4PVZ
 
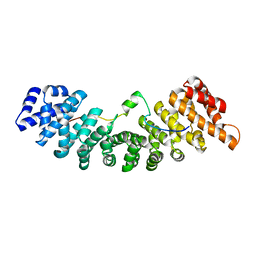 | | Structure of yeast importin a bound to the membrane protein Nuclear Localization Signal sequence of INM protein Heh2 | | Descriptor: | Importin subunit alpha, Inner nuclear membrane protein HEH2 | | Authors: | Lokareddy, R.K, Hapsari, A.R, van Rheenen, M, Bhardwaj, A, Veenhoff, L.M, Cingolani, C. | | Deposit date: | 2014-03-18 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Distinctive Properties of the Nuclear Localization Signals of Inner Nuclear Membrane Proteins Heh1 and Heh2.
Structure, 23, 2015
|
|
4HRF
 
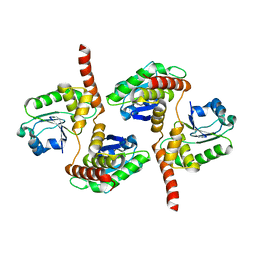 | | Atomic structure of DUSP26 | | Descriptor: | Dual specificity protein phosphatase 26 | | Authors: | Lokareddy, R.K, Bhardwaj, A, Cingolani, G. | | Deposit date: | 2012-10-27 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Atomic structure of dual-specificity phosphatase 26, a novel p53 phosphatase.
Biochemistry, 52, 2013
|
|
4XZR
 
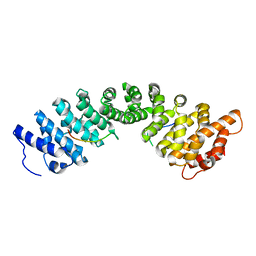 | |
4YI0
 
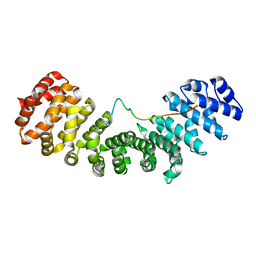 | | Structure of mouse importin a1 bound to Pom121NLS | | Descriptor: | Importin subunit alpha-1, Nuclear envelope pore membrane protein POM 121 | | Authors: | Lokareddy, R.K, Cingolani, G. | | Deposit date: | 2015-02-27 | | Release date: | 2016-01-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.806 Å) | | Cite: | Mammalian Pom121 and yeast Heh2 share IBB-like NLSs that support targeting to the inner nuclear membrane
To Be Published
|
|
3GZ2
 
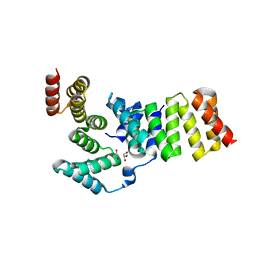 | | Crystal structure of IpgC in complex with an IpaB peptide | | Descriptor: | Chaperone protein ipgC, GLYCEROL, IMIDAZOLE, ... | | Authors: | Lokareddy, R.K, Lunelli, M, Kolbe, M. | | Deposit date: | 2009-04-06 | | Release date: | 2010-04-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Combination of two separate binding domains defines stoichiometry between type III secretion system chaperone IpgC and translocator protein IpaB
J.Biol.Chem., 285, 2010
|
|
5JJ3
 
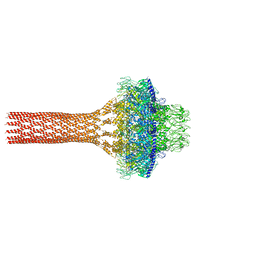 | |
5JJ1
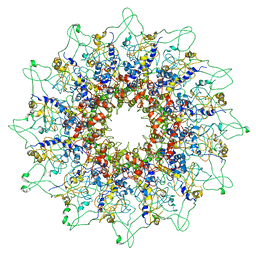 
 | |
7UXE
 
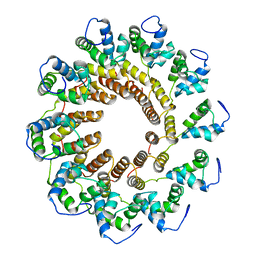 | | Pseudomonas phage E217 small terminase (TerS) | | Descriptor: | Small terminase | | Authors: | Lokareddy, R.K, Hou, C.-F.D, Doll, S.G, Li, F, Gillilan, R, Forti, F, Briani, F, Cingolani, G. | | Deposit date: | 2022-05-05 | | Release date: | 2022-09-28 | | Method: | ELECTRON MICROSCOPY (3.38 Å) | | Cite: | Terminase Subunits from the Pseudomonas-Phage E217.
J.Mol.Biol., 434, 2022
|
|
8DKR
 
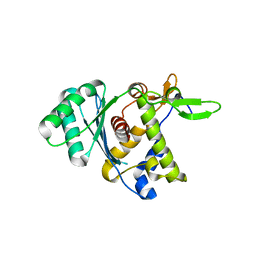 | |
7N9H
 
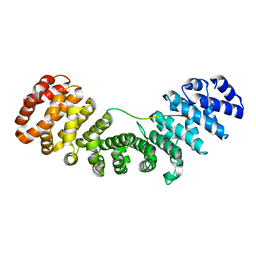 | |
6RGV
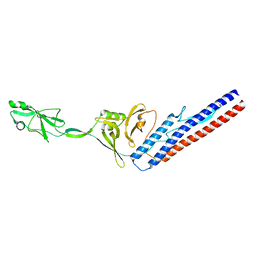 
 | |
6E3B
 
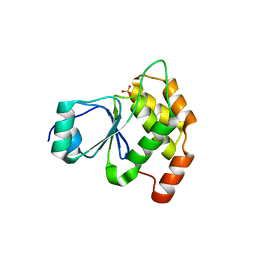 | |
6VI1
 
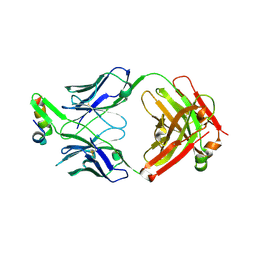 | | Structure of Fab4 bound to P22 TerL(1-33) | | Descriptor: | Synthetic Fab4 heavy chain, Synthetic Fab4 light chain, Terminase, ... | | Authors: | Cingolani, G, Lokareddy, R, Ko, Y. | | Deposit date: | 2020-01-11 | | Release date: | 2020-09-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Recognition of an alpha-helical hairpin in P22 large terminase by a synthetic antibody fragment.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
6W7T
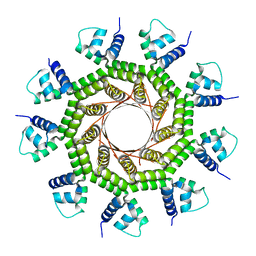 
 | | Structure of PaP3 small terminase | | Descriptor: | small terminase subunit | | Authors: | Cingolani, G, Lokareddy, R. | | Deposit date: | 2020-03-19 | | Release date: | 2020-11-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Biophysical analysis of Pseudomonas-phage PaP3 small terminase suggests a mechanism for sequence-specific DNA-binding by lateral interdigitation.
Nucleic Acids Res., 48, 2020
|
|
4NYH
 
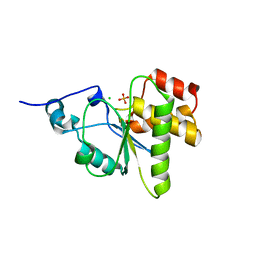 | | Orthorhombic crystal form of pir1 dual specificity phosphatase core | | Descriptor: | CHLORIDE ION, PHOSPHATE ION, RNA/RNP complex-1-interacting phosphatase | | Authors: | Sankhala, R.S, Lokareddy, R.K, Cingolani, G. | | Deposit date: | 2013-12-10 | | Release date: | 2014-01-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structure of Human PIR1, an Atypical Dual-Specificity Phosphatase.
Biochemistry, 53, 2014
|
|
6VI2
 
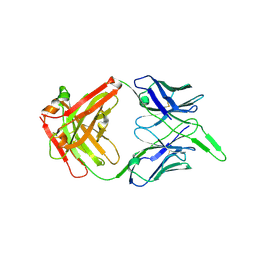 | | Structure of the unaligned Fab4 | | Descriptor: | FAB4 heavy chain, FAB4 light chain | | Authors: | Cingolani, G, Lokareddy, R, Ko, Y. | | Deposit date: | 2020-01-11 | | Release date: | 2020-09-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Recognition of an alpha-helical hairpin in P22 large terminase by a synthetic antibody fragment.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
6XMI
 
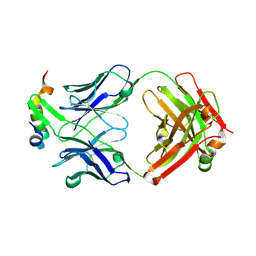 | | Structure of Fab4 bound to P22 TerL(1-33) | | Descriptor: | Fab Heavy chain, Fab Light chain, Terminase, ... | | Authors: | Cingolani, G, Lokareddy, R, Ko, Y. | | Deposit date: | 2020-06-30 | | Release date: | 2020-08-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Recognition of an alpha-helical hairpin in P22 large terminase by a synthetic antibody fragment.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
3GYZ
 
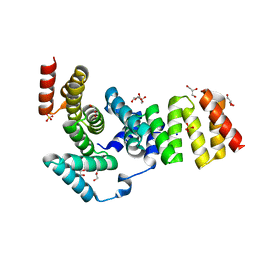 | | Crystal structure of IpgC from Shigella flexneri | | Descriptor: | Chaperone protein ipgC, GLYCEROL, SODIUM ION, ... | | Authors: | Lunelli, M, Lokareddy, R.K, Zychlinsky, A, Kolbe, M. | | Deposit date: | 2009-04-06 | | Release date: | 2009-06-16 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | IpaB-IpgC interaction defines binding motif for type III secretion translocator
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3GZ1
 
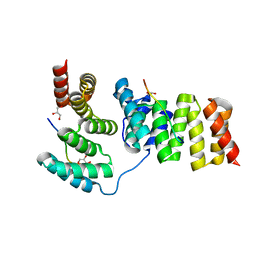 | | Crystal structure of IpgC in complex with the chaperone binding region of IpaB | | Descriptor: | Chaperone protein ipgC, GLYCEROL, Invasin ipaB | | Authors: | Lunelli, M, Lokareddy, R.K, Zychlinsky, A, Kolbe, M. | | Deposit date: | 2009-04-06 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | IpaB-IpgC interaction defines binding motif for type III secretion translocator
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
7JOQ
 
 | | Structure of NV1 small terminase | | Descriptor: | Small Terminase subunit | | Authors: | Cingolani, G, Lokareddy, R. | | Deposit date: | 2020-08-07 | | Release date: | 2020-11-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.95 Å) | | Cite: | Biophysical analysis of Pseudomonas-phage PaP3 small terminase suggests a mechanism for sequence-specific DNA-binding by lateral interdigitation.
Nucleic Acids Res., 48, 2020
|
|
5HUY
 
 | |
5HUW
 
 | |
5T94
 
 | | Crystal structure of Kap60 bound to yeast RCC1 (Prp20) | | Descriptor: | Guanine nucleotide exchange factor SRM1, Importin subunit alpha | | Authors: | Sankhala, R.S, Lokareddy, R.K, Pumroy, R.A, Cingolani, G. | | Deposit date: | 2016-09-09 | | Release date: | 2017-09-13 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.631 Å) | | Cite: | Three-dimensional context rather than NLS amino acid sequence determines importin alpha subtype specificity for RCC1.
Nat Commun, 8, 2017
|
|
4MBB
 
 | | Cubic crystal form of PIR1 dual specificity phosphatase core | | Descriptor: | CHLORIDE ION, PHOSPHATE ION, RNA/RNP complex-1-interacting phosphatase | | Authors: | Sankhala, R.S, Lokareddy, R.K, Cingolani, G. | | Deposit date: | 2013-08-19 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.849 Å) | | Cite: | Structure of Human PIR1, an Atypical Dual-Specificity Phosphatase.
Biochemistry, 53, 2014
|
|
5TBK
 
 | | Crystal structure of human importin a3 bound to RCC1 | | Descriptor: | Importin subunit alpha-3, Regulator of chromosome condensation | | Authors: | Sankhala, R.S, Lokareddy, R.K, Pumroy, R.A, Cingolani, G. | | Deposit date: | 2016-09-12 | | Release date: | 2017-09-13 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.45 Å) | | Cite: | Three-dimensional context rather than NLS amino acid sequence determines importin alpha subtype specificity for RCC1.
Nat Commun, 8, 2017
|
|
