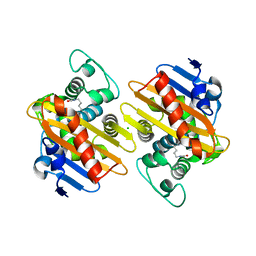
6GOA
 
 | |
5ODZ
 
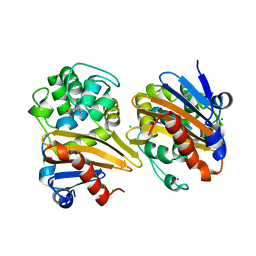 | | CRYSTAL STRUCTURE OF THE BETA-LACTAMASE OXA-163 | | Descriptor: | Beta-lactamase, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Lund, B.A, Carlsen, T.J.O, Leiros, H.K.S. | | Deposit date: | 2017-07-07 | | Release date: | 2017-10-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Structure, activity and thermostability investigations of OXA-163, OXA-181 and OXA-245 using biochemical analysis, crystal structures and differential scanning calorimetry analysis.
Acta Crystallogr F Struct Biol Commun, 73, 2017
|
|
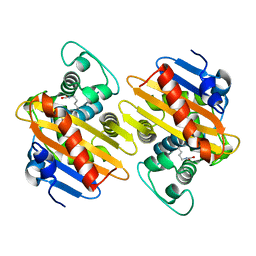
5OFT
 
 | |
7QLB
 
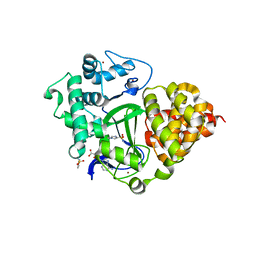 | | SMYD3 in complex with fragment FL06268 | | Descriptor: | 1-methylimidazole-4-sulfonamide, Histone-lysine N-methyltransferase SMYD3, S-ADENOSYLMETHIONINE, ... | | Authors: | Lund, B.A, Cederfelt, D, Dobritzsch, D. | | Deposit date: | 2021-12-20 | | Release date: | 2023-03-29 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification of fragments targeting SMYD3 using highly sensitive kinetic and multiplexed biosensor-based screening
Rsc Med Chem, 2024
|
|
7QNR
 
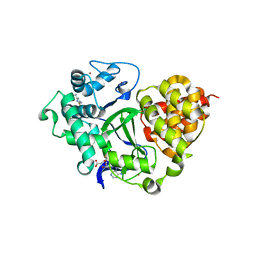 | | SMYD3 in complex with fragment FL01791 | | Descriptor: | 3-propan-2-yl-1,2,4-thiadiazol-5-amine, Histone-lysine N-methyltransferase SMYD3, S-ADENOSYLMETHIONINE, ... | | Authors: | Lund, B.A, Cederfelt, D, Dobritzsch, D. | | Deposit date: | 2021-12-22 | | Release date: | 2023-04-05 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Identification of fragments targeting SMYD3 using highly sensitive kinetic and multiplexed biosensor-based screening
Rsc Med Chem, 2024
|
|
7QNU
 
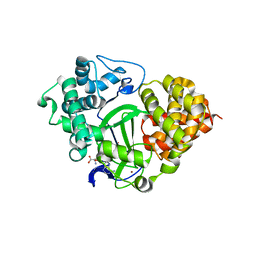 | | SMYD3 in complex with fragment FL08619 | | Descriptor: | BENZOYL-FORMIC ACID, Histone-lysine N-methyltransferase SMYD3, S-ADENOSYLMETHIONINE, ... | | Authors: | Lund, B.A, Cederfelt, D, Dobritzsch, D. | | Deposit date: | 2021-12-22 | | Release date: | 2023-04-05 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Identification of fragments targeting SMYD3 using highly sensitive kinetic and multiplexed biosensor-based screening
Rsc Med Chem, 2024
|
|
7R1K
 
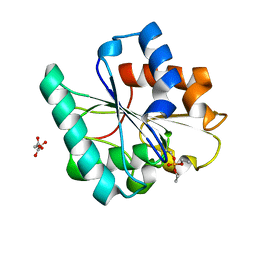 | | Phosphorylated Bacillus pumilus Lipase A | | Descriptor: | DIETHYL PHOSPHONATE, Lipase, OXALOACETATE ION | | Authors: | Lund, B.A. | | Deposit date: | 2022-02-03 | | Release date: | 2022-05-25 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure and Mechanism of a Cold-Adapted Bacterial Lipase
Biochemistry, 61, 2022
|
|
7R25
 
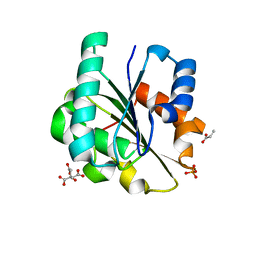 | | Bacillus pumilus Lipase A | | Descriptor: | CITRIC ACID, Lipase, PHOSPHATE ION, ... | | Authors: | Lund, B.A. | | Deposit date: | 2022-02-04 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (0.87 Å) | | Cite: | Structure and Mechanism of a Cold-Adapted Bacterial Lipase
Biochemistry, 61, 2022
|
|
7R4B
 
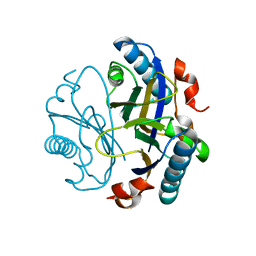 | | The Bacillus pumilus chorismate mutase | | Descriptor: | 1,2-ETHANEDIOL, Chorismate mutase AroH | | Authors: | Lund, B.A. | | Deposit date: | 2022-02-08 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | The Bacillus pumilus chorismate mutase
To Be Published
|
|
4Z84
 
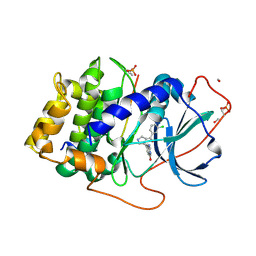 | | PKAB3 in complex with pyrrolidine inhibitor 34a | | Descriptor: | 7-[(3S,4R)-4-(3-chlorophenyl)carbonylpyrrolidin-3-yl]-3H-quinazolin-4-one, METHANOL, cAMP-dependent protein kinase catalytic subunit alpha, ... | | Authors: | Lund, B.A, Alam, K.A, Engh, R.A. | | Deposit date: | 2015-04-08 | | Release date: | 2015-12-02 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.554 Å) | | Cite: | Addressing the Glycine-Rich Loop of Protein Kinases by a Multi-Facetted Interaction Network: Inhibition of PKA and a PKB Mimic.
Chemistry, 22, 2016
|
|
4Z83
 
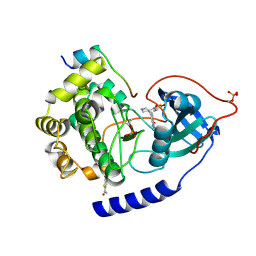 | | PKAB3 in complex with pyrrolidine inhibitor 47a | | Descriptor: | 7-{(3S,4R)-4-[(5-bromothiophen-2-yl)carbonyl]pyrrolidin-3-yl}quinazolin-4(3H)-one, cAMP-dependent protein kinase catalytic subunit alpha, cAMP-dependent protein kinase inhibitor alpha | | Authors: | Lund, B.A, Alam, K.A, Engh, R.A. | | Deposit date: | 2015-04-08 | | Release date: | 2015-12-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Addressing the Glycine-Rich Loop of Protein Kinases by a Multi-Facetted Interaction Network: Inhibition of PKA and a PKB Mimic.
Chemistry, 22, 2016
|
|
5OE2
 
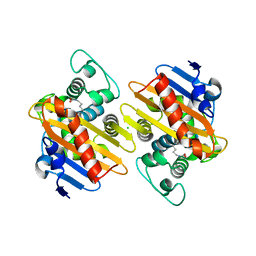 | | CRYSTAL STRUCTURE OF THE BETA-LACTAMASE OXA-245 | | Descriptor: | Beta-lactamase, CHLORIDE ION, SODIUM ION | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-07 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure, activity and thermostability investigations of OXA-163, OXA-181 and OXA-245 using biochemical analysis, crystal structures and differential scanning calorimetry analysis.
Acta Crystallogr F Struct Biol Commun, 73, 2017
|
|
5OE0
 
 | | CRYSTAL STRUCTURE OF THE BETA-LACTAMASE OXA-181 | | Descriptor: | Beta-lactamase, CHLORIDE ION, SULFATE ION | | Authors: | Lund, B.A, Carlsen, T.J.O, Leiros, H.K.S, Thomassen, A.M. | | Deposit date: | 2017-07-07 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.0500083 Å) | | Cite: | Structure, activity and thermostability investigations of OXA-163, OXA-181 and OXA-245 using biochemical analysis, crystal structures and differential scanning calorimetry analysis.
Acta Crystallogr F Struct Biol Commun, 73, 2017
|
|
8OWO
 
 | | SMYD3 in complex with fragment FL01507 | | Descriptor: | 3-oxidanylbenzenecarbonitrile, GLYCEROL, Histone-lysine N-methyltransferase SMYD3, ... | | Authors: | Lund, B.A, Cederfelt, D, Dobritzsch, D. | | Deposit date: | 2023-04-28 | | Release date: | 2023-08-30 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification of fragments targeting SMYD3 using highly sensitive kinetic and multiplexed biosensor-based screening.
Rsc Med Chem, 15, 2024
|
|
7ODA
 
 | | OXA-48-like Beta-lactamase OXA-436 | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactamase, CHLORIDE ION | | Authors: | Lund, B.A, Thomassen, A.M, Carlsen, T.J.W, Leiros, H.K.S. | | Deposit date: | 2021-04-29 | | Release date: | 2021-09-08 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.796 Å) | | Cite: | Biochemical and biophysical characterization of the OXA-48-like carbapenemase OXA-436.
Acta Crystallogr.,Sect.F, 77, 2021
|
|
5DVA
 
 | |
5DTK
 
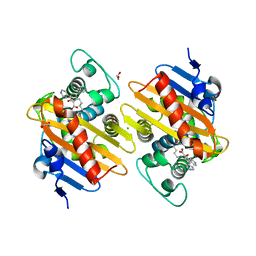 | | Fragments bound to the OXA-48 beta-lactamase: Compound 17 | | Descriptor: | 1,2-ETHANEDIOL, 3,5-di(pyridin-4-yl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Christopeit, T, Leiros, H.-K.S. | | Deposit date: | 2015-09-18 | | Release date: | 2016-05-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.60000241 Å) | | Cite: | Screening and Design of Inhibitor Scaffolds for the Antibiotic Resistance Oxacillinase-48 (OXA-48) through Surface Plasmon Resonance Screening.
J.Med.Chem., 59, 2016
|
|
5DTT
 
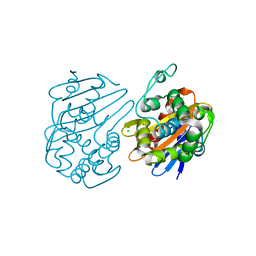 | | Fragments bound to the OXA-48 beta-lactamase: Compound 3 | | Descriptor: | 1,2-ETHANEDIOL, 3-(1,3-thiazol-2-yl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Christopeit, T, Leiros, H.-K.S. | | Deposit date: | 2015-09-18 | | Release date: | 2016-05-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.10000539 Å) | | Cite: | Screening and Design of Inhibitor Scaffolds for the Antibiotic Resistance Oxacillinase-48 (OXA-48) through Surface Plasmon Resonance Screening.
J.Med.Chem., 59, 2016
|
|
5DTS
 
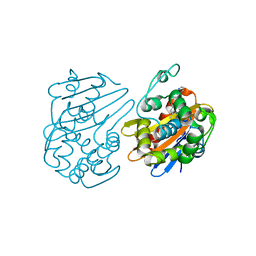 | |
2WHZ
 
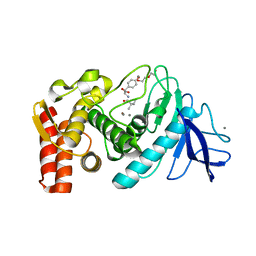 | |
2WI0
 
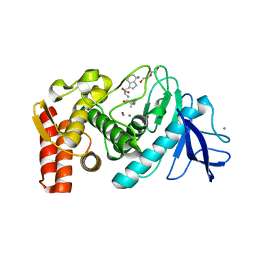 | |
5QAG
 
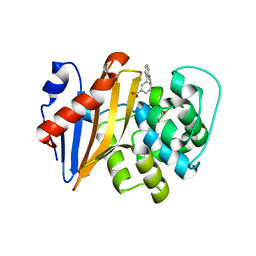 | | OXA-48 IN COMPLEX WITH COMPOUND 9a | | Descriptor: | 1,2-ETHANEDIOL, 3-(2-methoxycarbonylphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAW
 
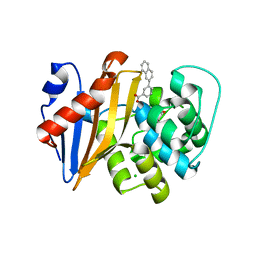 | | OXA-48 IN COMPLEX WITH COMPOUND 27 | | Descriptor: | 3-naphthalen-2-ylbenzoic acid, Beta-lactamase, CHLORIDE ION | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAI
 
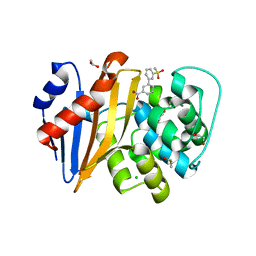 | | OXA-48 IN COMPLEX WITH COMPOUND 12a | | Descriptor: | 1,2-ETHANEDIOL, 3-(3-methylsulfonylphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAX
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 28 | | Descriptor: | 1,2-ETHANEDIOL, 3-quinolin-6-ylbenzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
