5C16
 
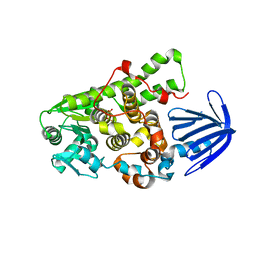 | | Myotubularin-related proetin 1 | | Descriptor: | Myotubularin-related protein 1, PHOSPHATE ION | | Authors: | Lee, B.I, Bong, S.M. | | Deposit date: | 2015-06-13 | | Release date: | 2016-04-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Crystal Structure of Human Myotubularin-Related Protein 1 Provides Insight into the Structural Basis of Substrate Specificity
Plos One, 11, 2016
|
|
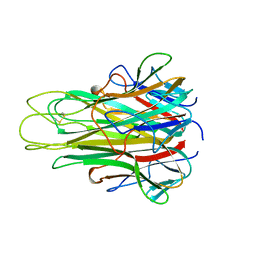
5TSW
 
 | |
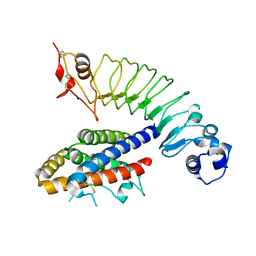
4J4L
 
 | |
3RFS
 
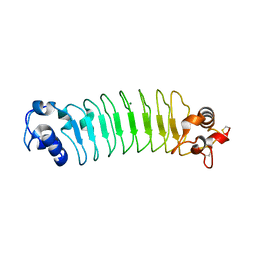 | | Design of a binding scaffold based on variable lymphocyte receptors of jawless vertebrates by module engineering | | Descriptor: | Internalin B, repeat modules, Variable lymphocyte receptor B, ... | | Authors: | Kim, H.J, Cheong, H.K, Jeon, Y.H. | | Deposit date: | 2011-04-06 | | Release date: | 2012-03-14 | | Last modified: | 2017-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Design of a binding scaffold based on variable lymphocyte receptors of jawless vertebrates by module engineering
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
5HZ2
 
 | | Crystal structure of PhaC1 from Ralstonia eutropha | | Descriptor: | GLYCEROL, Poly-beta-hydroxybutyrate polymerase, SULFATE ION | | Authors: | Kim, J, Kim, K.-J. | | Deposit date: | 2016-02-02 | | Release date: | 2016-12-07 | | Last modified: | 2017-04-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Ralstonia eutropha polyhydroxyalkanoate synthase C-terminal domain and reaction mechanisms.
Biotechnol J, 12, 2017
|
|
7MEJ
 
 | |
7MDW
 | |
7ME7
 
 | |
7N9C
 
 | |
7N9E
 
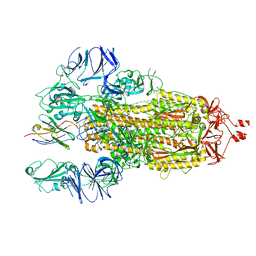 | |
7N9B
 
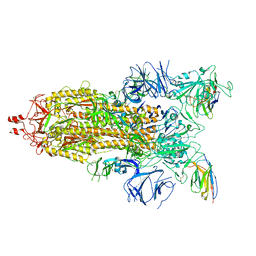 | |
7N9T
 
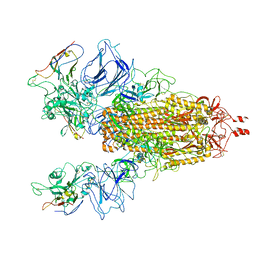 | |
4IKC
 
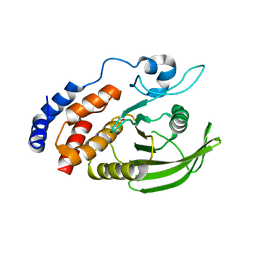 | | Crystal Structure of catalytic domain of PTPRQ | | Descriptor: | CHLORIDE ION, Phosphotidylinositol phosphatase PTPRQ, SULFATE ION | | Authors: | Yu, K.R, Ryu, S.E, Kim, S.J. | | Deposit date: | 2012-12-26 | | Release date: | 2013-07-31 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Structural basis for the dephosphorylating activity of PTPRQ towards phosphatidylinositide substrates
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4MFU
 
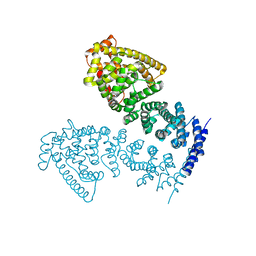 | | Crystal structure of human CTNNBL1(residues 77~563) | | Descriptor: | Beta-catenin-like protein 1 | | Authors: | Ahn, J.W, Kim, S, Kim, K.J. | | Deposit date: | 2013-08-28 | | Release date: | 2014-03-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.744 Å) | | Cite: | Structural insights into the novel ARM-repeat protein CTNNBL1 and its association with the hPrp19-CDC5L complex
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4MFV
 
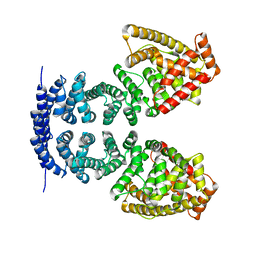 | | Crystal structure of human CTNNBL1(residues 33~563) | | Descriptor: | Beta-catenin-like protein 1 | | Authors: | Ahn, J.W, Kim, S, Kim, K.J. | | Deposit date: | 2013-08-28 | | Release date: | 2014-03-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.92 Å) | | Cite: | Structural insights into the novel ARM-repeat protein CTNNBL1 and its association with the hPrp19-CDC5L complex
Acta Crystallogr.,Sect.D, 70, 2014
|
|
7CYX
 
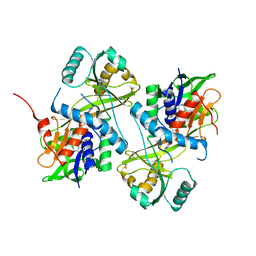 | |
8EZ6
 
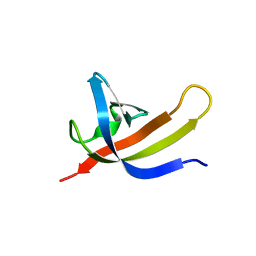 | |
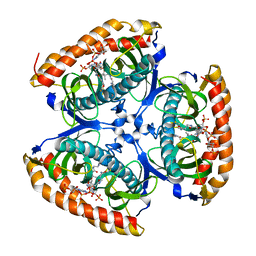
7CZ3
 
 | |
5IN4
 
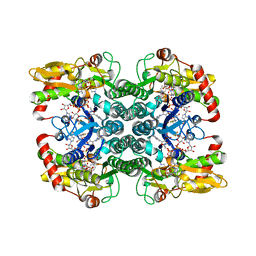 | | Crystal Structure of GDP-mannose 4,6 dehydratase bound to a GDP-fucose based inhibitor | | Descriptor: | GDP-mannose 4,6 dehydratase, GUANOSINE-5'-DIPHOSPHATE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Sickmier, E.A. | | Deposit date: | 2016-03-07 | | Release date: | 2016-08-17 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Facile Modulation of Antibody Fucosylation with Small Molecule Fucostatin Inhibitors and Cocrystal Structure with GDP-Mannose 4,6-Dehydratase.
Acs Chem.Biol., 11, 2016
|
|
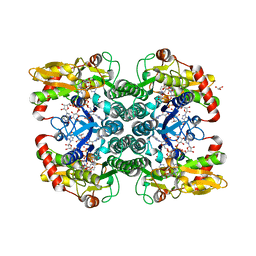
5IN5
 
 | |
7JTB
 
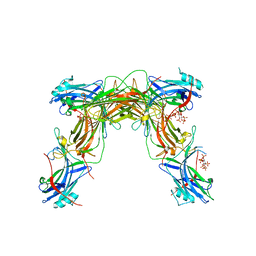 | |
7JSM
 
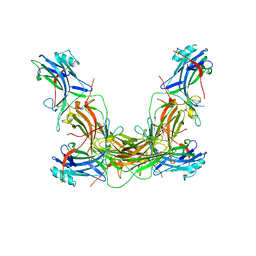 | |
7JXA
 
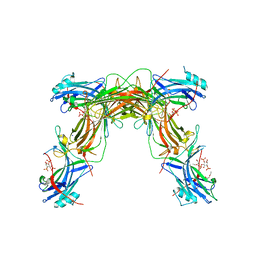 | | CRYSTAL STRUCTURE OF NATIVE BOVINE ARRESTIN 1 IN COMPLEX WITH INOSITOL 1,4,5-TRIPHOSPHATE | | Descriptor: | 2-ETHOXYETHANOL, D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, S-arrestin, ... | | Authors: | Sander, C.L, Palczewski, K, Kiser, P.D. | | Deposit date: | 2020-08-26 | | Release date: | 2021-10-13 | | Last modified: | 2022-02-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural evidence for visual arrestin priming via complexation of phosphoinositols.
Structure, 30, 2022
|
|
6KY2
 
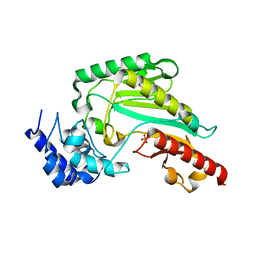 | | Crystal Structure of Arginine Kinase wild type from Daphnia magna | | Descriptor: | Arginine kinase, PHOSPHATE ION | | Authors: | Park, J.H, Rao, Z, Kim, S.Y, Kim, D.S. | | Deposit date: | 2019-09-16 | | Release date: | 2020-09-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Insight into Structural Aspects of Histidine 284 of Daphnia magna Arginine Kinase.
Mol.Cells, 43, 2020
|
|
6KY3
 
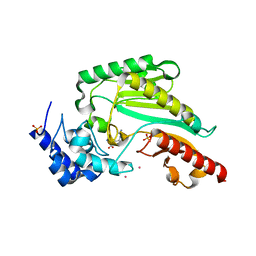 | | Structure of arginine kinase H284A mutant | | Descriptor: | ARGININE, Arginine kinase, PHOSPHATE ION, ... | | Authors: | Rao, Z, Park, J.H, Kim, S.Y, Kim, D.S. | | Deposit date: | 2019-09-16 | | Release date: | 2020-09-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Insight into Structural Aspects of Histidine 284 of Daphnia magna Arginine Kinase.
Mol.Cells, 43, 2020
|
|
