8H08
 
 | |
2KH5
 
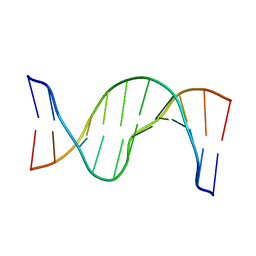 | | Solution Structure of cis-5R,6S-thymine glycol opposite complementary adenine in duplex DNA | | Descriptor: | 5'-D(*AP*CP*AP*AP*AP*CP*AP*CP*GP*CP*AP*C)-3', 5'-D(*GP*TP*GP*CP*GP*(CTG)P*GP*TP*TP*TP*GP*T)-3' | | Authors: | Brown, K.L. | | Deposit date: | 2009-03-24 | | Release date: | 2010-03-02 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Binding of the human nucleotide excision repair proteins XPA and XPC/HR23B to the 5R-thymine glycol lesion and structure of the cis-(5R,6S) thymine glycol epimer in the 5'-GTgG-3' sequence: destabilization of two base pairs at the lesion site
Nucleic Acids Res., 38, 2010
|
|
2KH6
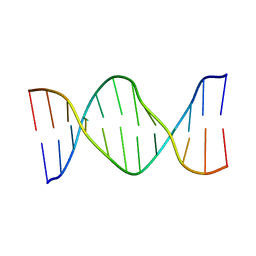 
 | | Solution Structure of cis-5R,6S-thymine glycol opposite complementary adenine in duplex DNA | | Descriptor: | 5'-D(*AP*CP*AP*AP*AP*CP*AP*CP*GP*CP*AP*C)-3', 5'-D(*GP*TP*GP*CP*GP*(CTG)P*GP*TP*TP*TP*GP*T)-3' | | Authors: | Brown, K.L. | | Deposit date: | 2009-03-24 | | Release date: | 2010-03-02 | | Last modified: | 2020-02-26 | | Method: | SOLUTION NMR | | Cite: | Binding of the human nucleotide excision repair proteins XPA and XPC/HR23B to the 5R-thymine glycol lesion and structure of the cis-(5R,6S) thymine glycol epimer in the 5'-GTgG-3' sequence: destabilization of two base pairs at the lesion site
Nucleic Acids Res., 38, 2010
|
|
8ERB
 
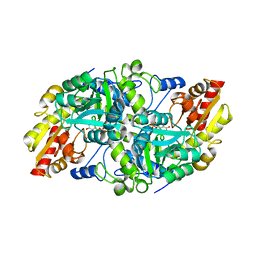 | | Crystal structure of Fub7 in complex with vinylglycine ketimine | | Descriptor: | (2E)-2-[({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)imino]but-3-enoic acid, Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-11 | | Release date: | 2023-10-18 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
8EQW
 
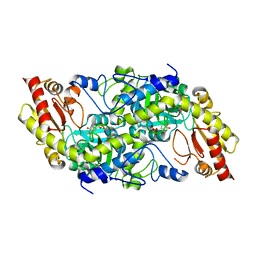 | | Crystal structure of Fub7 | | Descriptor: | Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-10 | | Release date: | 2023-10-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
8ERJ
 
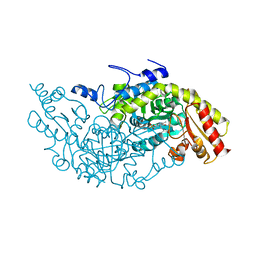 | | Crystal structure of Fub7 in complex with E-2-aminocrotonate | | Descriptor: | (2E)-2-{[(1E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}but-2-enoic acid, (2S)-2-[({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)amino]but-3-enoic acid, Sulfhydrylase FUB7 | | Authors: | Hai, Y. | | Deposit date: | 2022-10-12 | | Release date: | 2023-10-25 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Molecular and Structural Basis for C gamma-C Bond Formation by PLP-Dependent Enzyme Fub7.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
3WOH
 
 | |
3RER
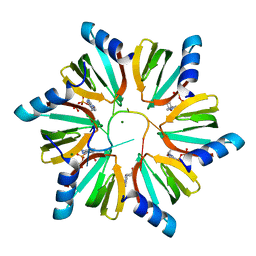 
 | | Crystal structure of E. coli Hfq in complex with AU6A RNA and ADP | | Descriptor: | 5'-R(*AP*UP*UP*UP*UP*UP*UP*A)-3', ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Wang, W.W, Wu, J.H, Shi, Y.Y. | | Deposit date: | 2011-04-05 | | Release date: | 2011-10-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Cooperation of Escherichia coli Hfq hexamers in DsrA binding.
Genes Dev., 25, 2011
|
|
8DL5
 
 | |
5L30
 
 | | Factor VIIa in complex with the inhibitor (2R,15R)-2-[(1-aminoisoquinolin-6-yl)amino]-4,15,17-trimethyl-7-[1-(1H-tetrazol-5-yl)cyclopropyl]-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6(21),7,9,16,19-hexaene-3,12-dione | | Descriptor: | (2R,15R)-2-[(1-aminoisoquinolin-6-yl)amino]-4,15,17-trimethyl-7-[1-(1H-tetrazol-5-yl)cyclopropyl]-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6(21),7,9,16,19-hexaene-3,12-dione, CALCIUM ION, Coagulation factor VII (Heavy Chain), ... | | Authors: | Wei, A. | | Deposit date: | 2016-08-02 | | Release date: | 2016-09-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Synthesis and P1' SAR exploration of potent macrocyclic tissue factor-factor VIIa inhibitors.
Bioorg.Med.Chem.Lett., 26, 2016
|
|
5L2Y
 
 | | Factor VIIa in complex with the inhibitor 1-[(2R,15R)-2-[(1-amino-4-fluoroisoquinolin-6-yl)amino]-4,15,20-trimethyl-3,12-dioxo-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6,8,10(21),16,19-hexaen-7-yl] cyclobutane-1-carboxylic acid | | Descriptor: | 1-[(2R,15R)-2-[(1-amino-4-fluoroisoquinolin-6-yl)amino]-4,15,17-trimethyl-3,12-dioxo-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6(21),7,9,16,19-hexaen-7-yl]cyclobutane-1-carboxylic acid, CALCIUM ION, Coagulation factor VII (Heavy Chain), ... | | Authors: | Wei, A. | | Deposit date: | 2016-08-02 | | Release date: | 2016-09-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Synthesis and P1' SAR exploration of potent macrocyclic tissue factor-factor VIIa inhibitors.
Bioorg.Med.Chem.Lett., 26, 2016
|
|
5L2Z
 
 | | Factor VIIa in complex with the inhibitor 1-[(2R,15R)-2-[(1-amino-4-fluoroisoquinolin-6-yl)amino]-4,15,17-trimethyl-3,12-dioxo-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6(21),7,9,16,19-hexaen-7-yl]cyclohexane-1-carboxylic acid | | Descriptor: | 1-[(2R,15R)-2-[(1-amino-4-fluoroisoquinolin-6-yl)amino]-4,15,17-trimethyl-3,12-dioxo-13-oxa-4,11-diazatricyclo[14.2.2.1~6,10~]henicosa-1(18),6(21),7,9,16,19-hexaen-7-yl]cyclohexane-1-carboxylic acid, CALCIUM ION, Coagulation factor VII (Heavy Chain), ... | | Authors: | Wei, A. | | Deposit date: | 2016-08-02 | | Release date: | 2016-09-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Synthesis and P1' SAR exploration of potent macrocyclic tissue factor-factor VIIa inhibitors.
Bioorg.Med.Chem.Lett., 26, 2016
|
|
5MEZ
 
 | | Crystal structure of Smad4-MH1 bound to the GGCT site. | | Descriptor: | CHLORIDE ION, DNA (5'-D(P*GP*CP*AP*GP*GP*CP*TP*AP*GP*CP*CP*TP*GP*CP*A)-3'), MH1 domain of human Smad4, ... | | Authors: | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | Deposit date: | 2016-11-16 | | Release date: | 2017-11-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
5MEY
 
 | | Crystal structure of Smad4-MH1 bound to the GGCGC site. | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | Deposit date: | 2016-11-16 | | Release date: | 2017-11-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
5MF0
 
 | | Crystal structure of Smad4-MH1 bound to the GGCCG site. | | Descriptor: | CHLORIDE ION, DNA (5'-D(P*AP*CP*GP*GP*GP*CP*CP*GP*CP*GP*GP*CP*CP*CP*GP*T)-3'), MH1 domain of human Smad4, ... | | Authors: | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | Deposit date: | 2016-11-16 | | Release date: | 2017-11-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
5NM9
 
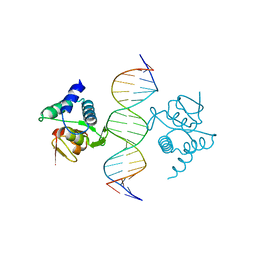 | | Crystal structure of the placozoa Trichoplax adhaerens Smad4-MH1 bound to the GGCGC site. | | Descriptor: | DNA (5'-D(P*AP*TP*GP*CP*GP*GP*GP*CP*GP*CP*GP*CP*CP*CP*GP*CP*AP*T)-3'), Mothers against decapentaplegic homolog, ZINC ION | | Authors: | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | Deposit date: | 2017-04-05 | | Release date: | 2017-11-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
4JWF
 
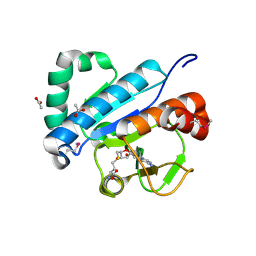 | | Crystal structure of spTrm10(74)-SAH complex | | Descriptor: | ACETIC ACID, S-ADENOSYL-L-HOMOCYSTEINE, tRNA (guanine(9)-N1)-methyltransferase | | Authors: | Yan, W, Shao, Z. | | Deposit date: | 2013-03-27 | | Release date: | 2013-10-16 | | Last modified: | 2014-02-12 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate.
Nucleic Acids Res., 42, 2014
|
|
4JWH
 
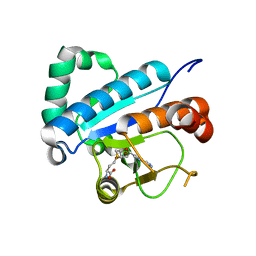 | | Crystal structure of spTrm10(Full length)-SAH complex | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, tRNA (guanine(9)-N1)-methyltransferase | | Authors: | Yan, W, Shao, Z. | | Deposit date: | 2013-03-27 | | Release date: | 2013-10-16 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate.
Nucleic Acids Res., 42, 2014
|
|
4JWJ
 
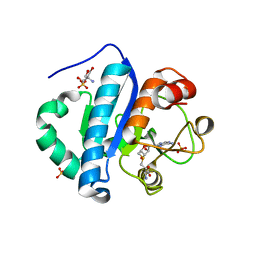 | | Crystal structure of scTrm10(84)-SAH complex | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, S-ADENOSYL-L-HOMOCYSTEINE, SULFATE ION, ... | | Authors: | Yan, W, Shao, Z. | | Deposit date: | 2013-03-27 | | Release date: | 2013-10-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate.
Nucleic Acids Res., 42, 2014
|
|
4JWG
 
 | | Crystal structure of spTrm10(74) | | Descriptor: | ACETIC ACID, tRNA (guanine(9)-N1)-methyltransferase | | Authors: | Yan, W, Shao, Z. | | Deposit date: | 2013-03-27 | | Release date: | 2013-10-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate.
Nucleic Acids Res., 42, 2014
|
|
3RES
 
 | |
6H3R
 
 | |
5GXO
 
 | | Discovery of a compound that activates SIRT3 to deacetylate Manganese Superoxide Dismutase | | Descriptor: | MANGANESE (II) ION, Superoxide dismutase [Mn], mitochondrial | | Authors: | Lu, J, Li, J, Wu, M, Wang, J, Xia, Q. | | Deposit date: | 2016-09-19 | | Release date: | 2017-08-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A small molecule activator of SIRT3 promotes deacetylation and activation of manganese superoxide dismutase.
Free Radic. Biol. Med., 112, 2017
|
|
7X2H
 
 | |
7XD2
 
 | |
