure of the 50S ribosomal subunit from Deinococcus radiodurans in complex with tiamulin.
Mol.Microbiol., 54, 2004
|
|
1Z58
 
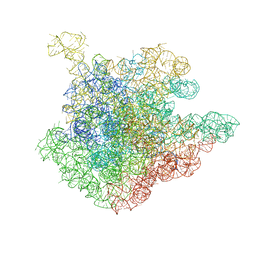 | | Crystal structure of a complex of the ribosome large subunit with rapamycin | | Descriptor: | 23S RIBOSOMAL RNA, RAPAMYCIN IMMUNOSUPPRESSANT DRUG | | Authors: | Amit, M, Berisio, R, Baram, D, Harms, J, Bashan, A, Yonath, A. | | Deposit date: | 2005-03-17 | | Release date: | 2005-06-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | A crevice adjoining the ribosome tunnel: Hints for cotranslational folding.
Febs Lett., 579, 2005
|
|
1SM1
 
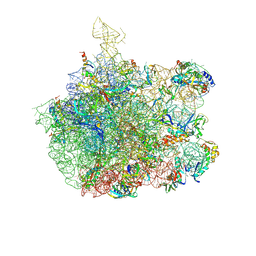 | | COMPLEX OF THE LARGE RIBOSOMAL SUBUNIT FROM DEINOCOCCUS RADIODURANS WITH QUINUPRISTIN AND DALFOPRISTIN | | Descriptor: | 23S RIBOSOMAL RNA, 5-(2-DIETHYLAMINO-ETHANESULFONYL)-21-HYDROXY-10-ISOPROPYL-11,19-DIMETHYL-9,26-DIOXA-3,15,28-TRIAZA-TRICYCLO[23.2.1.00,255]OCTACOSA-1(27),12,17,19,25(28)-PENTAENE-2,8,14,23-TETRAONE, 50S RIBOSOMAL PROTEIN L11, ... | | Authors: | Harms, J.M, Schluenzen, F, Fucini, P, Bartels, H, Yonath, A. | | Deposit date: | 2004-03-08 | | Release date: | 2004-08-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.42 Å) | | Cite: | Alterations at the Peptidyl Transferase Centre of the Ribosome Induced by the Synergistic Action of the Streptogramins Dalfopristin and Quinupristin.
Bmc Biol., 2, 2004
|
|
1CN1
 
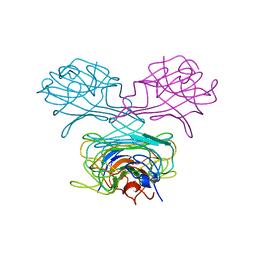 | | CRYSTAL STRUCTURE OF DEMETALLIZED CONCANAVALIN A. THE METAL-BINDING REGION | | Descriptor: | CONCANAVALIN A | | Authors: | Shoham, M, Yonath, A, Sussman, J.L, Moult, J, Traub, W, Gilboa(Kalb), A.J. | | Deposit date: | 1981-12-21 | | Release date: | 1982-03-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of demetallized concanavalin A: the metal-binding region.
J.Mol.Biol., 131, 1979
|
|
7A0R
 
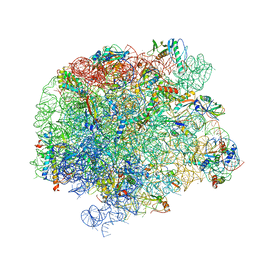 | | 50S Deinococcus radiodurans ribosome bounded with mycinamicin I | | Descriptor: | 50S ribosomal protein L13, 50S ribosomal protein L14, 50S ribosomal protein L15, ... | | Authors: | Breiner, E, Eyal, Z, Matzov, D, Halfon, Y, Cimicata, G, Rozenberg, H, Zimmerman, E, Bashan, A, Yonath, A. | | Deposit date: | 2020-08-10 | | Release date: | 2021-08-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Ribosome-binding and anti-microbial studies of the mycinamicins, 16-membered macrolide antibiotics from Micromonospora griseorubida.
Nucleic Acids Res., 49, 2021
|
|
1J5A
 
 | | STRUCTURAL BASIS FOR THE INTERACTION OF ANTIBIOTICS WITH THE PEPTIDYL TRANSFERASE CENTER IN EUBACTERIA | | Descriptor: | 23S RRNA, CLARITHROMYCIN, MAGNESIUM ION, ... | | Authors: | Schluenzen, F, Zarivach, R, Harms, J, Bashan, A, Tocilj, A, Albrecht, R, Yonath, A, Franceschi, F. | | Deposit date: | 2002-03-06 | | Release date: | 2002-03-08 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria.
Nature, 413, 2001
|
|
1FKA
 
 | | STRUCTURE OF FUNCTIONALLY ACTIVATED SMALL RIBOSOMAL SUBUNIT AT 3.3 A RESOLUTION | | Descriptor: | 16S RIBOSOMAL RNA, 30S RIBOSOMAL PROTEIN S10, 30S RIBOSOMAL PROTEIN S11, ... | | Authors: | Schluenzen, F, Tocilj, A, Zarivach, R, Harms, J, Gluehmann, M, Janell, D, Bashan, A, Bartels, H, Agmon, I, Franceschi, F, Yonath, A. | | Deposit date: | 2000-08-09 | | Release date: | 2000-09-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of functionally activated small ribosomal subunit at 3.3 angstroms resolution.
Cell(Cambridge,Mass.), 102, 2000
|
|
7A0S
 
 | | 50S Deinococcus radiodurans ribosome bounded with mycinamicin I | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 50S ribosomal protein L13, 50S ribosomal protein L14, ... | | Authors: | Breiner, E, Eyal, Z, Matzov, D, Halfon, Y, Cimicata, G, Rozenberg, H, Zimmerman, E, Bashan, A, Yonath, A. | | Deposit date: | 2020-08-10 | | Release date: | 2021-08-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.22 Å) | | Cite: | Ribosome-binding and anti-microbial studies of the mycinamicins, 16-membered macrolide antibiotics from Micromonospora griseorubida.
Nucleic Acids Res., 49, 2021
|
|
7A18
 
 | | 50S Deinococcus radiodurans ribosome bounded with mycinamicin IV | | Descriptor: | 50S ribosomal protein L13, 50S ribosomal protein L14, 50S ribosomal protein L15, ... | | Authors: | Breiner, E, Eyal, Z, Matzov, D, Halfon, Y, Cimicata, G, Rozenberg, H, Zimmerman, E, Bashan, A, Yonath, A. | | Deposit date: | 2020-08-12 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Ribosome-binding and anti-microbial studies of the mycinamicins, 16-membered macrolide antibiotics from Micromonospora griseorubida.
Nucleic Acids Res., 49, 2021
|
|
7ASP
 
 | |
7ASO
 
 | |
2O45
 
 | | Structure of the 23S rRNA of the large ribosomal subunit from Deinococcus Radiodurans in complex with the macrolide RU-69874 | | Descriptor: | (3AS,4R,7R,8S,9S,10R,11R,13R,15R,15AR)-4-ETHYL-11-METHOXY-3A,7,9,11,13,15-HEXAMETHYL-2,6,14-TRIOXO-1-[4-(4-PYRIDIN-3-YL-1H-IMIDAZOL-1-YL)BUTYL]-10-{[3,4,6-TRIDEOXY-3-(DIMETHYLAMINO)-BETA-D-XYLO-HEXOPYRANOSYL]OXY}TETRADECAHYDRO-2H-OXACYCLOTETRADECINO[4,3-D][1,3]OXAZOL-8-YL 2,6-DIDEOXY-3-C-METHYL-3-O-METHYL-ALPHA-L-RIBO-HEXOPYRANOSIDE, 23S rRNA | | Authors: | Baram, D, Pyetan, E, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Chemical Parameters Influencing Fine Tuning in the Binding of Macrolide Antibiotics to the Ribosomal Tunnel
TO BE PUBLISHED
|
|
2O43
 
 | | Structure of 23S rRNA of the large ribosomal subunit from Deinococcus radiodurans in complex with the macrolide erythromycylamine | | Descriptor: | (3R,4S,5S,6R,7R,9R,10S,11S,12R,13S,14R)-10-AMINO-6-{[(2S,3R,4S,6R)-4-(DIMETHYLAMINO)-3-HYDROXY-6-METHYLTETRAHYDRO-2H-PY RAN-2-YL]OXY}-14-ETHYL-7,12,13-TRIHYDROXY-4-{[(2R,4R,5S,6S)-5-HYDROXY-4-METHOXY-4,6-DIMETHYLTETRAHYDRO-2H-PYRAN-2-YL]OXY }-3,5,7,9,11,13-HEXAMETHYLOXACYCLOTETRADECAN-2-ONE, 23S rRNA | | Authors: | Pyetan, E, Baram, D, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Chemical parameters influencing fine tuning in the binding of Macrolide antibiotics to the ribosomal tunnel
TO BE PUBLISHED
|
|
2O44
 
 | | Structure of 23S rRNA of the large ribosomal subunit from Deinococcus radiodurans in complex with the macrolide josamycin | | Descriptor: | (2S,3S,4R,6S)-6-{[(2R,3S,4R,5R,6S)-6-{[(4R,5S,6S,7R,9R,10S,12E,14Z,16R)-4-(ACETYLOXY)-10-HYDROXY-5-METHOXY-9,16-DIMETHYL-2-OXO-7-(2-OXOETHYL)OXACYCLOHEXADECA-12,14-DIEN-6-YL]OXY}-4-(DIMETHYLAMINO)-5-HYDROXY-2-METHYLTETRAHYDRO-2H-PYRAN-3-YL]OXY}-4-HYDROXY-2,4-DIMETHYLTETRAHYDRO-2H-PYRAN-3-YL 3-METHYLBUTANOATE, 23S rRNA | | Authors: | Pyetan, E, Daram, D, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Chemical parameters influencing fine tuning in the binding of Macrolide antibiotics to the ribosomal tunnel
TO BE PUBLISHED
|
|
1NKW
 
 | | Crystal Structure Of The Large Ribosomal Subunit From Deinococcus Radiodurans | | Descriptor: | 23S ribosomal RNA, 50S ribosomal protein L11, 50S ribosomal protein L13, ... | | Authors: | Harms, J.M, Schluenzen, F, Zarivach, R, Bashan, A, Gat, S, Agmon, I, Bartels, H, Franceschi, F, Yonath, A. | | Deposit date: | 2003-01-05 | | Release date: | 2003-02-11 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | High resolution structure of the large ribosomal subunit from a mesophilic eubacterium
Cell(Cambridge,Mass.), 107, 2001
|
|
1OND
 
 | | THE CRYSTAL STRUCTURE OF THE 50S LARGE RIBOSOMAL SUBUNIT FROM DEINOCOCCUS RADIODURANS COMPLEXED WITH TROLEANDOMYCIN MACROLIDE ANTIBIOTIC | | Descriptor: | 23S RIBOSOMAL RNA, 50S ribosomal protein L22, 50S ribosomal protein L32, ... | | Authors: | Berisio, R, Schluenzen, F, Harms, J, Bashan, A, Auerbach, T, Baram, D, Yonath, A. | | Deposit date: | 2003-02-27 | | Release date: | 2003-04-15 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structural insight into the role of the ribosomal tunnel in cellular regulation
Nat.Struct.Biol., 10, 2003
|
|
2OGN
 
 | | The crystal structure of the large ribosomal subunit from Deinococcus radiodurans complexed with the pleuromutilin derivative SB-280080 | | Descriptor: | (3AS,4R,5S,6S,8R,9R,9AR,10R)-5-HYDROXY-4,6,9,10-TETRAMETHYL-1-OXO-6-VINYLDECAHYDRO-3A,9-PROPANOCYCLOPENTA[8]ANNULEN-8-YL (PIPERIDIN-4-YLTHIO)ACETATE, 23S ribosomal RNA, 50S ribosomal protein L3 | | Authors: | Davidovich, C, Bashan, A, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2007-01-07 | | Release date: | 2007-05-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.56 Å) | | Cite: | Induced-fit tightens pleuromutilins binding to ribosomes and remote interactions enable their selectivity.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2OGO
 
 | | The crystal structure of the large ribosomal subunit from Deinococcus radiodurans complexed with the pleuromutilin derivative retapamulin (SB-275833) | | Descriptor: | 23S ribosomal RNA, 50S ribosomal protein L3, Retapamulin | | Authors: | Davidovich, C, Bashan, A, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2007-01-07 | | Release date: | 2007-05-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.66 Å) | | Cite: | Induced-fit tightens pleuromutilins binding to ribosomes and remote interactions enable their selectivity.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2OGM
 
 | | The crystal structure of the large ribosomal subunit from Deinococcus radiodurans complexed with the pleuromutilin derivative SB-571519 | | Descriptor: | (2S,3AR,4R,5S,6S,8R,9R,9AR,10R)-2,5-DIHYDROXY-4,6,9,10-TETRAMETHYL-1-OXO-6-VINYLDECAHYDRO-3A,9-PROP[1]ENOCYCLOPENTA[8]ANNULEN-8-YL [(6-AMINOPYRIDAZIN-3-YL)CARBONYL]CARBAMATE, 23S ribosomal RNA, 50S ribosomal protein L3 | | Authors: | Davidovich, C, Bashan, A, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2007-01-07 | | Release date: | 2007-05-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Induced-fit tightens pleuromutilins binding to ribosomes and remote interactions enable their selectivity.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
1P9X
 
 | | THE CRYSTAL STRUCTURE OF THE 50S LARGE RIBOSOMAL SUBUNIT FROM DEINOCOCCUS RADIODURANS COMPLEXED WITH TELITHROMYCIN KETOLIDE ANTIBIOTIC | | Descriptor: | 23S RIBOSOMAL RNA, TELITHROMYCIN | | Authors: | Berisio, R, Harms, J, Schluenzen, F, Zarivach, R, Hansen, H.A, Fucini, P, Yonath, A. | | Deposit date: | 2003-05-13 | | Release date: | 2003-08-05 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structural insight into the antibiotic action of telithromycin against resistant mutants
J.Bacteriol., 185, 2003
|
|
1JZY
 
 | | Structural Basis for the Interaction of Antibiotics with the Peptidyl Transferase Center in Eubacteria | | Descriptor: | 23S rRNA, ERYTHROMYCIN A, MAGNESIUM ION, ... | | Authors: | Schluenzen, F, Zarivach, R, Harms, J, Bashan, A, Tocilj, A, Albrecht, R, Yonath, A, Franceschi, F. | | Deposit date: | 2001-09-17 | | Release date: | 2001-10-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria.
Nature, 413, 2001
|
|
1JZX
 
 | | Structural Basis for the Interaction of Antibiotics with the Peptidyl Transferase Center in Eubacteria | | Descriptor: | 23S rRNA, CLINDAMYCIN, MAGNESIUM ION, ... | | Authors: | Schluenzen, F, Zarivach, R, Harms, J, Bashan, A, Tocilj, A, Albrecht, R, Yonath, A, Franceschi, F. | | Deposit date: | 2001-09-17 | | Release date: | 2001-10-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria.
Nature, 413, 2001
|
|
