8A67
 
 | |
7NPO
 
 | | Branched K48-K63-Ub3 | | Descriptor: | GLYCEROL, Polyubiquitin-B | | Authors: | Lange, S.M, Kwasna, D, Kulathu, Y. | | Deposit date: | 2021-02-27 | | Release date: | 2022-08-10 | | Last modified: | 2024-07-31 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | VCP/p97-associated proteins are binders and debranching enzymes of K48-K63-branched ubiquitin chains.
Nat.Struct.Mol.Biol., 2024
|
|
7NBB
 
 | |
6RUU
 
 | | Pseudokinase domain of human IRAK3 | | Descriptor: | GLYCEROL, Interleukin-1 receptor-associated kinase 3, MERCURY (II) ION, ... | | Authors: | Lange, S.M, Kulathu, Y, Cohen, P. | | Deposit date: | 2019-05-29 | | Release date: | 2020-09-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Dimeric Structure of the Pseudokinase IRAK3 Suggests an Allosteric Mechanism for Negative Regulation.
Structure, 29, 2021
|
|
7NPI
 
 | | Crystal structure of Mindy2 (C266A) in complex with Lys48-linked penta-ubiquitin (K48-Ub5) | | Descriptor: | CHLORIDE ION, Polyubiquitin-C, SODIUM ION, ... | | Authors: | Lange, S.M, Armstrong, L.A, Kulathu, Y. | | Deposit date: | 2021-02-26 | | Release date: | 2021-09-15 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
6Z7V
 
 | | Crystal structure of Mindy2 (C266A) in complex with Lys48 linked di-ubiquitin (K48-Ub2) | | Descriptor: | POTASSIUM ION, Polyubiquitin-C, TETRAETHYLENE GLYCOL, ... | | Authors: | Abdul Rehman, S.A, Lange, S.M, Kulathu, Y. | | Deposit date: | 2020-06-01 | | Release date: | 2021-06-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
5HNH
 
 | | Crystal structure of pyrene- and phenanthrene-modified DNA in complex with the BpuJ1 endonuclease binding domain | | Descriptor: | DNA (5'-D(*GP*(YPY)P*AP*CP*CP*CP*GP*TP*GP*GP*A)-3'), DNA (5'-D(*TP*CP*CP*AP*CP*GP*GP*GP*T*(YPY)*(YPY)*C)-3'), Restriction endonuclease R.BpuJI | | Authors: | Probst, M, Aeschimann, W, Chau, T.-T.-H, Langenegger, S.M, Stocker, A, Haener, R. | | Deposit date: | 2016-01-18 | | Release date: | 2016-08-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.876 Å) | | Cite: | Structural insight into DNA-assembled oligochromophores: crystallographic analysis of pyrene- and phenanthrene-modified DNA in complex with BpuJI endonuclease.
Nucleic Acids Res., 44, 2016
|
|
5HLT
 
 | | Crystal structure of pyrene- and phenanthrene-modified DNA in complex with the BpuJ1 endonuclease binding domain | | Descriptor: | DNA (5'-D(*GP*YPY*TP*AP*CP*CP*CP*GP*TP*GP*GP*A)-3'), DNA (5'-D(*TP*CP*CP*AP*CP*GP*GP*GP*TP*AP*YPY*C)-3'), Restriction endonuclease R.BpuJI | | Authors: | Probst, M, Aeschimann, W, Chau, T.-T.-H, Langenegger, S.M, Stocker, A, Haener, R. | | Deposit date: | 2016-01-15 | | Release date: | 2016-08-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.672 Å) | | Cite: | Structural insight into DNA-assembled oligochromophores: crystallographic analysis of pyrene- and phenanthrene-modified DNA in complex with BpuJI endonuclease.
Nucleic Acids Res., 44, 2016
|
|
5HNF
 
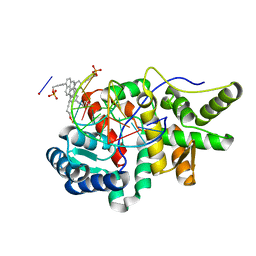 | | Crystal structure of pyrene- and phenanthrene-modified DNA in complex with the BpuJ1 endonuclease binding domain | | Descriptor: | DNA (5'-D(*GP*(YPE)P*AP*CP*CP*CP*GP*TP*GP*GP*A)-3'), DNA (5'-D(*TP*CP*CP*AP*CP*GP*GP*GP*TP*(YPF)P*C)-3'), Restriction endonuclease R.BpuJI | | Authors: | Probst, M, Aeschimann, W, Chau, T.-T.-H, Langenegger, S.M, Stocker, A, Haener, R. | | Deposit date: | 2016-01-18 | | Release date: | 2016-08-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.546 Å) | | Cite: | Structural insight into DNA-assembled oligochromophores: crystallographic analysis of pyrene- and phenanthrene-modified DNA in complex with BpuJI endonuclease.
Nucleic Acids Res., 44, 2016
|
|
6Y6R
 
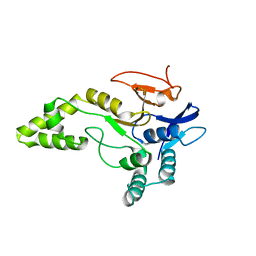 | | Crystal structure of MINDY1 T335D mutant | | Descriptor: | Ubiquitin carboxyl-terminal hydrolase MINDY-1 | | Authors: | Abdul Rehman, S.A, Kulathu, Y. | | Deposit date: | 2020-02-27 | | Release date: | 2021-03-31 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
6YJG
 
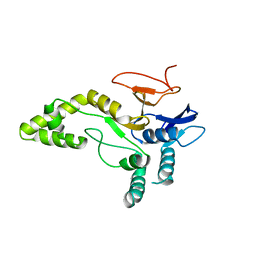 | | Crystal structure of MINDY1 mutant-Y114F | | Descriptor: | Ubiquitin carboxyl-terminal hydrolase MINDY1 | | Authors: | Abdul Rehman, S.A, Kulathu, Y. | | Deposit date: | 2020-04-03 | | Release date: | 2021-04-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.28 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
6Z49
 
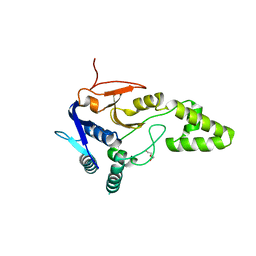 | | Crystal structure of deubiquitinase Mindy2 | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, TETRAETHYLENE GLYCOL, ... | | Authors: | Abdul Rehman, S.A, Kulathu, Y. | | Deposit date: | 2020-05-23 | | Release date: | 2021-06-02 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
6Z90
 
 | | Crystal structure of MINDY1 mutant-P138A | | Descriptor: | Ubiquitin carboxyl-terminal hydrolase MINDY-1 | | Authors: | Abdul Rehman, S.A, Kulathu, Y. | | Deposit date: | 2020-06-03 | | Release date: | 2021-06-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.59 Å) | | Cite: | Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2.
Mol.Cell, 81, 2021
|
|
6TXB
 
 | |
6TUV
 
 | |
4ZI3
 
 | | BART-like domain of BARTL1/CCDC104 aa1-133 in complex with Arl3FL bound to GppNHp in P1 21 1 | | Descriptor: | ADP-ribosylation factor-like protein 3, Cilia- and flagella-associated protein 36, MAGNESIUM ION, ... | | Authors: | Lokaj, M, Koerner, C, Koesling, S, Wittinghofer, A. | | Deposit date: | 2015-04-27 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Interaction of CCDC104/BARTL1 with Arl3 and Implications for Ciliary Function.
Structure, 23, 2015
|
|
4ZI2
 
 | | BART-like domain of BARTL1/CCDC104 in complex with Arl3FL bound to GppNHp in P21 21 21 | | Descriptor: | ADP-ribosylation factor-like protein 3, Cilia- and flagella-associated protein 36, MAGNESIUM ION, ... | | Authors: | Lokaj, M, Koerner, C, Koesling, S, Wittinghofer, A. | | Deposit date: | 2015-04-27 | | Release date: | 2015-11-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Interaction of CCDC104/BARTL1 with Arl3 and Implications for Ciliary Function.
Structure, 23, 2015
|
|
8B5W
 
 | | Crystal structure of the E3 module from UBR4 | | Descriptor: | 1,2-ETHANEDIOL, ZINC ION, cDNA FLJ12511 fis, ... | | Authors: | Virdee, S, Mabbitt, P.D, Barnsby-Greer, L. | | Deposit date: | 2022-09-24 | | Release date: | 2023-10-04 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | UBE2A and UBE2B are recruited by an atypical E3 ligase module in UBR4.
Nat.Struct.Mol.Biol., 31, 2024
|
|
8BTL
 
 | | Crystal structure of a complex between the E2 conjugating enzyme UBE2A and the E3 ligase module from UBR4 | | Descriptor: | Ubiquitin conjugating enzyme E2 A, ZINC ION, cDNA FLJ12511 fis, ... | | Authors: | Virdee, S, Mabbitt, P.D, Barnsby-Greer, L. | | Deposit date: | 2022-11-29 | | Release date: | 2023-12-13 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | UBE2A and UBE2B are recruited by an atypical E3 ligase module in UBR4.
Nat.Struct.Mol.Biol., 31, 2024
|
|
