4LFU
 
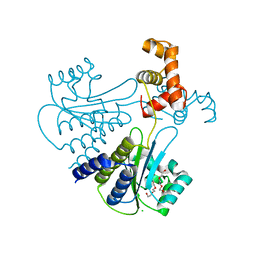 | | Crystal structure of Escherichia coli SdiA in the space group C2 | | Descriptor: | CHLORIDE ION, Regulatory protein SdiA, TETRAETHYLENE GLYCOL | | Authors: | Kim, T, Duong, T, Wu, C.A, Choi, J, Lan, N, Kang, S.W, Lokanath, N.K, Shin, D, Hwang, H.Y, Kim, K.K. | | Deposit date: | 2013-06-27 | | Release date: | 2014-03-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structural insights into the molecular mechanism of Escherichia coli SdiA, a quorum-sensing receptor
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4LGW
 
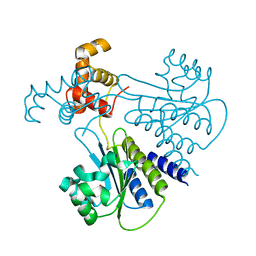 | | Crystal structure of Escherichia coli SdiA in the space group P6522 | | Descriptor: | GLYCEROL, Regulatory protein SdiA | | Authors: | Kim, T, Duong, T, Wu, C.A, Choi, J, Lan, N, Kang, S.W, Lokanath, N.K, Shin, D, Hwang, H.Y, Kim, K.K. | | Deposit date: | 2013-06-28 | | Release date: | 2014-03-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural insights into the molecular mechanism of Escherichia coli SdiA, a quorum-sensing receptor
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4PKN
 
 | | Crystal structure of the football-shaped GroEL-GroES2-(ADPBeFx)14 complex containing substrate Rubisco | | Descriptor: | 10 kDa chaperonin, 60 kDa chaperonin, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Fei, X, Ye, X, Laronde-Leblanc, N, Lorimer, G.H. | | Deposit date: | 2014-05-15 | | Release date: | 2014-08-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.66 Å) | | Cite: | Formation and structures of GroEL:GroES2 chaperonin footballs, the protein-folding functional form.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
2BT9
 
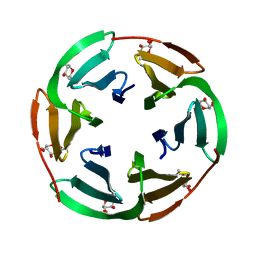 | | Lectin from Ralstonia solanacearum complexed with Me-fucoside | | Descriptor: | LECTIN, methyl alpha-L-fucopyranoside | | Authors: | Mitchell, E.P, Kostlanova, N, Wimmerova, M, Imberty, A. | | Deposit date: | 2005-05-27 | | Release date: | 2005-06-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (0.94 Å) | | Cite: | The fucose-binding lectin from Ralstonia solanacearum. A new type of beta-propeller architecture formed by oligomerization and interacting with fucoside, fucosyllactose, and plant xyloglucan.
J. Biol. Chem., 280, 2005
|
|
2BS6
 
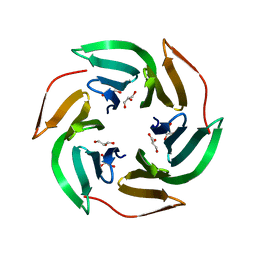 | | LECTIN FROM RALSTONIA SOLANACEARUM COMPLEXED WITH XYLOGLUCAN FRAGMENT | | Descriptor: | GLYCEROL, LECTIN, alpha-L-fucopyranose, ... | | Authors: | Mitchell, E.P, Kostlanova, N, Wimmerova, M, Imberty, A. | | Deposit date: | 2005-05-18 | | Release date: | 2005-05-19 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The fucose-binding lectin from Ralstonia solanacearum. A new type of beta-propeller architecture formed by oligomerization and interacting with fucoside, fucosyllactose, and plant xyloglucan.
J. Biol. Chem., 280, 2005
|
|
5A4D
 
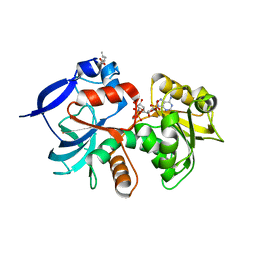 | | Crystal structure of the chloroplastic gamma-ketol reductase from Arabidopsis thaliana bound to 13KOTE and NADP | | Descriptor: | (13-oxo-9(Z),11(E),15(Z)-octadecatrienoic acid), NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PUTATIVE QUINONE-OXIDOREDUCTASE HOMOLOG, ... | | Authors: | Mas-y-mas, S, Curien, G, Giustini, C, Rolland, N, Ferrer, J.L, Cobessi, D. | | Deposit date: | 2015-06-08 | | Release date: | 2016-09-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.807 Å) | | Cite: | Crystal Structure of the Chloroplastic Oxoene Reductase ceQORH from Arabidopsis thaliana.
Front Plant Sci, 8, 2017
|
|
5A3V
 
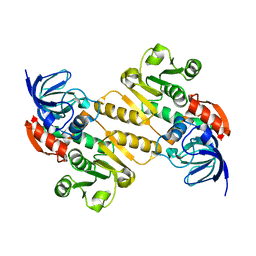 | | Crystal structure of the chloroplastic gamma-ketol reductase from Arabidopsis thaliana | | Descriptor: | PUTATIVE QUINONE-OXIDOREDUCTASE HOMOLOG, CHLOROPLASTIC | | Authors: | Mas-y-mas, S, Curien, G, Giustini, C, Rolland, N, Ferrer, J.L, Cobessi, D. | | Deposit date: | 2015-06-03 | | Release date: | 2016-09-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Crystal Structure of the Chloroplastic Oxoene Reductase ceQORH from Arabidopsis thaliana.
Front Plant Sci, 8, 2017
|
|
7PUE
 
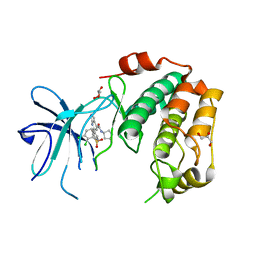 | | Human serum and glucocorticoid-regulated kinase 1 in complex with pyrazolopyridine inhibitor 3a | | Descriptor: | 6-[4-[[2,3-bis(chloranyl)phenyl]sulfonylamino]phenyl]-~{N}-[(3~{R})-pyrrolidin-3-yl]-2~{H}-pyrazolo[3,4-b]pyridine-4-carboxamide, GLYCEROL, Serine/threonine-protein kinase Sgk1 | | Authors: | Dreyer, M.K, Halland, N, Nazare, M. | | Deposit date: | 2021-09-29 | | Release date: | 2021-12-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.506 Å) | | Cite: | Rational Design of Highly Potent, Selective, and Bioavailable SGK1 Protein Kinase Inhibitors for the Treatment of Osteoarthritis.
J.Med.Chem., 65, 2022
|
|
2JHJ
 
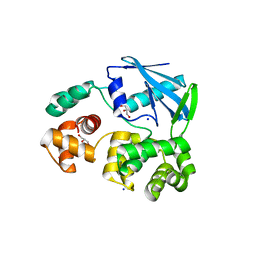 | | 3-methyladenine dna-glycosylase from Archaeoglobus fulgidus | | Descriptor: | 3-METHYLADENINE DNA-GLYCOSYLASE, GLYCEROL, SODIUM ION | | Authors: | Leiros, I, Nabong, M.P, Grosvik, K, Ringvoll, J, Haugland, G.T, Uldal, L, Reite, K, Olsbu, I.K, Knaevelsrud, I, Moe, E, Andersen, O.A, Birkeland, N.K, Ruoff, P, Klungland, A, Bjelland, S. | | Deposit date: | 2007-02-22 | | Release date: | 2007-04-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis for Enzymatic Excision of N1-Methyladenine and N3-Methylcytosine from DNA
Embo J., 26, 2007
|
|
7PAF
 
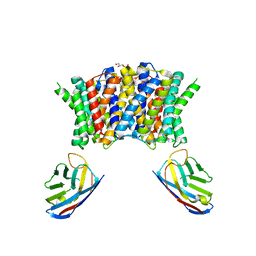 | | Streptococcus pneumoniae choline importer LicB in lipid nanodiscs | | Descriptor: | (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, LicB protein, Nanobody | | Authors: | Perez, C, Baerland, N. | | Deposit date: | 2021-07-29 | | Release date: | 2022-03-16 | | Method: | ELECTRON MICROSCOPY (3.75 Å) | | Cite: | Mechanistic basis of choline import involved in teichoic acids and lipopolysaccharide modification.
Sci Adv, 8, 2022
|
|
2JHN
 
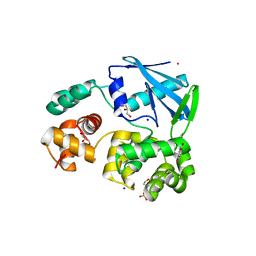 | | 3-methyladenine dna-glycosylase from Archaeoglobus fulgidus | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 3-METHYLADENINE DNA-GLYCOSYLASE, GLYCEROL, ... | | Authors: | Leiros, I, Nabong, M.P, Grosvik, K, Ringvoll, J, Haugland, G.T, Uldal, L, Reite, K, Olsbu, I.K, Knaevelsrud, I, Moe, E, Andersen, O.A, Birkeland, N.K, Ruoff, P, Klungland, A, Bjelland, S. | | Deposit date: | 2007-02-22 | | Release date: | 2007-04-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Basis for Enzymatic Excision of N1-Methyladenine and N3-Methylcytosine from DNA
Embo J., 26, 2007
|
|
3G56
 
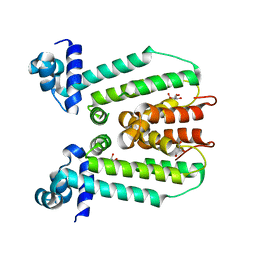 | | Structure of the macrolide biosensor protein, MphR(A) | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, Regulator of macrolide 2'-phosphotransferase I | | Authors: | Zheng, J, Sagar, V, Smolinsky, A, Bourke, C, LaRonde-LeBlanc, N, Cropp, T.A. | | Deposit date: | 2009-02-04 | | Release date: | 2009-03-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and function of the macrolide biosensor protein, MphR(A), with and without erythromycin
J.Mol.Biol., 387, 2009
|
|
3FRQ
 
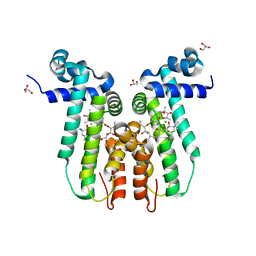 | | Structure of the macrolide biosensor protein, MphR(A), with erythromcyin | | Descriptor: | CHLORIDE ION, ERYTHROMYCIN A, GLYCEROL, ... | | Authors: | Zheng, J, Sagar, V, Smolinsky, A, Bourke, C, LaRonde-LeBlanc, N, Cropp, T.A. | | Deposit date: | 2009-01-08 | | Release date: | 2009-03-24 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structure and function of the macrolide biosensor protein, MphR(A), with and without erythromycin
J.Mol.Biol., 387, 2009
|
|
3GB5
 
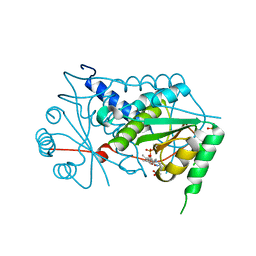 | | Crystal structure of Mus musculus iodotyrosine deiodinase (IYD) bound to FMN | | Descriptor: | ACETATE ION, FLAVIN MONONUCLEOTIDE, Iodotyrosine dehalogenase 1, ... | | Authors: | Thomas, S.R, McTamney, P.M, Adler, J.M, LaRonde-LeBlanc, N, Rokita, S.E. | | Deposit date: | 2009-02-18 | | Release date: | 2009-05-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of iodotyrosine deiodinase, a novel flavoprotein responsible for iodide salvage in thyroid glands.
J.Biol.Chem., 284, 2009
|
|
3GH8
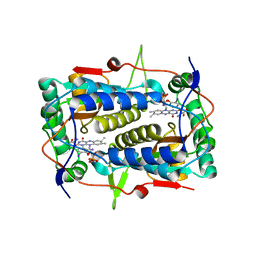 
 | | Crystal structure of Mus musculus iodotyrosine deiodinase (IYD) bound to FMN and di-iodotyrosine (DIT) | | Descriptor: | 3,5-DIIODOTYROSINE, FLAVIN MONONUCLEOTIDE, Iodotyrosine dehalogenase 1, ... | | Authors: | Thomas, S.R, McTamney, P.M, Adler, J.M, LaRonde-LeBlanc, N, Rokita, S.E. | | Deposit date: | 2009-03-03 | | Release date: | 2009-05-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Crystal structure of iodotyrosine deiodinase, a novel flavoprotein responsible for iodide salvage in thyroid glands.
J.Biol.Chem., 284, 2009
|
|
3GFD
 
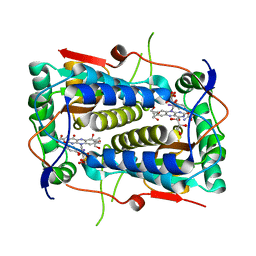 | | Crystal structure of Mus musculus iodotyrosine deiodinase (IYD) bound to FMN and mono-iodotyrosine (MIT) | | Descriptor: | 3-IODO-TYROSINE, FLAVIN MONONUCLEOTIDE, GLYCEROL, ... | | Authors: | Thomas, S.R, McTamney, P.M, Adler, J.M, LaRonde-LeBlanc, N, Rokita, S.E. | | Deposit date: | 2009-02-26 | | Release date: | 2009-05-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structure of iodotyrosine deiodinase, a novel flavoprotein responsible for iodide salvage in thyroid glands.
J.Biol.Chem., 284, 2009
|
|
3DLA
 
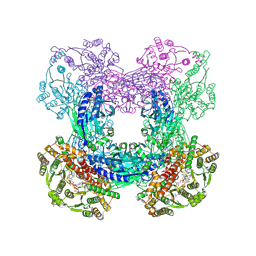 | | X-ray crystal structure of glutamine-dependent NAD+ synthetase from Mycobacterium tuberculosis bound to NaAD+ and DON | | Descriptor: | 5-OXO-L-NORLEUCINE, GLYCEROL, Glutamine-dependent NAD(+) synthetase, ... | | Authors: | LaRonde-LeBlanc, N.A, Resto, M, Gerratana, B. | | Deposit date: | 2008-06-26 | | Release date: | 2009-03-10 | | Last modified: | 2019-10-23 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Regulation of active site coupling in glutamine-dependent NAD(+) synthetase.
Nat.Struct.Mol.Biol., 16, 2009
|
|
8RCW
 
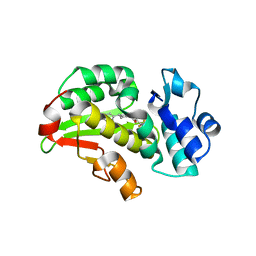 | | Crystal structure of the Mycobacterium tuberculosis regulator VirS (N-terminal fragment 4-208) in complex with the lead compound SMARt751 | | Descriptor: | 4,4,4-tris(fluoranyl)-1-[4-(4-fluorophenyl)piperidin-1-yl]butan-1-one, HTH-type transcriptional regulator VirS | | Authors: | Grosse, C, Sigoillot, M, Megalizzi, V, Tanina, A, Willand, N, Baulard, A.R, Wintjens, R. | | Deposit date: | 2023-12-07 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.692 Å) | | Cite: | Crystal structure of the Mycobacterium tuberculosis VirS regulator reveals its interaction with the lead compound SMARt751.
J.Struct.Biol., 216, 2024
|
|
2NSZ
 
 | |
8F5N
 
 | | Identification of an Immunodominant region on a Group A Streptococcus T-antigen Reveals Temperature-Dependent Motion in Pili | | Descriptor: | CALCIUM ION, Mouse-Human Fab heavy chain, Mouse-Human Fab light chain, ... | | Authors: | Raynes, J.M, Young, P.G, Moreland, N.J. | | Deposit date: | 2022-11-14 | | Release date: | 2023-03-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of an immunodominant region on a group A Streptococcus T-antigen reveals temperature-dependent motion in pili.
Virulence, 14, 2023
|
|
8F70
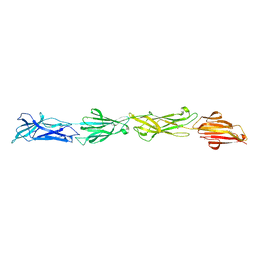 
 | |
6N0A
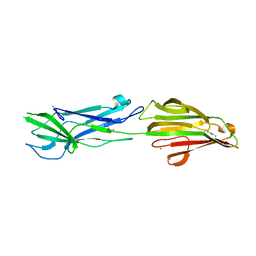 
 | | Structure of the major pilin protein (T-18.1) from Streptococcus pyogenes serotype MGAS8232 | | Descriptor: | CALCIUM ION, Major pilin backbone protein T-antigen | | Authors: | Young, P.G, Raynes, J.M, Loh, J.M, Proft, T, Baker, E.N, Moreland, N.J. | | Deposit date: | 2018-11-06 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Group AStreptococcusT Antigens Have a Highly Conserved Structure Concealed under a Heterogeneous Surface That Has Implications for Vaccine Design.
Infect.Immun., 87, 2019
|
|
1ZAO
 
 | | Crystal Structure of A.fulgidus Rio2 Kinase Complexed With ATP and Manganese Ions | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, MANGANESE (II) ION, ... | | Authors: | Laronde-Leblanc, N, Guszczynski, T, Copeland, T, Wlodawer, A. | | Deposit date: | 2005-04-06 | | Release date: | 2005-06-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Autophosphorylation of Archaeoglobus fulgidus Rio2 and crystal structures of its nucleotide-metal ion complexes.
Febs J., 272, 2005
|
|
2ION
 
 | |
2IOL
 
 | |
