6H43
 
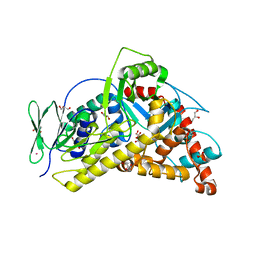 | | Flavin-dependent Tryptophan 6-halogenase Thal | | Descriptor: | GLYCEROL, PHOSPHATE ION, POTASSIUM ION, ... | | Authors: | Moritzer, A, Minges, H, Frese, M, Sewald, N, Niemann, H.H. | | Deposit date: | 2018-07-20 | | Release date: | 2018-12-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based switch of regioselectivity in the flavin-dependent tryptophan 6-halogenase Thal.
J. Biol. Chem., 294, 2019
|
|
6H44
 
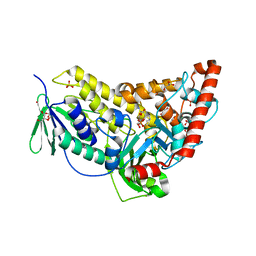 | | Flavin-dependent Tryptophan 6-halogenase Thal in complex with tryptophan | | Descriptor: | GLYCEROL, PHOSPHATE ION, POTASSIUM ION, ... | | Authors: | Moritzer, A, Minges, H, Sewald, N, Niemann, H.H. | | Deposit date: | 2018-07-20 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structure-based switch of regioselectivity in the flavin-dependent tryptophan 6-halogenase Thal.
J. Biol. Chem., 294, 2019
|
|
6SLT
 
 | |
6FRL
 
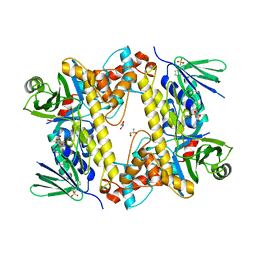 | | BrvH, a flavin-dependent halogenase from Brevundimonas sp. BAL3 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Widmann, C, Neubauer, P.R, Sewald, N, Niemann, H.H. | | Deposit date: | 2018-02-16 | | Release date: | 2018-05-23 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A flavin-dependent halogenase from metagenomic analysis prefers bromination over chlorination.
PLoS ONE, 13, 2018
|
|
5G36
 
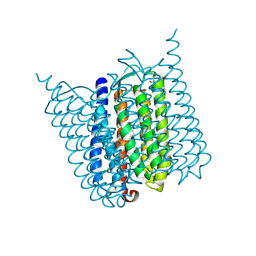 | | Yellow form of Halorhodopsin from Halobacterium salinarum in a new rhombohedral crystal form | | Descriptor: | CHLORIDE ION, HALORHODOPSIN, RETINAL, ... | | Authors: | Schreiner, M, Schlesinger, R, Heberle, J, Niemann, H.H. | | Deposit date: | 2016-04-20 | | Release date: | 2016-08-31 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of Halobacterium Salinarum Halorhodopsin with Partially Depopulated Primary Chloride Binding Site
Acta Crystallogr.,Sect.F, 72, 2016
|
|
1XDU
 
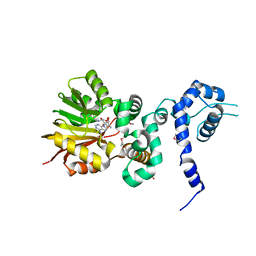 | | Crystal structure of Aclacinomycin-10-hydroxylase (RdmB) in complex with Sinefungin (SFG) | | Descriptor: | ACETATE ION, Protein RdmB, SINEFUNGIN | | Authors: | Jansson, A, Koskiniemi, H, Erola, A, Wang, J, Mantsala, P, Schneider, G, Niemi, J. | | Deposit date: | 2004-09-08 | | Release date: | 2004-11-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Aclacinomycin 10-Hydroxylase Is a Novel Substrate-assisted Hydroxylase Requiring S-Adenosyl-L-methionine as Cofactor
J.Biol.Chem., 280, 2005
|
|
5AHY
 
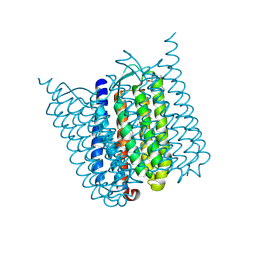 | | Halorhodopsin from Halobacterium salinarum in a new rhombohedral crystal form | | Descriptor: | CHLORIDE ION, HALORHODOPSIN, RETINAL, ... | | Authors: | Schreiner, M, Schlesinger, R, Heberle, J, Niemann, H.H. | | Deposit date: | 2015-02-11 | | Release date: | 2015-04-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structure of Halorhodopsin from Halobacterium Salinarum in a New Crystal Form that Imposes Little Restraint on the E-F Loop.
J.Struct.Biol., 190, 2015
|
|
5AHZ
 
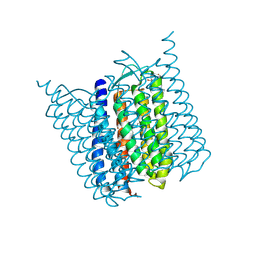 | | Bromide-bound form of Halorhodopsin from Halobacterium salinarum in a new rhombohedral crystal form | | Descriptor: | BROMIDE ION, HALORHODOPSIN, RETINAL, ... | | Authors: | Schreiner, M, Schlesinger, R, Heberle, J, Niemann, H.H. | | Deposit date: | 2015-02-11 | | Release date: | 2015-04-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structure of Halorhodopsin from Halobacterium Salinarum in a New Crystal Form that Imposes Little Restraint on the E-F Loop.
J.Struct.Biol., 190, 2015
|
|
1TW2
 
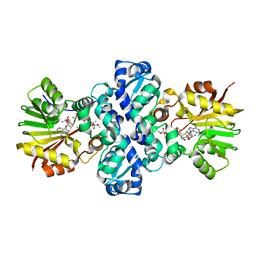 | | Crystal structure of Carminomycin-4-O-methyltransferase (DnrK) in complex with S-adenosyl-L-homocystein (SAH) and 4-methoxy-e-rhodomycin T (M-ET) | | Descriptor: | Carminomycin 4-O-methyltransferase, METHYL (4R)-2-ETHYL-2,5,12-TRIHYDROXY-7-METHOXY-6,11-DIOXO-4-{[2,3,6-TRIDEOXY-3-(DIMETHYLAMINO)-BETA-D-RIBO-HEXOPYRANOSYL]OXY}-1H,2H,3H,4H,6H,11H-TETRACENE-1-CARBOXYLATE, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Jansson, A, Koskiniemi, H, Mantsala, P, Niemi, J, Schneider, G, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2004-06-30 | | Release date: | 2004-09-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a ternary complex of DnrK, a methyltransferase in daunorubicin biosynthesis, with bound products
J.Biol.Chem., 279, 2004
|
|
1TW3
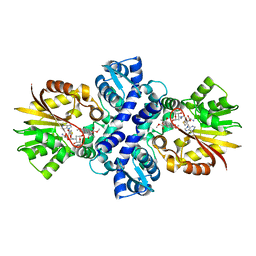 
 | | Crystal structure of Carminomycin-4-O-methyltransferase (DnrK) in complex with S-adenosyl-L-homocystein (SAH) and 4-methoxy-e-rhodomycin T (M-ET) | | Descriptor: | Carminomycin 4-O-methyltransferase, METHYL (4R)-2-ETHYL-2,5,12-TRIHYDROXY-7-METHOXY-6,11-DIOXO-4-{[2,3,6-TRIDEOXY-3-(DIMETHYLAMINO)-BETA-D-RIBO-HEXOPYRANOSYL]OXY}-1H,2H,3H,4H,6H,11H-TETRACENE-1-CARBOXYLATE, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Jansson, A, Koskiniemi, H, Mantsala, P, Niemi, J, Schneider, G. | | Deposit date: | 2004-06-30 | | Release date: | 2004-09-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of a ternary complex of DnrK, a methyltransferase in daunorubicin biosynthesis, with bound products
J.Biol.Chem., 279, 2004
|
|
8BBQ
 
 | | Determination of the structure of active tyrosinase from bacterium Verrucomicrobium spinosum | | Descriptor: | COPPER (II) ION, Core tyrosinase, GLYCEROL, ... | | Authors: | Fekry, M, Dave, K, Badgujar, D, Aurelius, O, Hamnevik, E, Dobritzsch, D, Danielson, H. | | Deposit date: | 2022-10-14 | | Release date: | 2023-09-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | The Crystal Structure of Tyrosinase from Verrucomicrobium spinosum Reveals It to Be an Atypical Bacterial Tyrosinase.
Biomolecules, 13, 2023
|
|
8BBR
 
 | | Determination of the structure of active tyrosinase from bacterium Verrucomicrobium spinosum | | Descriptor: | COPPER (II) ION, Core tyrosinase, SULFATE ION | | Authors: | Fekry, M, Dave, K, Badgujar, D, Aurelius, O, Hamnevik, E, Dobritzsch, D, Danielson, H. | | Deposit date: | 2022-10-14 | | Release date: | 2023-09-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | The Crystal Structure of Tyrosinase from Verrucomicrobium spinosum Reveals It to Be an Atypical Bacterial Tyrosinase.
Biomolecules, 13, 2023
|
|
6GCU
 
 | | MET receptor in complex with InlB internalin domain and DARPin A3A | | Descriptor: | DARPin A3A, Hepatocyte growth factor receptor, Internalin B | | Authors: | Meyer, T, Andres, F, Iamele, L, Gherardi, E, Pluckthun, A, Niemann, H.H. | | Deposit date: | 2018-04-19 | | Release date: | 2019-05-15 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (6.001 Å) | | Cite: | Inhibition of the MET Kinase Activity and Cell Growth in MET-Addicted Cancer Cells by Bi-Paratopic Linking.
J.Mol.Biol., 431, 2019
|
|
3IHG
 
 | | Crystal structure of a ternary complex of aklavinone-11 hydroxylase with FAD and aklavinone | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, RdmE, SULFATE ION, ... | | Authors: | Lindqvist, Y, Koskiniemi, H, Jansson, A, Sandalova, T, Schneider, G. | | Deposit date: | 2009-07-30 | | Release date: | 2009-09-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Structural basis for substrate recognition and specificity in aklavinone-11-hydroxylase from rhodomycin biosynthesis.
J.Mol.Biol., 393, 2009
|
|
2WQW
 
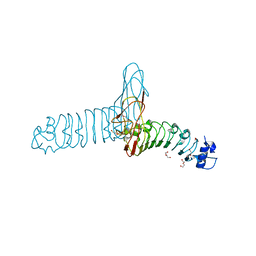 | |
2HYB
 
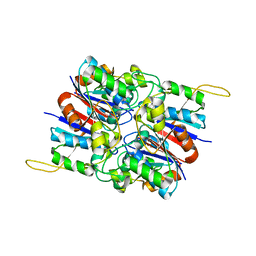 | | Crystal Structure of Hexameric DsrEFH | | Descriptor: | DsrH, Intracellular sulfur oxidation protein dsrF, Putative sulfurtransferase dsrE | | Authors: | Shin, D.H, Connie, H, Schulte, A, Dahl, C, Kim, R, Kim, S.H, Berkeley Structural Genomics Center (BSGC) | | Deposit date: | 2006-08-04 | | Release date: | 2007-07-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of hexameric DsrEFH
To be Published
|
|
3UVC
 
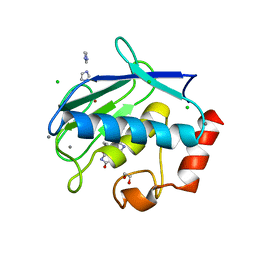 | | MMP12 in a complex with the dimeric adduct: 5-(5-phenylhydantoin)-5-phenylhydantoin | | Descriptor: | (4R,4'S)-4,4'-diphenyl-4,4'-biimidazolidine-2,2',5,5'-tetrone, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Derbyshire, D.J, Danielson, H, Nystrum, S. | | Deposit date: | 2011-11-29 | | Release date: | 2013-01-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Characterization of fragments interacting with MMP-12
To be Published
|
|
3ZCJ
 
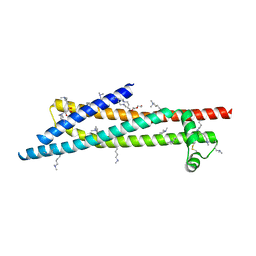 | |
3ZCI
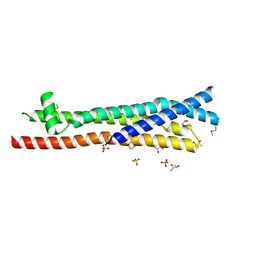 
 | |
2BSJ
 
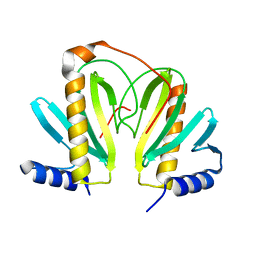 | | Native Crystal Structure of the Type III Secretion chaperone SycT from Yersinia enterocolitica | | Descriptor: | CHAPERONE PROTEIN SYCT, CHLORIDE ION | | Authors: | Buttner, C.R, Cornelis, G.R, Heinz, D.W, Niemann, H.H. | | Deposit date: | 2005-05-21 | | Release date: | 2005-09-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Crystal Structure of Yersinia Enterocolitica Type III Secretion Chaperone Syct.
Protein Sci., 14, 2005
|
|
2BSH
 
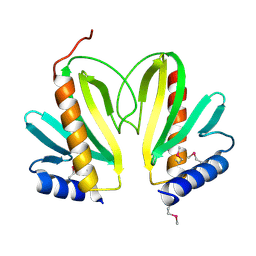 | |
4AM9
 
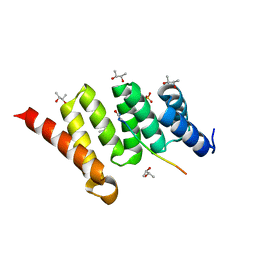 | | CRYSTAL STRUCTURE OF THE YERSINIA ENTEROCOLITICA TYPE III SECRETION CHAPERONE SYCD IN COMPLEX WITH A PEPTIDE OF THE TRANSLOCATOR YOPD | | Descriptor: | (R,R)-2,3-BUTANEDIOL, CHAPERONE SYCD, SULFATE ION, ... | | Authors: | Schreiner, M, Niemann, H.H. | | Deposit date: | 2012-03-08 | | Release date: | 2012-05-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of the Yersinia Enterocolitica Type III Secretion Chaperone Sycd in Complex with a Peptide of the Minor Translocator Yopd
Bmc Struct.Biol., 12, 2012
|
|
2BSI
 
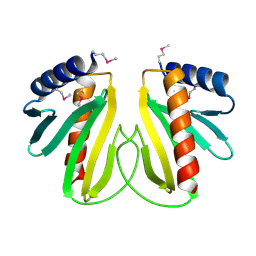 | |
2Y6V
 
 | |
2Y5P
 
 | |
