4JPH
 
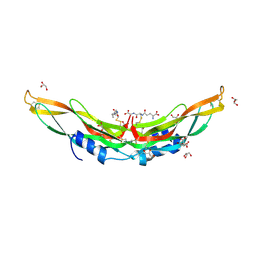 | | Crystal structure of Protein Related to DAN and Cerberus (PRDC) | | Descriptor: | CITRIC ACID, GLUTATHIONE, GLYCEROL, ... | | Authors: | Deng, X, Nolan, K.T, Kattamuri, C, Thompson, T.B. | | Deposit date: | 2013-03-19 | | Release date: | 2013-07-24 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of protein related to dan and cerberus: insights into the mechanism of bone morphogenetic protein antagonism.
Structure, 21, 2013
|
|
8E3G
 
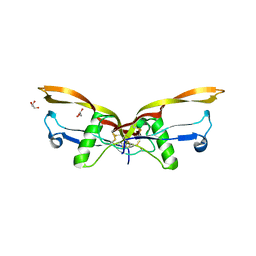 | | BMP2/GDF5 heterodimer | | Descriptor: | Bone morphogenetic protein 2, GLYCEROL, Growth/differentiation factor 5 | | Authors: | Gipson, G.R, Nolan, K.T, Thompson, T.B. | | Deposit date: | 2022-08-17 | | Release date: | 2023-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Formation and characterization of BMP2/GDF5 and BMP4/GDF5 heterodimers.
Bmc Biol., 21, 2023
|
|
6MAA
 
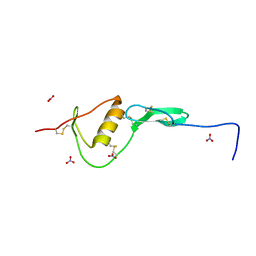 | | WFIKKN2 Follistatin Domain | | Descriptor: | NITRATE ION, WAP, Kazal, ... | | Authors: | McCoy, J.C, Walker, R.G, Thomas, T.B. | | Deposit date: | 2018-08-27 | | Release date: | 2019-03-06 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (1.394 Å) | | Cite: | Crystal structure of the WFIKKN2 follistatin domain reveals insight into how it inhibits growth differentiation factor 8 (GDF8) and GDF11.
J.Biol.Chem., 294, 2019
|
|
7OLY
 
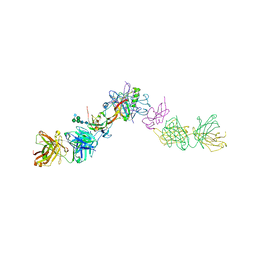 | | Structure of activin A in complex with an ActRIIB-Alk4 fusion reveal insight into activin receptor interactions | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-6)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Activin receptor type-1B, ... | | Authors: | Hakansson, M, Rose, N.C, Castonguay, R, Logan, D.T, Krishnan, L. | | Deposit date: | 2021-05-20 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.265 Å) | | Cite: | Structures of activin ligand traps using natural sets of type I and type II TGF beta receptors.
Iscience, 25, 2022
|
|
1LUC
 
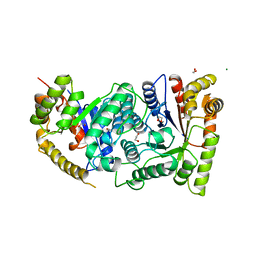 | | BACTERIAL LUCIFERASE | | Descriptor: | 1,2-ETHANEDIOL, BACTERIAL LUCIFERASE, MAGNESIUM ION | | Authors: | Fisher, A.J, Rayment, I. | | Deposit date: | 1996-05-10 | | Release date: | 1996-12-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The 1.5-A resolution crystal structure of bacterial luciferase in low salt conditions.
J.Biol.Chem., 271, 1996
|
|
5UHM
 
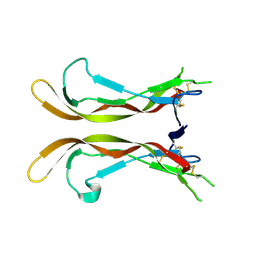 | |
5NTU
 
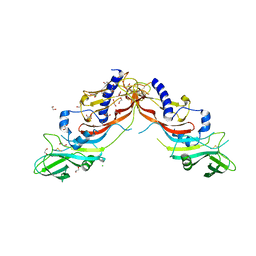 | | Crystal Structure of human Pro-myostatin Precursor at 2.6 A Resolution | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Growth/differentiation factor 8 | | Authors: | Cotton, T.R, Fischer, G, Hyvonen, M. | | Deposit date: | 2017-04-28 | | Release date: | 2018-01-17 | | Last modified: | 2018-02-21 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Structure of the human myostatin precursor and determinants of growth factor latency.
EMBO J., 37, 2018
|
|
5NXS
 
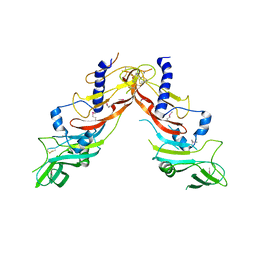 | |
1BSL
 
 | |
1G64
 
 | | THE THREE-DIMENSIONAL STRUCTURE OF ATP:CORRINOID ADENOSYLTRANSFERASE FROM SALMONELLA TYPHIMURIUM. COBALAMIN/ATP TERNARY COMPLEX | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, COB(I)ALAMIN ADENOSYLTRANSFERASE, COBALAMIN, ... | | Authors: | Rayment, I, Escalante-Semerena, J.C, Bauer, C.B. | | Deposit date: | 2000-11-03 | | Release date: | 2000-12-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Three-dimensional structure of ATP:corrinoid adenosyltransferase from Salmonella typhimurium in its free state, complexed with MgATP, or complexed with hydroxycobalamin and MgATP.
Biochemistry, 40, 2001
|
|
1G5R
 
 | | THE THREE-DIMENSIONAL STRUCTURE OF ATP:CORRINOID ADENOSYLTRANSFERASE FROM SALMONELLA TYPHIMURIUM. APO FORM | | Descriptor: | CHLORIDE ION, COB(I)ALAMIN ADENOSYLTRANSFERASE | | Authors: | Rayment, I, Escalante-Semerena, J.C, Bauer, C.B. | | Deposit date: | 2000-11-02 | | Release date: | 2000-11-22 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Three-dimensional structure of ATP:corrinoid adenosyltransferase from Salmonella typhimurium in its free state, complexed with MgATP, or complexed with hydroxycobalamin and MgATP.
Biochemistry, 40, 2001
|
|
1G5T
 
 | | THE THREE-DIMENSIONAL STRUCTURE OF ATP:CORRINOID ADENOSYLTRANSFERASE FROM SALMONELLA TYPHIMURIUM. APO-ATP FORM | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, COB(I)ALAMIN ADENOSYLTRANSFERASE, MAGNESIUM ION | | Authors: | Rayment, I, Escalante-Semerena, J.C, Bauer, C.B. | | Deposit date: | 2000-11-02 | | Release date: | 2000-11-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Three-dimensional structure of ATP:corrinoid adenosyltransferase from Salmonella typhimurium in its free state, complexed with MgATP, or complexed with hydroxycobalamin and MgATP.
Biochemistry, 40, 2001
|
|
