1TNF
 
 | |
4K11
 
 | | The structure of 1NA in complex with Src T338G | | Descriptor: | 1-tert-butyl-3-(naphthalen-1-yl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Eck, M.J, Yun, C.H. | | Deposit date: | 2013-04-04 | | Release date: | 2014-04-09 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The structure of 1NA in complex with Src T338G
To be Published
|
|
2XDQ
 
 | | Dark Operative Protochlorophyllide Oxidoreductase (ChlN-ChlB)2 Complex | | Descriptor: | 1-METHYLGUANIDINE, IRON/SULFUR CLUSTER, LIGHT-INDEPENDENT PROTOCHLOROPHYLLIDE REDUCTASE SUBUNIT B, ... | | Authors: | Broecker, M.J, Schomburg, S, Heinz, D.W, Jahn, D, Schubert, W.-D, Moser, J. | | Deposit date: | 2010-05-06 | | Release date: | 2010-06-16 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of the Nitrogenase-Like Dark Operative Protochlorophyllide Oxidoreductase Catalytic Complex (Chln/Chlb)2.
J.Biol.Chem., 285, 2010
|
|
4WYB
 
 | | Structure of the Bud6 flank domain in complex with actin | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Eck, M.J, Park, E, Zheng, W. | | Deposit date: | 2014-11-17 | | Release date: | 2015-08-19 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.493 Å) | | Cite: | Structure of a Bud6/Actin Complex Reveals a Novel WH2-like Actin Monomer Recruitment Motif.
Structure, 23, 2015
|
|
4ZJV
 | |
8DGT
 
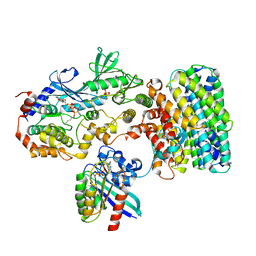 | | Cryo-EM structure of a RAS/RAF complex (state 2) | | Descriptor: | 14-3-3 protein zeta, 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, ... | | Authors: | Eck, M.J, Jeon, H, Park, E, Rawson, S. | | Deposit date: | 2022-06-24 | | Release date: | 2023-07-05 | | Last modified: | 2025-05-14 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Cryo-EM structure of a RAS/RAF recruitment complex.
Nat Commun, 14, 2023
|
|
8DGS
 
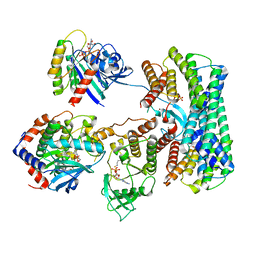 | | Cryo-EM structure of a RAS/RAF complex (state 1) | | Descriptor: | 14-3-3 protein zeta, 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, ... | | Authors: | Eck, M.J, Jeon, H, Park, E, Rawson, S. | | Deposit date: | 2022-06-24 | | Release date: | 2023-07-05 | | Last modified: | 2025-05-14 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Cryo-EM structure of a RAS/RAF recruitment complex.
Nat Commun, 14, 2023
|
|
5U1M
 
 | |
3O4X
 
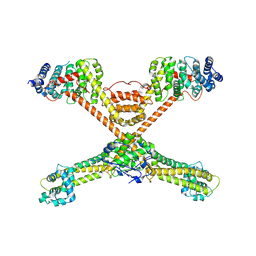 | |
1QZY
 
 | | Human Methionine Aminopeptidase in complex with bengamide inhibitor LAF153 and cobalt | | Descriptor: | (E)-(2R,3R,4S,5R)-3,4,5-TRIHYDROXY-2-METHOXY-8,8-DIMETHYL-NON-6-ENOIC ACID ((3S,6R)-6-HYDROXY-2-OXO-AZEPAN-3-YL)-AMIDE, COBALT (II) ION, Methionine aminopeptidase 2, ... | | Authors: | Eck, M.J, Song, H.K, Morollo, A. | | Deposit date: | 2003-09-18 | | Release date: | 2003-11-25 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Proteomics-based target identification: bengamides as a new class of methionine aminopeptidase inhibitors.
J.Biol.Chem., 278, 2003
|
|
3IKA
 
 | |
1EG4
 
 | | STRUCTURE OF A DYSTROPHIN WW DOMAIN FRAGMENT IN COMPLEX WITH A BETA-DYSTROGLYCAN PEPTIDE | | Descriptor: | BETA-DYSTROGLYCAN, DYSTROPHIN | | Authors: | Huang, X, Poy, F, Zhang, R, Joachimiak, A, Sudol, M, Eck, M.J. | | Deposit date: | 2000-02-11 | | Release date: | 2000-08-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a WW domain containing fragment of dystrophin in complex with beta-dystroglycan.
Nat.Struct.Biol., 7, 2000
|
|
6WXN
 
 | | EGFR(T790M/V948R) in complex with LN3844 | | Descriptor: | CHLORIDE ION, Epidermal growth factor receptor, MAGNESIUM ION, ... | | Authors: | Heppner, D.E, Eck, M.J. | | Deposit date: | 2020-05-11 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Design of a "Two-in-One" Mutant-Selective Epidermal Growth Factor Receptor Inhibitor That Spans the Orthosteric and Allosteric Sites.
J.Med.Chem., 65, 2022
|
|
4WD5
 
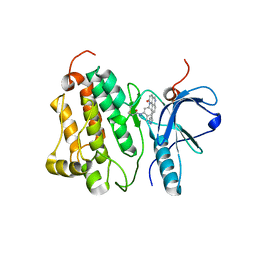 | | Crystal structure of EGFR 696-1022 T790M in complex with QL-X138 | | Descriptor: | CHLORIDE ION, Epidermal growth factor receptor, N-{2-methyl-5-[2-oxo-9-(1H-pyrazol-4-yl)benzo[h][1,6]naphthyridin-1(2H)-yl]phenyl}propanamide | | Authors: | Yun, C.H, Eck, M.J. | | Deposit date: | 2014-09-07 | | Release date: | 2016-05-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal structure of EGFR 696-1022 T790M in complex with QL-X138
To Be Published
|
|
7MYX
 
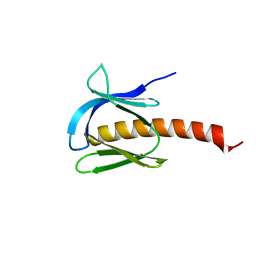 | |
9O0V
 
 | | Crystal structure of CRAF/MEK1 complex with PLX4720, CH5126766, and AMPPNP | | Descriptor: | Dual specificity mitogen-activated protein kinase kinase 1, N-(3-fluoro-4-{[4-methyl-2-oxo-7-(pyrimidin-2-yloxy)-2H-chromen-3-yl]methyl}pyridin-2-yl)-N'-methylsulfuric diamide, N-{3-[(5-chloro-1H-pyrrolo[2,3-b]pyridin-3-yl)carbonyl]-2,4-difluorophenyl}propane-1-sulfonamide, ... | | Authors: | Jang, D.M, Eck, M.J. | | Deposit date: | 2025-04-03 | | Release date: | 2025-10-15 | | Last modified: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Characterization and inhibitor sensitivity of ARAF, BRAF, and CRAF kinases.
J.Biol.Chem., 301, 2025
|
|
9O0U
 
 | | Crystal structure of CRAF/MEK1 complex with PLX4720 and CH5126766 | | Descriptor: | Dual specificity mitogen-activated protein kinase kinase 1, N-(3-fluoro-4-{[4-methyl-2-oxo-7-(pyrimidin-2-yloxy)-2H-chromen-3-yl]methyl}pyridin-2-yl)-N'-methylsulfuric diamide, N-{3-[(5-chloro-1H-pyrrolo[2,3-b]pyridin-3-yl)carbonyl]-2,4-difluorophenyl}propane-1-sulfonamide, ... | | Authors: | Jang, D.M, Eck, M.J. | | Deposit date: | 2025-04-03 | | Release date: | 2025-10-15 | | Last modified: | 2025-11-19 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Characterization and inhibitor sensitivity of ARAF, BRAF, and CRAF kinases.
J.Biol.Chem., 301, 2025
|
|
9ND3
 
 | | JAK2 pseudokinase domain in complex with diaminotriazole inhibitor | | Descriptor: | N-[(1r,4r)-4-{3-[5-amino-3-(4-sulfamoylanilino)-1H-1,2,4-triazole-1-carbonyl]-2,4-difluorophenyl}cyclohexyl]prop-2-enamide, Tyrosine-protein kinase JAK2 | | Authors: | Primi, M.C, Eck, M.J. | | Deposit date: | 2025-02-17 | | Release date: | 2025-11-05 | | Last modified: | 2025-11-12 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | JAK2 pseudokinase domain in complex with diaminotriazole inhibitor
To Be Published
|
|
9ND5
 
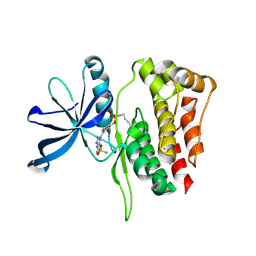 | |
9MMR
 
 | | Cryo-EM structure of CRAF/MEK1/14-3-3 complex (open monomer conformation, CRAF Y340D/Y341D mutant) | | Descriptor: | 14-3-3 protein zeta, 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, ... | | Authors: | Jang, D.M, Jeon, H, Eck, M.J. | | Deposit date: | 2024-12-20 | | Release date: | 2025-09-10 | | Last modified: | 2025-10-08 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Cryo-EM structures of CRAF/MEK1/14-3-3 complexes in autoinhibited and open-monomer states reveal features of RAF regulation.
Nat Commun, 16, 2025
|
|
9MMQ
 
 | | Cryo-EM structure of CRAF/MEK1 complex (kinase domain) | | Descriptor: | 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, MAGNESIUM ION, ... | | Authors: | Jang, D.M, Jeon, H, Eck, M.J. | | Deposit date: | 2024-12-20 | | Release date: | 2025-09-10 | | Last modified: | 2025-10-08 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Cryo-EM structures of CRAF/MEK1/14-3-3 complexes in autoinhibited and open-monomer states reveal features of RAF regulation.
Nat Commun, 16, 2025
|
|
9MMS
 
 | | Cryo-EM structure of CRAF/MEK1 complex (kinase domain, CRAF Y340D/Y341D mutant) | | Descriptor: | 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, MAGNESIUM ION, ... | | Authors: | Jang, D.M, Jeon, H, Eck, M.J. | | Deposit date: | 2024-12-20 | | Release date: | 2025-09-10 | | Last modified: | 2025-10-08 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Cryo-EM structures of CRAF/MEK1/14-3-3 complexes in autoinhibited and open-monomer states reveal features of RAF regulation.
Nat Commun, 16, 2025
|
|
9MMP
 
 | | Cryo-EM structure of CRAF/MEK1/14-3-3 complex (autoinhibited conformation) | | Descriptor: | 14-3-3 protein zeta, 5-[(2-fluoro-4-iodophenyl)amino]-N-(2-hydroxyethoxy)imidazo[1,5-a]pyridine-6-carboxamide, Dual specificity mitogen-activated protein kinase kinase 1, ... | | Authors: | Jang, D.M, Jeon, H, Eck, M.J. | | Deposit date: | 2024-12-20 | | Release date: | 2025-09-10 | | Last modified: | 2025-10-08 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM structures of CRAF/MEK1/14-3-3 complexes in autoinhibited and open-monomer states reveal features of RAF regulation.
Nat Commun, 16, 2025
|
|
4Z16
 
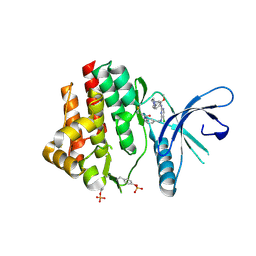 | | Crystal Structure of the Jak3 Kinase Domain Covalently Bound to N-(3-(((5-chloro-2-((2-methoxy-4-(4-methylpiperazin-1-yl)phenyl)amino)pyrimidin-4-yl)amino)methyl)phenyl)acrylamide | | Descriptor: | N-(3-{[(5-chloro-2-{[2-methoxy-4-(4-methylpiperazin-1-yl)phenyl]amino}pyrimidin-4-yl)amino]methyl}phenyl)prop-2-enamide, Tyrosine-protein kinase JAK3 | | Authors: | McNally, R, Tan, L, Gray, N.S, Eck, M.J. | | Deposit date: | 2015-03-26 | | Release date: | 2016-02-10 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Development of Selective Covalent Janus Kinase 3 Inhibitors.
J.Med.Chem., 58, 2015
|
|
8GB4
 
 | |
