6B36
 | |
6B3C
 | |
6B3H
 | | Crystal Structure of HIV Protease complexed with N-(2-(2-((6R,9S)-2,2-dioxido-2-thia-1,7-diazabicyclo[4.3.1]decan-9-yl)ethyl)-3-fluorophenyl)-3,3-bis(4-fluorophenyl)propanamide | | Descriptor: | CHLORIDE ION, HIV-1 Protease, N-(2-{2-[(6R,9S)-2,2-dioxo-2lambda~6~-thia-1,7-diazabicyclo[4.3.1]decan-9-yl]ethyl}-3-fluorophenyl)-3,3-bis(4-fluorophenyl)propanamide | | Authors: | Su, H.P. | | Deposit date: | 2017-09-21 | | Release date: | 2018-01-03 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Design and Synthesis of Piperazine Sulfonamide Cores Leading to Highly Potent HIV-1 Protease Inhibitors.
ACS Med Chem Lett, 8, 2017
|
|
6BLF
 
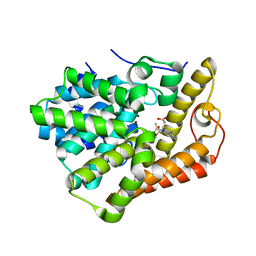 | | PDE2 complexed with 2-[6-fluoro-8-methylsulfonyl-9-[(1R)-1-[4-(trifluoromethyl)phenyl]ethyl]-1,2,3,4-tetrahydrocarbazol-1-yl]acetic acid | | Descriptor: | MAGNESIUM ION, ZINC ION, [(1S)-6-fluoro-8-(methylsulfonyl)-9-{(1R)-1-[4-(trifluoromethyl)phenyl]ethyl}-2,3,4,9-tetrahydro-1H-carbazol-1-yl]acetic acid, ... | | Authors: | Su, H.P. | | Deposit date: | 2017-11-10 | | Release date: | 2018-11-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Hydrolase complex
To Be Published
|
|
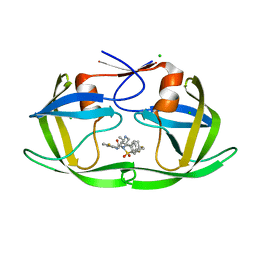
6B3F
 | |
3M9F
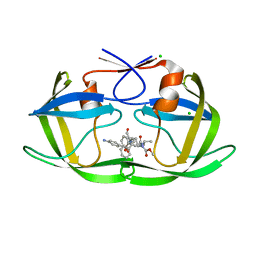 
 | | HIV protease complexed with compound 10b | | Descriptor: | CHLORIDE ION, HIV-1 protease, N-[(1S,5S)-5-{[(4-aminophenyl)sulfonyl](3-methylbutyl)amino}-1-methyl-6-oxohexyl]-Nalpha-(methoxycarbonyl)-beta-phenyl-L-phenylalaninamide | | Authors: | Su, H.P. | | Deposit date: | 2010-03-22 | | Release date: | 2010-06-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Epsilon substituted lysinol derivatives as HIV-1 protease inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3K7B
 
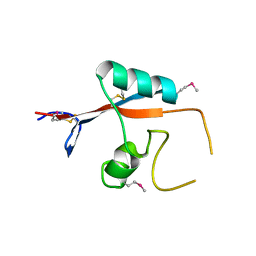 | |
4PMS
 
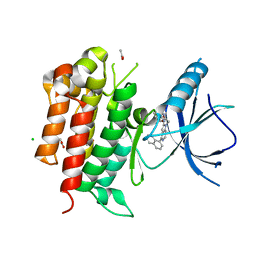 | | The structure of TrkA kinase bound to the inhibitor 4-naphthalen-1-yl-1-[(5-phenyl-1,2,4-oxadiazol-3-yl)methyl]-1H-pyrrolo[3,2-c]pyridine-2-carboxylic acid | | Descriptor: | 4-(naphthalen-1-yl)-1-[(5-phenyl-1,2,4-oxadiazol-3-yl)methyl]-1H-pyrrolo[3,2-c]pyridine-2-carboxylic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Su, H.P. | | Deposit date: | 2014-05-22 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Maximizing diversity from a kinase screen: identification of novel and selective pan-Trk inhibitors for chronic pain.
J.Med.Chem., 57, 2014
|
|
4PMT
 
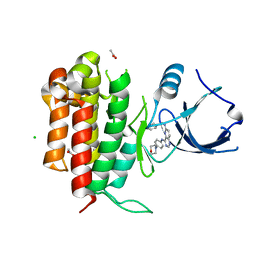 | |
4PMP
 
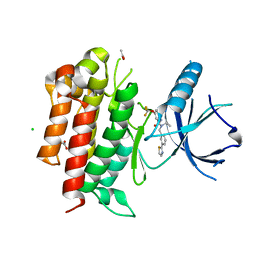 | |
4PMM
 
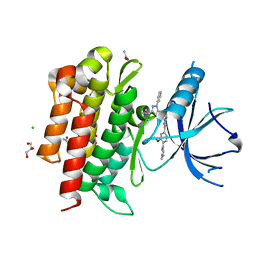 | |
5EAK
 
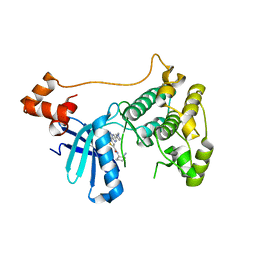 | |
3C64
 
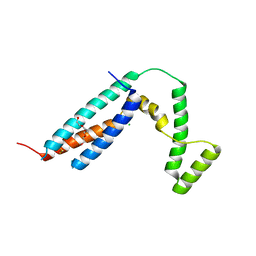 | | The MC179 portion of the Cysteine-rich Interdomain Region (CIDR) of a Plasmodium falciparum Erythrocyte Membrane Protein-1 (PfEMP1) | | Descriptor: | CHLORIDE ION, PfEMP1 variant 2 of strain MC, TETRAETHYLENE GLYCOL | | Authors: | Klein, M.M, Gittis, A.G, Su, H.P, Makobongo, M.O, Moore, J.M, Singh, S, Miller, L.H, Garboczi, D.N. | | Deposit date: | 2008-02-02 | | Release date: | 2008-09-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Cysteine-Rich Interdomain Region from the Highly Variable Plasmodium falciparum Erythrocyte Membrane Protein-1 Exhibits a Conserved Structure.
Plos Pathog., 4, 2008
|
|
1Z27
 
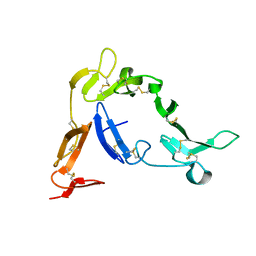 | | Crystal structure of Native Pvs25, an ookinete protein from Plasmodium vivax. | | Descriptor: | ookinete surface protein Pvs25 | | Authors: | Saxena, A.K, Singh, K, Su, H.P, Klein, M.M, Stowers, A.W, Saul, A.J, Long, C.A, Garboczi, D.N. | | Deposit date: | 2005-03-07 | | Release date: | 2005-12-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | The essential mosquito-stage P25 and P28 proteins from Plasmodium form tile-like triangular prisms
Nat.Struct.Mol.Biol., 13, 2006
|
|
1Z1Y
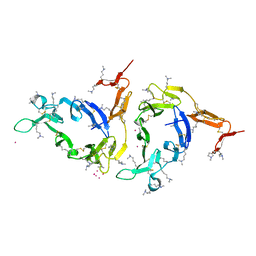 
 | | Crystal structure of Methylated Pvs25, an ookinete protein from Plasmodium vivax | | Descriptor: | YTTERBIUM (III) ION, ookinete surface protein Pvs25 | | Authors: | Saxena, A.K, Singh, K, Su, H.P, Klein, M.M, Stowers, A.W, Saul, A.J, Long, C.A, Garboczi, D.N. | | Deposit date: | 2005-03-07 | | Release date: | 2005-12-06 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The essential mosquito-stage P25 and P28 proteins from Plasmodium form tile-like triangular prisms
Nat.Struct.Mol.Biol., 13, 2006
|
|
1Z3G
 
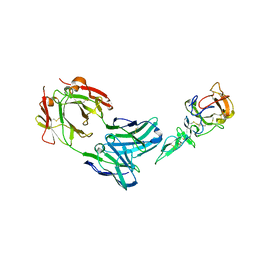 | | Crystal structure of complex between Pvs25 and Fab fragment of malaria transmission blocking antibody 2A8 | | Descriptor: | 2A8 Fab Heavy Chain, 2A8 Fab Light Chain, ookinete surface protein Pvs25 | | Authors: | Saxena, A.K, Singh, K, Su, H.P, Klein, M.M, Stowers, A.W, Saul, A.J, Long, C.A, Garboczi, D.N. | | Deposit date: | 2005-03-12 | | Release date: | 2005-12-06 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | The essential mosquito-stage P25 and P28 proteins from Plasmodium form tile-like triangular prisms
Nat.Struct.Mol.Biol., 13, 2006
|
|
5VZ6
 
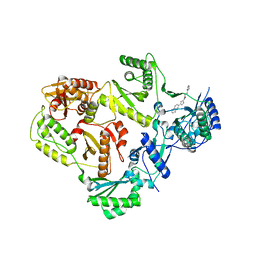 | | HIV Reverse Transcriptase complexed with (E)-3-(pyrimidin-2-yl)-N-(5-(5,6,7,8-tetrahydronaphthalen-2-yl)-1H-pyrazol-3-yl)acrylamide | | Descriptor: | 3-(pyrimidin-2-yl)-N-[3-(5,6,7,8-tetrahydronaphthalen-2-yl)-1H-pyrazol-5-yl]propanamide, HIV Reverse Transcriptase | | Authors: | Yan, Y, Su, H.P. | | Deposit date: | 2017-05-26 | | Release date: | 2018-05-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | HIV Reverse Transcriptase complexed with inhibitor
To Be Published
|
|
3BIK
 
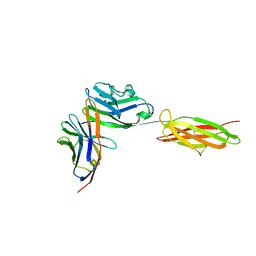 | | Crystal Structure of the PD-1/PD-L1 Complex | | Descriptor: | GLYCEROL, Programmed cell death 1 ligand 1, Programmed cell death protein 1 | | Authors: | Lin, D.Y, Tanaka, Y, Iwasaki, M, Gittis, A.G, Su, H.P, Mikami, B, Okazaki, T, Honjo, T, Minato, N, Garboczi, D.N. | | Deposit date: | 2007-11-30 | | Release date: | 2008-02-26 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | The PD-1/PD-L1 complex resembles the antigen-binding Fv domains of antibodies and T cell receptors.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3BIS
 
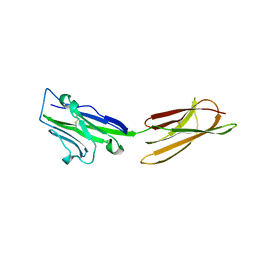 | | Crystal Structure of the PD-L1 | | Descriptor: | Programmed cell death 1 ligand 1 | | Authors: | Lin, D.Y, Tanaka, Y, Iwasaki, M, Gittis, A.G, Su, H.P, Mikami, B, Okazaki, T, Honjo, T, Minato, N, Garboczi, D.N. | | Deposit date: | 2007-11-30 | | Release date: | 2008-02-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | The PD-1/PD-L1 complex resembles the antigen-binding Fv domains of antibodies and T cell receptors.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3LP3
 
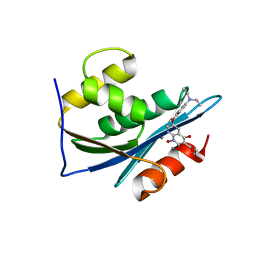 | | p15 HIV RNaseH domain with inhibitor MK3 | | Descriptor: | 3-[4-(diethylamino)phenoxy]-6-(ethoxycarbonyl)-5,8-dihydroxy-7-oxo-7,8-dihydro-1,8-naphthyridin-1-ium, MANGANESE (II) ION, p15 | | Authors: | Yan, Y, Munshi, S.K, Prasad, G.S, Su, H.P. | | Deposit date: | 2010-02-04 | | Release date: | 2010-06-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis for the inhibition of RNase H activity of HIV-1 reverse transcriptase by RNase H active site-directed inhibitors.
J.Virol., 84, 2010
|
|
3LP2
 
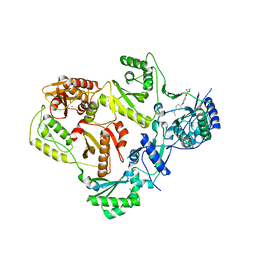 | | HIV-1 reverse transcriptase with inhibitor | | Descriptor: | 3-[4-(diethylamino)phenoxy]-6-(ethoxycarbonyl)-5,8-dihydroxy-7-oxo-7,8-dihydro-1,8-naphthyridin-1-ium, MANGANESE (II) ION, Reverse transcriptase/ribonuclease H, ... | | Authors: | Yan, Y, Munshi, S.K, Prasad, G.S, Su, H.P. | | Deposit date: | 2010-02-04 | | Release date: | 2010-06-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis for the inhibition of RNase H activity of HIV-1 reverse transcriptase by RNase H active site-directed inhibitors.
J.Virol., 84, 2010
|
|
3LP1
 
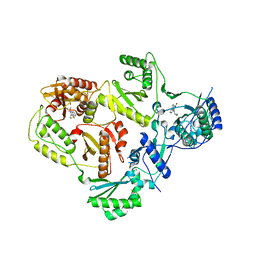 | | HIV-1 reverse transcriptase with inhibitor | | Descriptor: | 11-CYCLOPROPYL-5,11-DIHYDRO-4-METHYL-6H-DIPYRIDO[3,2-B:2',3'-E][1,4]DIAZEPIN-6-ONE, 3-cyclopentyl-1,4-dihydroxy-1,8-naphthyridin-2(1H)-one, MANGANESE (II) ION, ... | | Authors: | Yan, Y, Munshi, S.K, Prasad, G.S, Su, H.P. | | Deposit date: | 2010-02-04 | | Release date: | 2010-06-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Structural basis for the inhibition of RNase H activity of HIV-1 reverse transcriptase by RNase H active site-directed inhibitors.
J.Virol., 84, 2010
|
|
3LP0
 
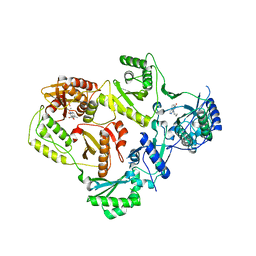 | | HIV-1 reverse transcriptase with inhibitor | | Descriptor: | 11-CYCLOPROPYL-5,11-DIHYDRO-4-METHYL-6H-DIPYRIDO[3,2-B:2',3'-E][1,4]DIAZEPIN-6-ONE, MANGANESE (II) ION, Reverse transcriptase/ribonuclease H, ... | | Authors: | Yan, Y, Munshi, S.K, Prasad, G.S, Su, H.P. | | Deposit date: | 2010-02-04 | | Release date: | 2010-06-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structural basis for the inhibition of RNase H activity of HIV-1 reverse transcriptase by RNase H active site-directed inhibitors.
J.Virol., 84, 2010
|
|
3RVG
 
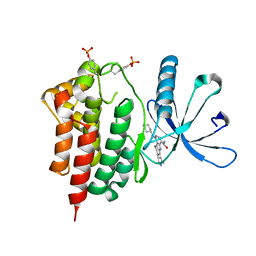 | | Crystals structure of Jak2 with a 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitor | | Descriptor: | 1-(cyclohexylamino)-7-(1-methyl-1H-pyrazol-4-yl)-5H-pyrido[4,3-b]indole-4-carboxamide, Tyrosine-protein kinase JAK2 | | Authors: | Lim, J, Taoka, B, Otte, R.D, Spencer, K, Dinsmore, C.J, Altman, M.D, Chan, G, Rosenstein, C, Sharma, S, Su, H.P, Szewczak, A.A, Xu, L, Yin, H, Zugay-Murphy, J, Marshall, C.G, Young, J.R. | | Deposit date: | 2011-05-06 | | Release date: | 2012-03-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | Discovery of 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitors of Janus kinase 2 (JAK2) for the treatment of myeloproliferative disorders.
J.Med.Chem., 54, 2011
|
|
5EJ0
 
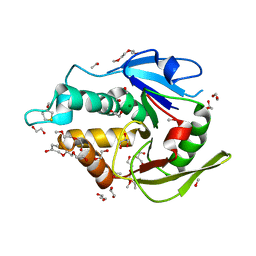 | | The vaccinia virus H3 envelope protein, a major target of neutralizing antibodies, exhibits a glycosyltransferase fold and binds UDP-Glucose | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, 2-(2-METHOXYETHOXY)ETHANOL, ... | | Authors: | Singh, K, Gittis, A.G, Gitti, R.K, Ostazesky, S.A, Su, H.P, Garboczi, D.N. | | Deposit date: | 2015-10-30 | | Release date: | 2016-03-16 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Vaccinia Virus H3 Envelope Protein, a Major Target of Neutralizing Antibodies, Exhibits a Glycosyltransferase Fold and Binds UDP-Glucose.
J.Virol., 90, 2016
|
|
