8CIJ
 
 | | CRYSTAL STRUCTURE OF HUMAN HPK1 (MAP4K1) COMPLEX WITH 2-[8-Amino-7-fluoro-6-(8-methyl-2,3-dihydro-1H-pyrido[2,3-b][1,4]oxazin-7-yl)-isoquinolin-3-ylamino]-6-isopropyl-5,6-dihydro-4H-1,6,8a-triaza-azulen-7-one | | Descriptor: | 2-[[8-azanyl-7-fluoranyl-6-(8-methyl-2,3-dihydro-1~{H}-pyrido[2,3-b][1,4]oxazin-7-yl)isoquinolin-3-yl]amino]-6-propan-2-yl-5,8-dihydro-4~{H}-pyrazolo[1,5-d][1,4]diazepin-7-one, Mitogen-activated protein kinase kinase kinase kinase 1 | | Authors: | Musil, D, Toure, M. | | Deposit date: | 2023-02-09 | | Release date: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.821 Å) | | Cite: | Discovery of quinazoline HPK1 inhibitors with high cellular potency.
Bioorg.Med.Chem., 92, 2023
|
|
6QEH
 
 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX WITH AN INHIBITOR 5-Chloro-quinolin-8-ol | | Descriptor: | 5-chloranylquinolin-8-ol, GLYCEROL, MANGANESE (II) ION, ... | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2019-01-07 | | Release date: | 2019-05-01 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Discovery and Structure-Based Optimization of Next-Generation Reversible Methionine Aminopeptidase-2 (MetAP-2) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6QEI
 
 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX WITH AN INHIBITOR 5,6-Difluoro-3-(2-isopropoxy-4-piperazin-1-yl-phenyl)-1H-indole-2-carboxylic acid amide | | Descriptor: | 1,2-ETHANEDIOL, 5,6-bis(fluoranyl)-3-(4-piperazin-1-yl-2-propan-2-yloxy-phenyl)-1~{H}-indole-2-carboxamide, DIMETHYL SULFOXIDE, ... | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2019-01-07 | | Release date: | 2019-05-01 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Discovery and Structure-Based Optimization of Next-Generation Reversible Methionine Aminopeptidase-2 (MetAP-2) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6QEJ
 
 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX WITH AN INHIBITOR Thiophene-2-sulfonic acid (4-fluoro-benzyl)-(4H-[1,2,4]triazol-3-ylmethyl)-amide | | Descriptor: | 1,2-ETHANEDIOL, MANGANESE (II) ION, Methionine aminopeptidase 2, ... | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2019-01-07 | | Release date: | 2019-05-01 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Discovery and Structure-Based Optimization of Next-Generation Reversible Methionine Aminopeptidase-2 (MetAP-2) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6QED
 
 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX; WITH AN INHIBITOR (S)-3-Hydroxy-2-oxo-1-(2-oxo-1,2,3,4-tetrahydro-quinolin-6-yl)-pyrrolidine-3-carboxylic acid 3-chloro-5-fluoro-benzylamide | | Descriptor: | (3~{S})-~{N}-[(3-chloranyl-5-fluoranyl-phenyl)methyl]-3-oxidanyl-2-oxidanylidene-1-(2-oxidanylidene-3,4-dihydro-1~{H}-quinolin-6-yl)pyrrolidine-3-carboxamide, 1,2-ETHANEDIOL, DIMETHYL SULFOXIDE, ... | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2019-01-07 | | Release date: | 2019-05-01 | | Last modified: | 2019-06-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Discovery and Structure-Based Optimization of Next-Generation Reversible Methionine Aminopeptidase-2 (MetAP-2) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6QEF
 
 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX WITH AN INHIBITOR (S)-3-Hydroxy-2-oxo-1-phenyl-pyrrolidine-3-carboxylic acid 3-chloro-5-fluoro-benzylamide | | Descriptor: | (3~{S})-~{N}-[(3-chloranyl-5-fluoranyl-phenyl)methyl]-3-oxidanyl-2-oxidanylidene-1-phenyl-pyrrolidine-3-carboxamide, 1,2-ETHANEDIOL, GLYCEROL, ... | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2019-01-07 | | Release date: | 2019-05-01 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Discovery and Structure-Based Optimization of Next-Generation Reversible Methionine Aminopeptidase-2 (MetAP-2) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6YR9
 
 | |
6YQ1
 
 | | FOCAL ADHESION KINASE CATALYTIC DOMAIN IN COMPLEX WITH N-Methyl-N-(3-{[2-(2-oxo-1,2,3,4-tetrahydro-quinolin-6-ylamino)-5-trifluoromethyl-pyrimidin-4-ylamino]-methyl}-pyridin-2-yl)-methanesulfonamide | | Descriptor: | Focal adhesion kinase 1, SODIUM ION, SULFATE ION, ... | | Authors: | Musil, D, Heinrich, T, Amaral, M. | | Deposit date: | 2020-04-16 | | Release date: | 2021-02-10 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.784 Å) | | Cite: | Structure-kinetic relationship reveals the mechanism of selectivity of FAK inhibitors over PYK2.
Cell Chem Biol, 28, 2021
|
|
6YOJ
 
 | |
6ZJZ
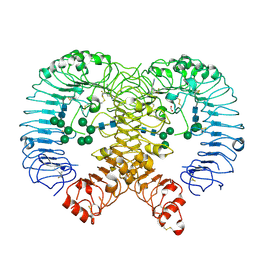 
 | | Discovery of M5049: a novel selective TLR7/8 inhibitor for treatment of autoimmunity | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 5-[(3~{R},5~{S})-3-azanyl-5-(trifluoromethyl)piperidin-1-yl]quinoline-8-carbonitrile, FORMIC ACID, ... | | Authors: | Musil, D, Lehman, M, Strauss, J. | | Deposit date: | 2020-06-29 | | Release date: | 2020-12-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.489 Å) | | Cite: | Discovery of M5049: A Novel Selective Toll-Like Receptor 7/8 Inhibitor for Treatment of Autoimmunity.
J.Pharmacol.Exp.Ther., 376, 2021
|
|
6YT6
 
 | |
6YVS
 
 | | FOCAL ADHESION KINASE CATALYTIC DOMAIN IN COMPLEX WITH 5-{4-[(Pyridin-3-ylmethyl)-amino]-5-trifluoromethyl-pyrimidin-2-ylamino}-1,3-dihydro-indol-2-one | | Descriptor: | 5-[[4-(pyridin-3-ylmethylamino)-5-(trifluoromethyl)pyrimidin-2-yl]amino]-1,3-dihydroindol-2-one, Focal adhesion kinase 1, SULFATE ION | | Authors: | Musil, D, Heinrich, T, Amaral, M. | | Deposit date: | 2020-04-28 | | Release date: | 2021-02-10 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structure-kinetic relationship reveals the mechanism of selectivity of FAK inhibitors over PYK2.
Cell Chem Biol, 28, 2021
|
|
6YVY
 
 | |
6YXV
 
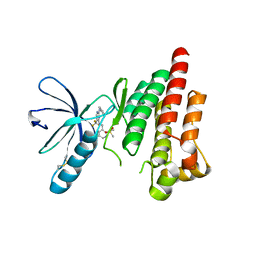 | |
5LYW
 
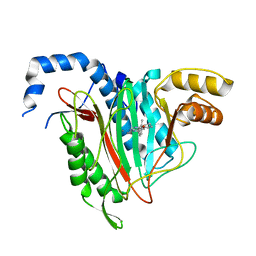 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX; WITH AN INHIBITOR 6-((R)-2-o-Tolyloxymethyl-pyrrolidin-1-yl)-9H-purine | | Descriptor: | 6-[(2~{R})-2-[(2-methylphenoxy)methyl]pyrrolidin-1-yl]-7~{H}-purine, MANGANESE (II) ION, Methionine aminopeptidase 2 | | Authors: | Musil, D, Heinrich, T, Knoechel, T, Lehmann, M. | | Deposit date: | 2016-09-28 | | Release date: | 2017-08-16 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Novel reversible methionine aminopeptidase-2 (MetAP-2) inhibitors based on purine and related bicyclic templates.
Bioorg. Med. Chem. Lett., 27, 2017
|
|
5LYX
 
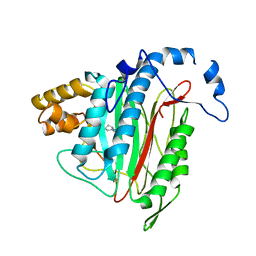 | | CRYSTAL STRUCTURE OF HUMAN METHIONINE AMINOPEPTIDASE-2 IN COMPLEX; WITH AN INHIBITOR 5-((R)-1-[1,2,4]Triazolo[1,5-a]pyrimidin-7-yl-pyrrolidin-2-ylmethoxy)-isoquinoline | | Descriptor: | 5-[[(2~{R})-1-([1,2,4]triazolo[1,5-a]pyrimidin-7-yl)pyrrolidin-2-yl]methoxy]isoquinoline, MANGANESE (II) ION, Methionine aminopeptidase 2 | | Authors: | Musil, D, Heinrich, T, Lehmann, M. | | Deposit date: | 2016-09-29 | | Release date: | 2017-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Novel reversible methionine aminopeptidase-2 (MetAP-2) inhibitors based on purine and related bicyclic templates.
Bioorg. Med. Chem. Lett., 27, 2017
|
|
4EEH
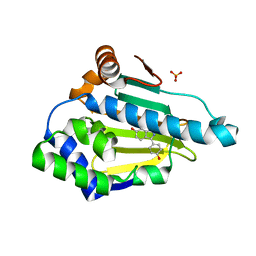 
 | | Hsp90 Alpha N-terminal Domain in Complex with an Inhibitor 3-(4-Hydroxy-phenyl)-1H-indazol-6-ol | | Descriptor: | 3-(4-hydroxyphenyl)-1H-indazol-6-ol, Heat shock protein HSP 90-alpha, SULFATE ION | | Authors: | Musil, D, Lehmann, M, Graedler, U, Buchstaller, H.-P. | | Deposit date: | 2012-03-28 | | Release date: | 2012-06-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Fragment-based discovery of hydroxy-indazole-carboxamides as novel small molecule inhibitors of Hsp90
Bioorg.Med.Chem.Lett., 22, 2012
|
|
4EFT
 
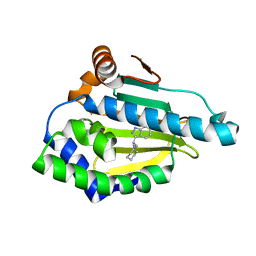 | | Hsp90 Alpha N-terminal Domain in Complex with an Inhibitor 3-Cyclohexyl-2-(6-hydroxy-1H-indazol-3-yl)-propionitrile | | Descriptor: | (2R)-3-cyclohexyl-2-(6-hydroxy-1H-indazol-3-yl)propanenitrile, Heat shock protein HSP 90-alpha | | Authors: | Musil, D, Lehmann, M, Graedler, U, Buchstaller, H.-P. | | Deposit date: | 2012-03-30 | | Release date: | 2012-06-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Fragment-based discovery of hydroxy-indazole-carboxamides as novel small molecule inhibitors of Hsp90
Bioorg.Med.Chem.Lett., 22, 2012
|
|
6ELP
 
 | |
6EL5
 
 | | Estimation of relative drug-target residence times by random acceleration molecular dynamics simulation | | Descriptor: | 8-(2-CHLORO-3,4,5-TRIMETHOXY-BENZYL)-2-FLUORO-9-PENT-4-YLNYL-9H-PURIN-6-YLAMINE, Heat shock protein HSP 90-alpha | | Authors: | Musil, D, Lehmann, M, Eggenweiler, H.-M. | | Deposit date: | 2017-09-27 | | Release date: | 2018-05-30 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Estimation of Drug-Target Residence Times by tau-Random Acceleration Molecular Dynamics Simulations.
J Chem Theory Comput, 14, 2018
|
|
6EI5
 
 | |
6ELO
 
 | |
6ELN
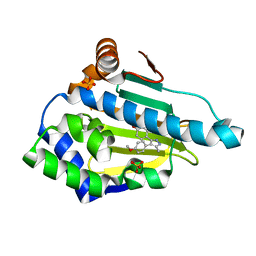 
 | | Estimation of relative drug-target residence times by random acceleration molecular dynamics simulation | | Descriptor: | 4-[4-(4-methoxyphenyl)-5-methyl-1H-pyrazol-3-yl]benzene-1,3-diol, Heat shock protein HSP 90-alpha, SULFATE ION | | Authors: | Musil, D, Lehmann, M, Eggenweiler, H.-M. | | Deposit date: | 2017-09-29 | | Release date: | 2018-05-30 | | Last modified: | 2018-07-18 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Estimation of Drug-Target Residence Times by tau-Random Acceleration Molecular Dynamics Simulations.
J Chem Theory Comput, 14, 2018
|
|
6EY9
 
 | |
6EY8
 
 | | Estimation of relative drug-target residence times by random acceleration molecular dynamics simulation | | Descriptor: | DIMETHYL SULFOXIDE, Heat shock protein HSP 90-alpha, SULFATE ION, ... | | Authors: | Musil, D, Lehmann, M, Buchstaller, H.-P. | | Deposit date: | 2017-11-11 | | Release date: | 2018-05-30 | | Last modified: | 2018-07-18 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Estimation of Drug-Target Residence Times by tau-Random Acceleration Molecular Dynamics Simulations.
J Chem Theory Comput, 14, 2018
|
|
