3E0H
 
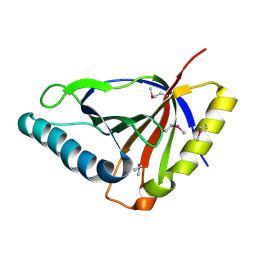 | | Crystal structure of an uncharacterized protein from Chlorobium tepidum. NorthEast Structural Genomics target CtR107. | | Descriptor: | uncharacterized protein | | Authors: | Seetharaman, J, Chen, Y, Wang, H, Janjua, H, Foote, E.L, Xiao, R, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-07-31 | | Release date: | 2008-09-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Crystal structure of an uncharacterized protein from Chlorobium tepidum. NorthEast Structural Genomics target CtR107.
To be Published
|
|
4H1Z
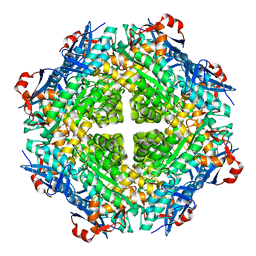 
 | | Crystal structure of putative isomerase from Sinorhizobium meliloti, open loop conformation (target EFI-502104) | | Descriptor: | CHLORIDE ION, Enolase Q92Zs5, FORMIC ACID, ... | | Authors: | Patskovsky, Y, Toro, R, Bhosle, R, Hillerich, B, Seidel, R.D, Washington, E, Scott Glenn, A, Chowdhury, S, Evans, B, Hammonds, J, Zencheck, W.D, Imker, H.J, Gerlt, J.A, Almo, S.C, Enzyme Function Initiative (EFI) | | Deposit date: | 2012-09-11 | | Release date: | 2012-09-26 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.012 Å) | | Cite: | Crystal Structure of Enolase Q92Zs5 (Target EFI-502104) from Sinorhizobium meliloti
To be Published
|
|
6XUR
 
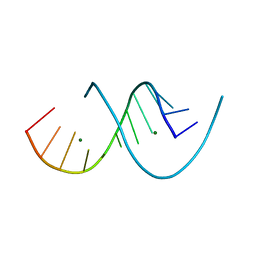 | | RNA dodecamer with a 6-hydrazino-2-aminopurine modified base | | Descriptor: | MAGNESIUM ION, RNA dodecamer with a 6-hydrazino-2-aminopurine modified base | | Authors: | Ennifar, E, Micura, R, Gasser, C, Brillet, K. | | Deposit date: | 2020-01-21 | | Release date: | 2020-02-19 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Thioguanosine Conversion Enables mRNA-Lifetime Evaluation by RNA Sequencing Using Double Metabolic Labeling (TUC-seq DUAL).
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6XY9
 
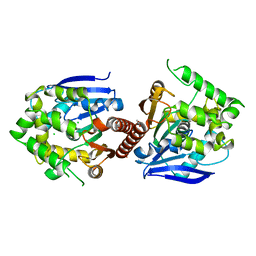 | | Crystal structure of haloalkane dehalogenase DbeA-M1 loop variant from Bradyrhizobium elkanii | | Descriptor: | ACETATE ION, CHLORIDE ION, Haloalkane dehalogenase | | Authors: | Prudnikova, T, Rezacova, P, Kuta Smatanova, I, Chaloupkova, R, Damborsky, J. | | Deposit date: | 2020-01-29 | | Release date: | 2020-07-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and catalytic effects of surface loop-helix transplantation within haloalkane dehalogenase family.
Comput Struct Biotechnol J, 18, 2020
|
|
4P1V
 
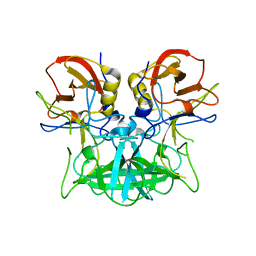 | | Structure of the P domain from a GI.7 Norovirus variant in complex with H-type 2 HBGA | | Descriptor: | P domain of VPI, alpha-L-fucopyranose-(1-2)-beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Shanker, S, Czako, R, Sankaran, B, Atmar, R, Estes, M, Prasad, B.V.V. | | Deposit date: | 2014-02-27 | | Release date: | 2014-04-02 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5497 Å) | | Cite: | Structural analysis of determinants of histo-blood group antigen binding specificity in genogroup I noroviruses.
J.Virol., 88, 2014
|
|
5GR3
 
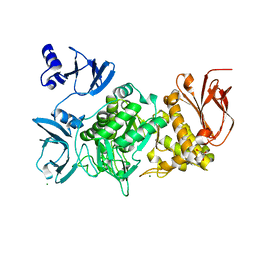 | |
6XIT
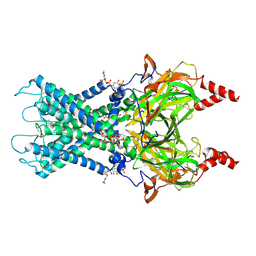 
 | | Cryo-EM structure of the G protein-gated inward rectifier K+ channel GIRK2 (Kir3.2) in complex with PIP2 | | Descriptor: | G protein-activated inward rectifier potassium channel 2, POTASSIUM ION, [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate | | Authors: | Niu, Y, Tao, X, MacKinnon, R. | | Deposit date: | 2020-06-21 | | Release date: | 2020-10-07 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Cryo-EM analysis of PIP 2 regulation in mammalian GIRK channels.
Elife, 9, 2020
|
|
6XD0
 
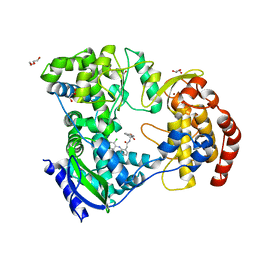 | | Dengue serotype 3 RNA-dependent RNA polymerase bound to NITD-434 | | Descriptor: | 2-[({2-[(2,6-dichlorophenyl)amino]phenyl}acetyl)amino]-2,3-dihydro-1H-indene-2-carboxylic acid, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Arora, R, Benson, T.E, Liew, C.W, Lescar, J. | | Deposit date: | 2020-06-09 | | Release date: | 2020-09-23 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.012 Å) | | Cite: | Two RNA Tunnel Inhibitors Bind in Highly Conserved Sites in Dengue Virus NS5 Polymerase: Structural and Functional Studies.
J.Virol., 94, 2020
|
|
6BGR
 
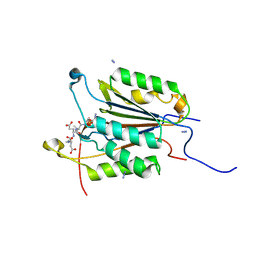 | | Caspase-3 Mutant - S150E | | Descriptor: | AZIDE ION, Ac-Asp-Glu-Val-Asp-CMK, Caspase-3 | | Authors: | Thomas, M.E, Grinshpon, R, Swartz, P.D, Clark, A.C. | | Deposit date: | 2017-10-29 | | Release date: | 2018-02-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Modifications to a common phosphorylation network provide individualized control in caspases.
J. Biol. Chem., 293, 2018
|
|
6XFM
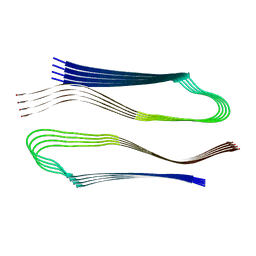 
 | | Molecular structure of the core of amyloid-like fibrils formed by residues 111-214 of FUS | | Descriptor: | RNA-binding protein FUS | | Authors: | Tycko, R, Lee, M, Ghosh, U, Thurber, K, Kato, M. | | Deposit date: | 2020-06-15 | | Release date: | 2020-10-07 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (2.62 Å) | | Cite: | Molecular structure and interactions within amyloid-like fibrils formed by a low-complexity protein sequence from FUS.
Nat Commun, 11, 2020
|
|
5GQU
 
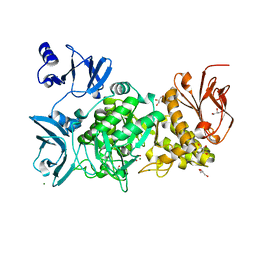 | |
5GR6
 
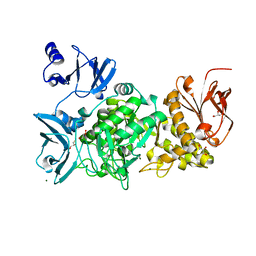 | |
6P10
 
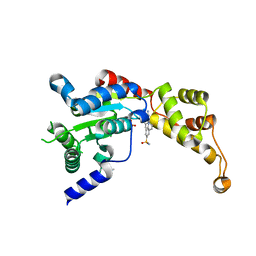 | | Structure of spastin AAA domain (N527C mutant) in complex with JNJ-7706621 inhibitor | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-({5-amino-1-[(2,6-difluorophenyl)carbonyl]-1H-1,2,4-triazol-3-yl}amino)benzenesulfonamide, Drosophila melanogaster Spastin AAA domain, ... | | Authors: | Pisa, R, Cupido, T, Kapoor, T.M. | | Deposit date: | 2019-05-17 | | Release date: | 2019-07-24 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.301 Å) | | Cite: | Analyzing Resistance to Design Selective Chemical Inhibitors for AAA Proteins.
Cell Chem Biol, 26, 2019
|
|
6BJC
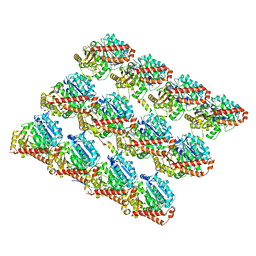 
 | | TPX2_mini decorated GMPCPP-microtubule | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER, ... | | Authors: | Zhang, R, Nogales, E. | | Deposit date: | 2017-11-05 | | Release date: | 2017-11-22 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structural insight into TPX2-stimulated microtubule assembly.
Elife, 6, 2017
|
|
6P4H
 
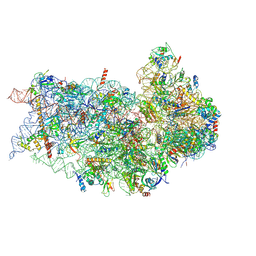 | | Structure of a mammalian small ribosomal subunit in complex with the Israeli Acute Paralysis Virus IRES (Class 2) | | Descriptor: | 18S rRNA, IAPV-IRES, RACK1, ... | | Authors: | Acosta-Reyes, F.J, Neupane, R, Frank, J, Fernandez, I.S. | | Deposit date: | 2019-05-27 | | Release date: | 2019-09-18 | | Last modified: | 2020-01-08 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The Israeli acute paralysis virus IRES captures host ribosomes by mimicking a ribosomal state with hybrid tRNAs.
Embo J., 38, 2019
|
|
6XX7
 
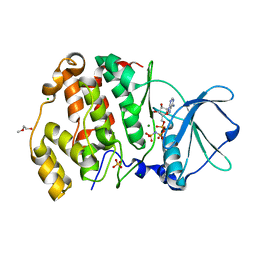 | | Arabidopsis thaliana Casein Kinase 2 (CK2) alpha-1 crystal in complex with ANP | | Descriptor: | CHLORIDE ION, Casein kinase II subunit alpha-1, MAGNESIUM ION, ... | | Authors: | Demulder, M, De Veylder, L, Loris, R. | | Deposit date: | 2020-01-27 | | Release date: | 2020-04-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of Arabidopsis thaliana casein kinase 2 alpha 1.
Acta Crystallogr.,Sect.F, 76, 2020
|
|
2OTD
 
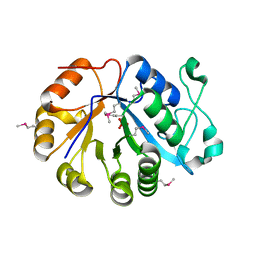 | | The crystal structure of the glycerophosphodiester phosphodiesterase from Shigella flexneri 2a | | Descriptor: | Glycerophosphodiester phosphodiesterase, PHOSPHATE ION | | Authors: | Zhang, R, Wu, R, Clancy, S, Jiang, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2007-02-07 | | Release date: | 2007-03-06 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The crystal structure of the glycerophosphodiester phosphodiesterase from Shigella flexneri 2a
To be Published
|
|
2Y39
 
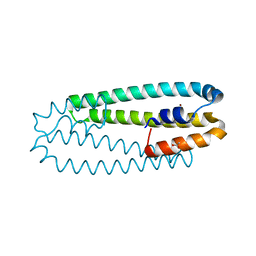 | | Ni-bound form of Cupriavidus metallidurans CH34 CnrXs | | Descriptor: | ACETATE ION, NICKEL (II) ION, NICKEL AND COBALT RESISTANCE PROTEIN CNRR | | Authors: | Trepreau, J, Girard, E, Maillard, A.P, de Rosny, E, Petit-Haertlein, I, Kahn, R, Coves, J. | | Deposit date: | 2010-12-20 | | Release date: | 2011-03-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Structural Basis for Metal Sensing by Cnrx.
J.Mol.Biol., 408, 2011
|
|
6BPM
 
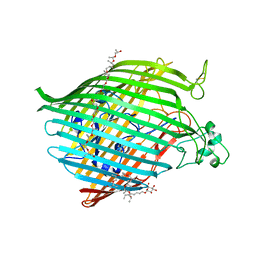 | | The crystal structure of the Ferric-Catecholate import receptor Fiu from K12 E. coli: Closed form (C21) | | Descriptor: | (20S)-2,5,8,11,14,17-HEXAMETHYL-3,6,9,12,15,18-HEXAOXAHENICOSANE-1,20-DIOL, Catecholate siderophore receptor Fiu, octyl beta-D-glucopyranoside | | Authors: | Grinter, R. | | Deposit date: | 2017-11-23 | | Release date: | 2018-11-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The structure of the bacterial iron-catecholate transporter Fiu suggests that it imports substrates via a two-step mechanism.
J.Biol.Chem., 2019
|
|
6P5I
 
 | | Structure of a mammalian 80S ribosome in complex with the Israeli Acute Paralysis Virus IRES (Class 1) | | Descriptor: | 18S rRNA, 28S rRNA, 5.8S rRNA, ... | | Authors: | Acosta-Reyes, F.J, Neupane, R, Frank, J, Fernandez, I.S. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2020-02-26 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | The Israeli acute paralysis virus IRES captures host ribosomes by mimicking a ribosomal state with hybrid tRNAs.
Embo J., 38, 2019
|
|
6P4G
 
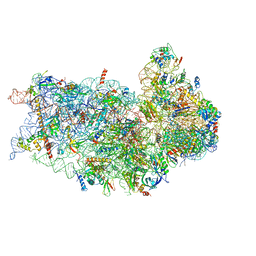 | | Structure of a mammalian small ribosomal subunit in complex with the Israeli Acute Paralysis Virus IRES (Class 1) | | Descriptor: | 18S rRNA, IAPV-IRES, RACK1, ... | | Authors: | Acosta-Reyes, F.J, Neupane, R, Frank, J, Fernandez, I.S. | | Deposit date: | 2019-05-27 | | Release date: | 2019-09-18 | | Last modified: | 2020-01-08 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | The Israeli acute paralysis virus IRES captures host ribosomes by mimicking a ribosomal state with hybrid tRNAs.
Embo J., 38, 2019
|
|
6BJ5
 
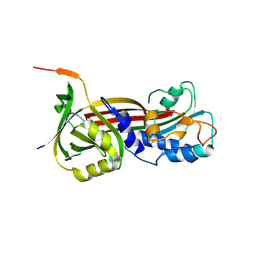 | | Structure of the Clinically used Myxomaviral Serine Protease Inhibitor 1 (SERP-1) | | Descriptor: | 2,5,8,11,14,17-HEXAOXANONADECAN-19-OL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mahon, B.P, Lomelino, C.L, McKenna, R. | | Deposit date: | 2017-11-05 | | Release date: | 2018-11-07 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the Clinically used Myxomaviral Serine Protease Inhibitor 1 (SERP-1)
To Be Published
|
|
3DM3
 
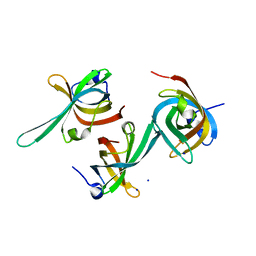 | | Crystal structure of a domain of a Replication factor A protein, from Methanocaldococcus jannaschii. NorthEast Structural Genomics target MjR118E | | Descriptor: | Replication factor A, SODIUM ION | | Authors: | Seetharaman, J, Su, M, Maglaqui, M, Janjua, H, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-26 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of a domain of a Replication factor A protein, from Methanocaldococcus jannaschii. NorthEast Structural Genomics target MjR118E
To be Published
|
|
6XXW
 
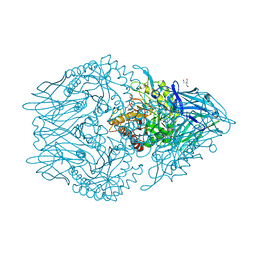 | | Structure of beta-D-Glucuronidase for Dictyoglomus thermophilum. | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Beta-glucuronidase, ... | | Authors: | Lafite, P, Daniellou, R. | | Deposit date: | 2020-01-28 | | Release date: | 2020-07-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.851 Å) | | Cite: | Thioglycoligation of aromatic thiols using a natural glucuronide donor.
Org.Biomol.Chem., 18, 2020
|
|
6XXY
 
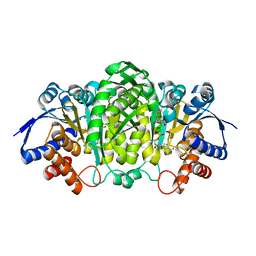 | | Crystal structure of Haemophilus influenzae 3-isopropylmalate dehydrogenase in complex with O-isobutenyl oxalylhydroxamate. | | Descriptor: | 2-(2-methylprop-2-enoxyamino)-2-oxidanylidene-ethanoic acid, 3-isopropylmalate dehydrogenase, MAGNESIUM ION, ... | | Authors: | Miggiano, R, Rossi, F, Martignon, S, Rizzi, M. | | Deposit date: | 2020-01-28 | | Release date: | 2020-02-26 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystal structure of Haemophilus influenzae 3-isopropylmalate dehydrogenase (LeuB) in complex with the inhibitor O-isobutenyl oxalylhydroxamate.
Biochem.Biophys.Res.Commun., 524, 2020
|
|
