2KHT
 | | NMR Structure of human alpha defensin HNP-1 | | 分子名称: | Neutrophil defensin 1 | | 著者 | Zhang, Y, Li, S, Doherty, T.F, Lubkowski, J, Lu, W, Li, J, Barinka, C, Hong, M. | | 登録日 | 2009-04-11 | | 公開日 | 2010-02-09 | | 最終更新日 | 2024-05-01 | | 実験手法 | SOLID-STATE NMR | | 主引用文献 | Resonance assignment and three-dimensional structure determination of a human alpha-defensin, HNP-1, by solid-state NMR.
J.Mol.Biol., 397, 2010
|
|
4X83
 
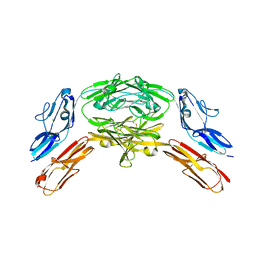 | | Crystal structure of Dscam1 isoform 7.44, N-terminal four Ig domains | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | 著者 | Chen, Q, Yu, Y, Li, S.A, Cheng, L. | | 登録日 | 2014-12-10 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.902 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
4X9B
 
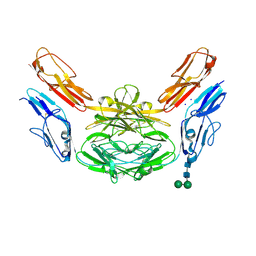 | | Crystal structure of Dscam1 isoform 4.44, N-terminal four Ig domains | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Down syndrome cell adhesion molecule, isoform 4.44, ... | | 著者 | Chen, Q, Yu, Y, Li, S.A, Cheng, L. | | 登録日 | 2014-12-11 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
4X9G
 | | Crystal structure of Dscam1 isoform 6.44, N-terminal four Ig domains | | 分子名称: | Down Syndrome Cell Adhesion Molecule isoform 6.44, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | 著者 | Chen, Q, Yu, Y, Li, S.A, Cheng, L. | | 登録日 | 2014-12-11 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (3.403 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
4X9I
 | | Crystal structure of Dscam1 isoform 9.44, N-terminal four Ig domains | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Down Syndrome Cell Adhesion Molecule, isoform 9.44, ... | | 著者 | Chen, Q, Yu, Y, Li, S.A, cheng, L. | | 登録日 | 2014-12-11 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (2.904 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
4XB7
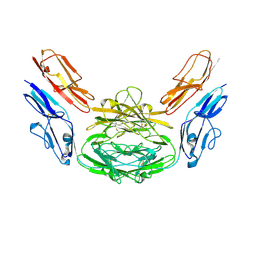 
 | | Crystal structure of Dscam1 isoform 4.4, N-terminal four Ig domains | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Down syndrome cell adhesion molecule, isoform 4.4, ... | | 著者 | Chen, Q, Yu, Y, Li, S.A, Cheng, L. | | 登録日 | 2014-12-16 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (4.004 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
8K58
 
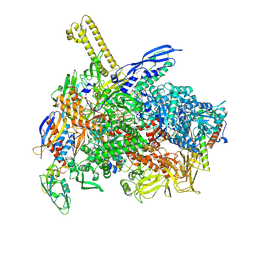 | | The cryo-EM map of close TIEA-TEC complex | | 分子名称: | 15 kDa RNA polymerase-binding protein, DNA (29-MER), DNA-directed RNA polymerase subunit alpha, ... | | 著者 | Zhang, K.N, Liu, Y, Chen, M, Wang, Y, Lin, W, Li, M, Zhang, X, Gao, Y, Gong, Q, Chen, H, Steve, M, Li, S, Zhang, K, Liu, B. | | 登録日 | 2023-07-21 | | 公開日 | 2024-07-24 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3.15 Å) | | 主引用文献 | TIEA inhibits Sigma70-dependent transcriptions, accelerates elongation speed and elevates transcription error
To Be Published
|
|
8K5A
 
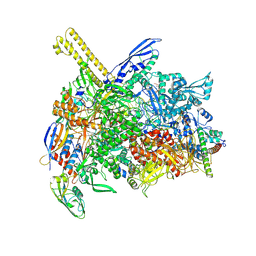 | | The cryo-EM map of open TIEA-TEC complex | | 分子名称: | 15 kDa RNA polymerase-binding protein, DNA (29-MER), DNA-directed RNA polymerase subunit alpha, ... | | 著者 | Zhang, K.N, Liu, Y, Chen, M, Wang, Y, Lin, W, Li, M, Zhang, X, Gao, Y, Gong, Q, Chen, H, Steve, M, Li, S, Zhang, K, Liu, B. | | 登録日 | 2023-07-21 | | 公開日 | 2024-07-24 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | TIEA inhibits sigma70-dependent transcriptions, accelerates elongation speed and elevates transcription error
To Be Published
|
|
8K0E
 
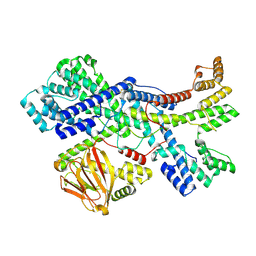 | | Human collagen prolyl processing enzyme complex, P3H1/CRTAP heterodimer | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Cartilage-associated protein, FE (III) ION, ... | | 著者 | Li, W, Peng, J, Yao, D, Rao, B, Xia, Y, Wang, Q, Li, S, Cao, M, Shen, Y, Ma, P, Liao, R, Qin, A, Zhao, J, Cao, Y. | | 登録日 | 2023-07-08 | | 公開日 | 2024-09-18 | | 最終更新日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (3.65 Å) | | 主引用文献 | The structural basis for the collagen processing by human P3H1/CRTAP/PPIB ternary complex.
Nat Commun, 15, 2024
|
|
8K59
 
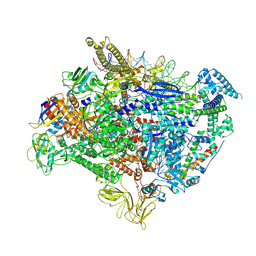 | | The cryo-EM map of TIC-TIEA complex | | 分子名称: | 15 kDa RNA polymerase-binding protein, DNA (61-MER), DNA (63-MER), ... | | 著者 | Zhang, K.N, Liu, Y, Chen, M, Wang, Y, Lin, W, Li, M, Zhang, X, Gao, Y, Gong, Q, Chen, H, Steve, M, Li, S, Zhang, K, Liu, B. | | 登録日 | 2023-07-21 | | 公開日 | 2024-11-13 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | TIEA inhibits sigma70-dependent transcriptions, accelerates elongation speed and elevates transcription error
To Be Published
|
|
6UES
 
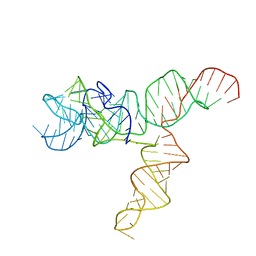 | | Apo SAM-IV Riboswitch | | 分子名称: | RNA (119-MER) | | 著者 | Zhang, K, Li, S, Kappel, K, Pintilie, G, Su, Z, Mou, T, Schmid, M, Das, R, Chiu, W. | | 登録日 | 2019-09-23 | | 公開日 | 2019-12-18 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Cryo-EM structure of a 40 kDa SAM-IV riboswitch RNA at 3.7 angstrom resolution.
Nat Commun, 10, 2019
|
|
4XB8
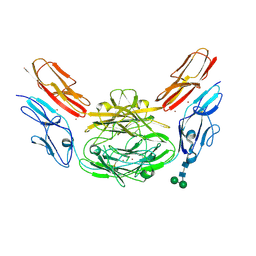 
 | | Crystal structure of Dscam1 isoform 9.44, N-terminal four Ig domains (with zinc) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Down Syndrome Cell Adhesion Molecule, ... | | 著者 | Chen, Q, Yu, Y, Li, S.A, cheng, L. | | 登録日 | 2014-12-16 | | 公開日 | 2015-12-16 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (3.202 Å) | | 主引用文献 | Structural basis of Dscam1 homodimerization: Insights into context constraint for protein recognition
Sci Adv, 2, 2016
|
|
3SLI
 
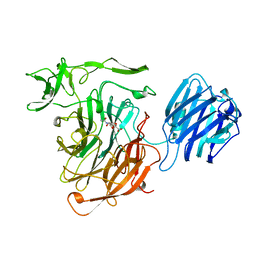 | | LEECH INTRAMOLECULAR TRANS-SIALIDASE COMPLEXED WITH 2,7-ANHYDRO-NEU5AC PREPARED BY SOAKING WITH 3'-SIALYLLACTOSE | | 分子名称: | 2-ACETYLAMINO-7-(1,2-DIHYDROXY-ETHYL)-3-HYDROXY-6,8-DIOXA-BICYCLO[3.2.1]OCTANE-5-CARBOXYLIC ACID, INTRAMOLECULAR TRANS-SIALIDASE | | 著者 | Luo, Y, Li, S.C, Li, Y.T, Luo, M. | | 登録日 | 1998-10-03 | | 公開日 | 1999-04-27 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | The 1.8 A structures of leech intramolecular trans-sialidase complexes: evidence of its enzymatic mechanism.
J.Mol.Biol., 285, 1999
|
|
6F2Z
 
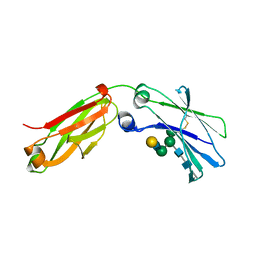 | | Structure of a Fc mutant | | 分子名称: | Fc fragment of antibody, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | 著者 | Zhou, A, Li, S. | | 登録日 | 2017-11-27 | | 公開日 | 2018-12-12 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structure of Fc mutant in monomeric form
To Be Published
|
|
7R7S
 
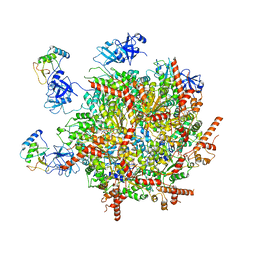 | | p47-bound p97-R155H mutant with ATPgammaS | | 分子名称: | NSFL1 cofactor p47, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Transitional endoplasmic reticulum ATPase | | 著者 | Nandi, P, Li, S, Coulmbres, R.C.A, Wang, F, Williams, D.R, Malyutin, A.G, Poh, Y.-P, Chou, T.-F, Chiu, P.-L. | | 登録日 | 2021-06-25 | | 公開日 | 2021-08-04 | | 最終更新日 | 2024-06-05 | | 実験手法 | ELECTRON MICROSCOPY (4.23 Å) | | 主引用文献 | Structural and Functional Analysis of Disease-Linked p97 ATPase Mutant Complexes.
Int J Mol Sci, 22, 2021
|
|
6J7V
 
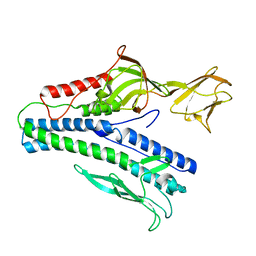 | |
5GZO
 
 | | Structure of neutralizing antibody bound to Zika envelope protein | | 分子名称: | Antibody heavy chain, Antibody light chain, Genome polyprotein | | 著者 | Wang, Q, Yang, H, Liu, X, Dai, L, Ma, T, Qi, J, Wong, G, Peng, R, Liu, S, Li, J, Li, S, Song, J, Liu, J, He, J, Yuan, H, Xiong, Y, Liao, Y, Li, J, Yang, J, Tong, Z, Griffin, B, Bi, Y, Liang, M, Xu, X, Cheng, G, Wang, P, Qiu, X, Kobinger, G, Shi, Y, Yan, J, Gao, G.F. | | 登録日 | 2016-09-29 | | 公開日 | 2017-01-18 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (2.755 Å) | | 主引用文献 | Molecular determinants of human neutralizing antibodies isolated from a patient infected with Zika virus
Sci Transl Med, 8, 2016
|
|
1T7S
 
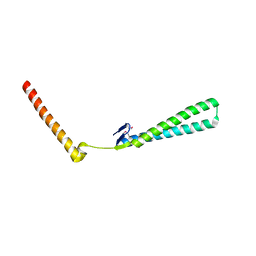 | | Structural Genomics of Caenorhabditis elegans: Structure of BAG-1 protein | | 分子名称: | BAG-1 cochaperone | | 著者 | Symersky, J, Zhang, Y, Schormann, N, Li, S, Bunzel, R, Pruett, P, Luan, C.-H, Luo, M, Southeast Collaboratory for Structural Genomics (SECSG) | | 登録日 | 2004-05-10 | | 公開日 | 2004-05-18 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structural genomics of Caenorhabditis elegans: structure of the BAG domain.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
5K81
 
 | | Crystal Structure of a Primate APOBEC3G N-Terminal Domain | | 分子名称: | Apolipoprotein B mRNA editing enzyme, catalytic peptide-like 3G,Apolipoprotein B mRNA editing enzyme, catalytic peptide-like 3G, ... | | 著者 | Xiao, X, Li, S.-X, Yang, H, Chen, X.S. | | 登録日 | 2016-05-27 | | 公開日 | 2016-08-10 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (2.006 Å) | | 主引用文献 | Crystal structures of APOBEC3G N-domain alone and its complex with DNA.
Nat Commun, 7, 2016
|
|
5K82
 
 | | Crystal Structure of a Primate APOBEC3G N-Terminal Domain | | 分子名称: | Apolipoprotein B mRNA editing enzyme, catalytic peptide-like 3G,Apolipoprotein B mRNA editing enzyme, catalytic peptide-like 3G, ... | | 著者 | Xiao, X, Li, S.-X, Yang, H, Chen, X.S. | | 登録日 | 2016-05-27 | | 公開日 | 2016-08-10 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (2.913 Å) | | 主引用文献 | Crystal structures of APOBEC3G N-domain alone and its complex with DNA.
Nat Commun, 7, 2016
|
|
6NUD
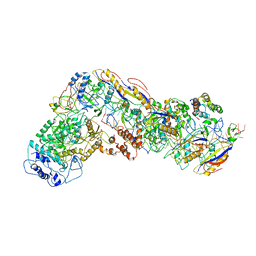 
 | | Small conformation of ssRNA-bound CRISPR_Csm complex | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CRISPR system Cms protein Csm2, CRISPR system single-strand-specific deoxyribonuclease Cas10/Csm1 (subtype III-A), ... | | 著者 | Zhang, K, Pintilie, G, Li, S, Zhu, Y, Chiu, W, Huang, Z. | | 登録日 | 2019-01-31 | | 公開日 | 2019-03-13 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Coupling of ssRNA cleavage with DNase activity in type III-A CRISPR-Csm revealed by cryo-EM and biochemistry.
Cell Res., 29, 2019
|
|
8GQP
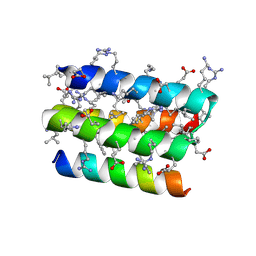 
 | | Complex of D-protein binder D-19437 and L-target L-Pep-1 | | 分子名称: | D-binder, L-pep1 | | 著者 | Liang, M.F, Li, S.C, Wang, T.Y, Liu, L, Lu, P.L. | | 登録日 | 2022-08-30 | | 公開日 | 2023-09-13 | | 最終更新日 | 2024-11-20 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Accurate de novo design of heterochiral protein-protein interactions
Cell Res., 2024
|
|
6NUE
 
 | | Small conformation of apo CRISPR_Csm complex | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CRISPR system Cms protein Csm2, CRISPR system single-strand-specific deoxyribonuclease Cas10/Csm1 (subtype III-A), ... | | 著者 | Zhang, K, Pintilie, G, Li, S, Zhu, Y, Chiu, W, Huang, Z. | | 登録日 | 2019-01-31 | | 公開日 | 2019-03-13 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | Coupling of ssRNA cleavage with DNase activity in type III-A CRISPR-Csm revealed by cryo-EM and biochemistry.
Cell Res., 29, 2019
|
|
2NNZ
 
 | | Solution structure of the hypothetical protein AF2241 from Archaeoglobus fulgidus | | 分子名称: | Hypothetical protein | | 著者 | Ai, X, Semesib, A, Yee, A, Arrowsmith, C.H, Li, S.S.C, Choy, W.Y, Ontario Centre for Structural Proteomics (OCSP) | | 登録日 | 2006-10-24 | | 公開日 | 2007-08-28 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Hypothetical protein AF2241 from Archaeoglobus fulgidus adopts a cyclophilin-like fold.
J.Biomol.Nmr, 38, 2007
|
|
1LXK
 
 | | Streptococcus pneumoniae Hyaluronate Lyase in Complex with Tetrasaccharide Hyaluronan Substrate | | 分子名称: | Hyaluronate Lyase, beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose | | 著者 | Jedrzejas, M.J, Mello, L.V, De Groot, B.L, Li, S. | | 登録日 | 2002-06-05 | | 公開日 | 2002-08-07 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (1.53 Å) | | 主引用文献 | Mechanism of hyaluronan degradation by Streptococcus pneumoniae hyaluronate lyase. Structures of complexes with the substrate.
J.Biol.Chem., 277, 2002
|
|
