8ETF
 
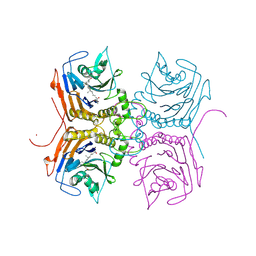 | | Bile Salt Hydrolase B from Lactobacillus gasseri with covalent inhibitor bound | | Descriptor: | (5R)-1-fluoro-5-[(1R,3aS,3bR,5aR,7R,9aS,9bS,11aR)-7-hydroxy-9a,11a-dimethylhexadecahydro-1H-cyclopenta[a]phenanthren-1-yl]hexan-2-one (non-preferred name), Choloylglycine hydrolase, NICKEL (II) ION | | Authors: | Walker, M.E, Redinbo, M.R. | | Deposit date: | 2022-10-17 | | Release date: | 2023-11-01 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structural diversity of bile salt hydrolases reveals rationale for substrate selectivity
To Be Published
|
|
8EWT
 
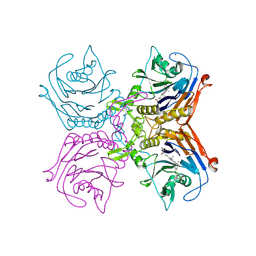 | | Bile salt hydrolase A from Lactobacillus gasseri bound to covalent probe | | Descriptor: | (5R)-5-{(1R,3aS,3bR,5aR,7R,9aS,9bS,11aR)-9a,11a-dimethyl-7-[(prop-2-yn-1-yl)oxy]hexadecahydro-1H-cyclopenta[a]phenanthren-1-yl}-1-fluorohexan-2-one (non-preferred name), Conjugated bile salt hydrolase, SODIUM ION | | Authors: | Walker, M.E, Redinbo, M.R. | | Deposit date: | 2022-10-24 | | Release date: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Structural diversity of bile salt hydrolases reveals rationale for substrate selectivity
To Be Published
|
|
2O9I
 
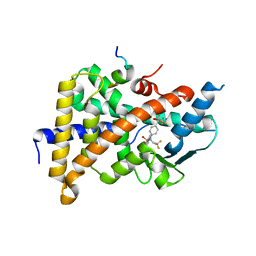 | | Crystal Structure of the Human Pregnane X Receptor LBD in complex with an SRC-1 coactivator peptide and T0901317 | | Descriptor: | N-(2,2,2-TRIFLUOROETHYL)-N-{4-[2,2,2-TRIFLUORO-1-HYDROXY-1-(TRIFLUOROMETHYL)ETHYL]PHENYL}BENZENESULFONAMIDE, Nuclear Receptor Coactivator 1 isoform 3, Orphan nuclear receptor PXR | | Authors: | Xue, Y, Redinbo, M.R. | | Deposit date: | 2006-12-13 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the PXR-T1317 complex provides a scaffold to examine the potential for receptor antagonism.
Bioorg.Med.Chem., 15, 2007
|
|
2O6L
 
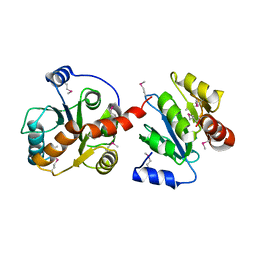 | |
8FAO
 
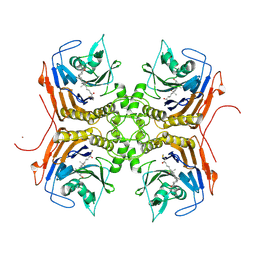 | | Bile Salt Hydrolase B from Lactobacillus gasseri with covalent inhibitor bound | | Descriptor: | (5R)-5-[(1R,3aS,3bR,5aR,7R,9aS,9bS,11S,11aR)-7,11-dihydroxy-9a,11a-dimethylhexadecahydro-1H-cyclopenta[a]phenanthren-1-yl]-1-fluorohexan-2-one (non-preferred name), Choloylglycine hydrolase, NICKEL (II) ION | | Authors: | Grundy, M.K, Walker, M.E, Redinbo, M.R. | | Deposit date: | 2022-11-28 | | Release date: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structural diversity of bile salt hydrolases reveals rationale for substrate selectivity
To Be Published
|
|
1A36
 
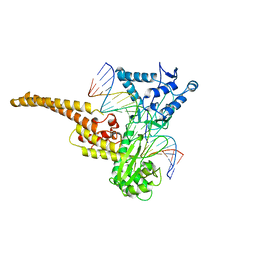 | | TOPOISOMERASE I/DNA COMPLEX | | Descriptor: | DNA (5'-D(*AP*AP*AP*AP*AP*GP*AP*CP*TP*TP*AP*GP*AP*AP*AP*AP*A P*TP*TP*TP*TP*T)- 3'), DNA (5'-D(*AP*AP*AP*AP*AP*TP*TP*TP*TP*TP*CP*TP*AP*AP*GP*TP*C P*TP*TP*TP*TP*T)- 3'), TOPOISOMERASE I | | Authors: | Stewart, L, Redinbo, M.R, Qiu, X, Champoux, J.J, Hol, W.G.J. | | Deposit date: | 1998-01-29 | | Release date: | 1998-08-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A model for the mechanism of human topoisomerase I.
Science, 279, 1998
|
|
1RR8
 
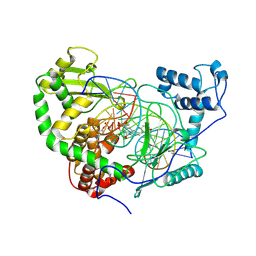 | | Structural Mechanisms of Camptothecin Resistance by Mutations in Human Topoisomerase I | | Descriptor: | (S)-10-[(DIMETHYLAMINO)METHYL]-4-ETHYL-4,9-DIHYDROXY-1H-PYRANO[3',4':6,7]INOLIZINO[1,2-B]-QUINOLINE-3,14(4H,12H)-DIONE, 2-(1-DIMETHYLAMINOMETHYL-2-HYDROXY-8-HYDROXYMETHYL-9-OXO-9,11-DIHYDRO-INDOLIZINO[1,2-B]QUINOLIN-7-YL)-2-HYDROXY-BUTYRIC ACID, 5'-D(*AP*AP*AP*AP*AP*GP*AP*CP*TP*T*GP*GP*AP*AP*AP*AP*AP*TP*TP*TP*TP*T)-3', ... | | Authors: | Chrencik, J.E, Staker, B.L, Burgin, A.B, Stewart, L, Redinbo, M.R. | | Deposit date: | 2003-12-08 | | Release date: | 2004-07-06 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Mechanisms of camptothecin resistance by human topoisomerase I mutations
J.Mol.Biol., 339, 2004
|
|
2HRR
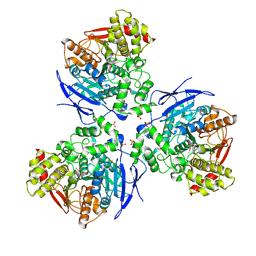 
 | |
1SKX
 
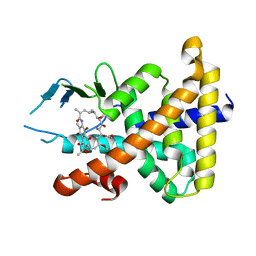 | | Structural Disorder in the Complex of Human PXR and the Macrolide Antibiotic Rifampicin | | Descriptor: | Orphan nuclear receptor PXR, RIFAMPICIN | | Authors: | Chrencik, J.E, Xue, Y, Orans, J.O, Redinbo, M.R. | | Deposit date: | 2004-03-05 | | Release date: | 2005-03-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural disorder in the complex of human pregnane x receptor and the macrolide antibiotic rifampicin
Mol.Endocrinol., 19, 2005
|
|
2H7C
 
 | | Crystal structure of human carboxylesterase in complex with Coenzyme A | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, COENZYME A, ... | | Authors: | Bencharit, S, Edwards, C.C, Morton, C.L, Howard-Williams, E.L, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2006-06-02 | | Release date: | 2006-08-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Multisite promiscuity in the processing of endogenous substrates by human carboxylesterase 1
J.Mol.Biol., 363, 2006
|
|
2Q7U
 
 | |
1ILH
 
 | |
1ILG
 
 | |
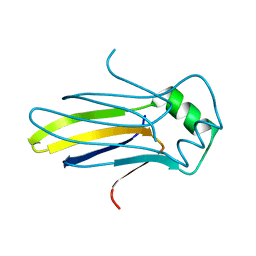
4K59
 
 | |
1K4Y
 
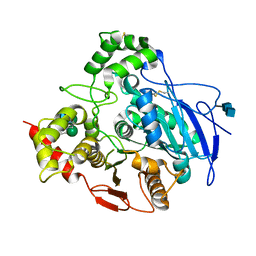 | | Crystal Structure of Rabbit Liver Carboxylesterase in Complex with 4-piperidino-piperidine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 4-PIPERIDINO-PIPERIDINE, LIVER CARBOXYLESTERASE, ... | | Authors: | Bencharit, S, Morton, C.L, Howard-Williams, E.L, Danks, M.K, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2001-10-09 | | Release date: | 2002-05-01 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into CPT-11 activation by mammalian carboxylesterases.
Nat.Struct.Biol., 9, 2002
|
|
1LPQ
 
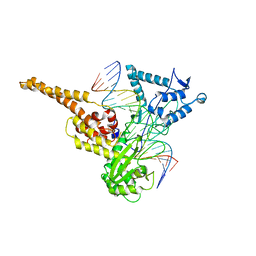 | | Human DNA Topoisomerase I (70 Kda) In Non-Covalent Complex With A 22 Base Pair DNA Duplex Containing an 8-oxoG Lesion | | Descriptor: | 5'-D(*AP*AP*AP*AP*AP*GP*AP*CP*TP*TP*(8OG)P*GP*AP*AP*AP*AP*AP*TP*TP*TP*TP*T)-3', 5'-D(*AP*AP*AP*AP*AP*TP*TP*TP*TP*TP*CP*CP*AP*AP*GP*TP*CP*TP*TP*TP*TP*T)-3', DNA topoisomerase I | | Authors: | Lesher, D.T, Pommier, Y, Stewart, L, Redinbo, M.R. | | Deposit date: | 2002-05-08 | | Release date: | 2002-08-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.14 Å) | | Cite: | 8-Oxoguanine rearranges the active site of human topoisomerase I
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
3LPF
 
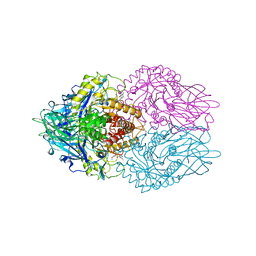 | | Structure of E. coli beta-Glucuronidase bound with a novel, potent inhibitor 1-((6,7-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)-1-(2-hydroxyethyl)-3-(3-methoxyphenyl)thiourea | | Descriptor: | 1-[(6,7-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl]-1-(2-hydroxyethyl)-3-(3-methoxyphenyl)thiourea, Beta-glucuronidase | | Authors: | Wallace, B.D, Redinbo, M.R. | | Deposit date: | 2010-02-05 | | Release date: | 2010-11-17 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Alleviating cancer drug toxicity by inhibiting a bacterial enzyme.
Science, 330, 2010
|
|
3LPG
 
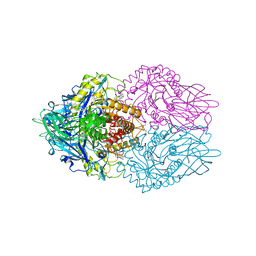 | |
1YUC
 
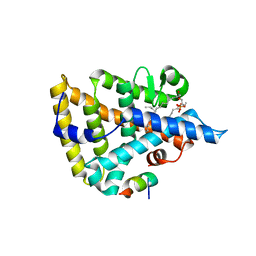 | | Human Nuclear Receptor Liver Receptor Homologue-1, LRH-1, Bound to Phospholipid and a Fragment of Human SHP | | Descriptor: | GLYCEROL, L-ALPHA-PHOSPHATIDYL-BETA-OLEOYL-GAMMA-PALMITOYL-PHOSPHATIDYLETHANOLAMINE, Nuclear receptor 0B2, ... | | Authors: | Ortlund, E.A, Yoonkwang, L, Solomon, I.H, Hager, J.M, Safi, R, Choi, Y, Guan, Z, Tripathy, A, Raetz, C.R.H, McDonnell, D.P, Moore, D.D, Redinbo, M.R. | | Deposit date: | 2005-02-13 | | Release date: | 2005-03-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Modulation of human nuclear receptor LRH-1 activity by phospholipids and SHP
Nat.Struct.Mol.Biol., 12, 2005
|
|
1YA4
 
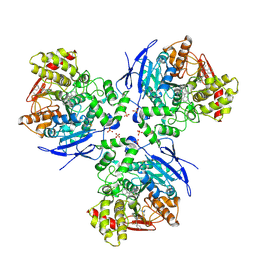 | | Crystal Structure of Human Liver Carboxylesterase 1 in complex with tamoxifen | | Descriptor: | (Z)-2-[4-(1,2)-DIPHENYL-1-BUTENYL)-PHENOXY]-N,N-DIMETHYLETHANAMINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, CES1 protein, ... | | Authors: | Fleming, C.D, Bencharit, S, Edwards, C.C, Hyatt, J.L, Morton, C.L, Howard-Williams, E.L, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2004-12-17 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural insights into drug processing by human carboxylesterase 1: tamoxifen, mevastatin, and inhibition by benzil.
J.Mol.Biol., 352, 2005
|
|
1YAJ
 
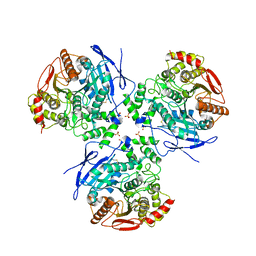 | | Crystal Structure of Human Liver Carboxylesterase in complex with benzil | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, BENZOIC ACID, CES1 protein, ... | | Authors: | Fleming, C.D, Bencharit, S, Edwards, C.C, Hyatt, J.L, Morton, C.M, Howard-Williams, E.L, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2004-12-17 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural insights into drug processing by human carboxylesterase 1: tamoxifen, mevastatin, and inhibition by benzil.
J.Mol.Biol., 352, 2005
|
|
1YAH
 
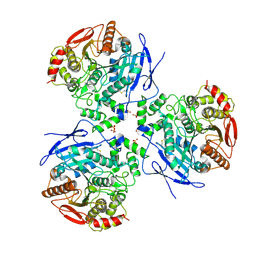 | | Crystal Structure of Human Liver Carboxylesterase complexed to Etyl Acetate; A Fatty Acid Ethyl Ester Analogue | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CES1 protein, ETHYL ACETATE, ... | | Authors: | Fleming, C.D, Bencahrit, S, Edwards, C.C, Hyatt, J.L, Morton, C.L, Howard-Williams, E.L, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2004-12-17 | | Release date: | 2005-08-02 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural insights into drug processing by human carboxylesterase 1: tamoxifen, mevastatin, and inhibition by benzil.
J.Mol.Biol., 352, 2005
|
|
3MBO
 
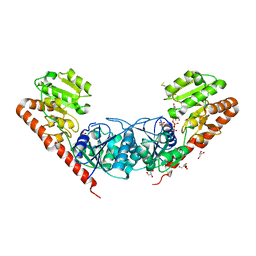 | | Crystal Structure of the Glycosyltransferase BaBshA bound with UDP and L-malate | | Descriptor: | D-MALATE, GLYCEROL, Glycosyltransferase, ... | | Authors: | Wallace, B.D, Claiborne, A, Redinbo, M.R. | | Deposit date: | 2010-03-25 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.307 Å) | | Cite: | Characterization of the N-Acetyl-alpha-d-glucosaminyl
l-Malate Synthase and Deacetylase Functions for Bacillithiol Biosynthesis
in Bacillus anthracis
Biochemistry, 49, 2010
|
|
1YA8
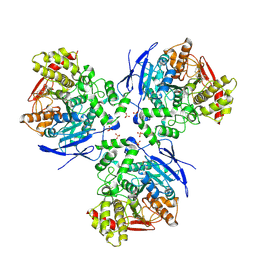 
 | | Crystal Structure of Human Liver Carboxylesterase in complex with cleavage products of Mevastatin | | Descriptor: | (1S,7S,8S,8AR)-1,2,3,7,8,8A-HEXAHYDRO-7-METHYL-8-[2-[(2R,4R)-TETRAHYDRO-4-HY DROXY-6-OXO-2H-PYRAN-2-YL]ETHYL]-1-NAPHTHALENOL, 2-METHYLBUTANOIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Fleming, C.D, Bencharit, S, Edwards, C.C, Hyatt, J.L, Morton, C.L, Howard-Williams, E.L, Potter, P.M, Redinbo, M.R. | | Deposit date: | 2004-12-17 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural insights into drug processing by human carboxylesterase 1: tamoxifen, mevastatin, and inhibition by benzil.
J.Mol.Biol., 352, 2005
|
|
3GS3
 
 | |
