2MT8
 
 | |
2N8E
 | |
2K1V
 
 | | R3/I5 relaxin chimera | | Descriptor: | Insulin-like peptide INSL5, Relaxin-3 | | Authors: | Rosengren, K, Haugaard-Jonsson, L.M. | | Deposit date: | 2008-03-17 | | Release date: | 2008-04-08 | | Last modified: | 2019-12-25 | | Method: | SOLUTION NMR | | Cite: | Structure of the R3/I5 Chimeric Relaxin Peptide, a Selective GPCR135 and GPCR142 Agonist.
J.Biol.Chem., 283, 2008
|
|
2KBC
 
 | |
2K7G
 
 | | Solution Structure of varv F | | Descriptor: | Varv peptide F | | Authors: | Wang, C.K. | | Deposit date: | 2008-08-10 | | Release date: | 2009-02-10 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Combined X-ray and NMR analysis of the stability of the cyclotide cystine knot fold that underpins its insecticidal activity and potential use as a drug scaffold
J.Biol.Chem., 284, 2009
|
|
2KCG
 
 | | Solution structure of cycloviolacin O2 | | Descriptor: | Cycloviolacin-O2 | | Authors: | Wang, C.K. | | Deposit date: | 2008-12-22 | | Release date: | 2009-07-21 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Despite a conserved cystine knot motif, different cyclotides have different membrane binding modes.
Biophys.J., 97, 2009
|
|
2KNP
 | |
2KCH
 
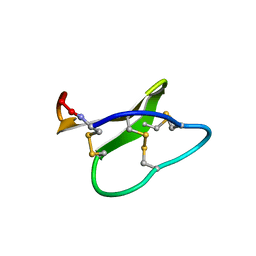 | | Solution structure of micelle-bound kalata B2 | | Descriptor: | Kalata-B2 | | Authors: | Wang, C.K. | | Deposit date: | 2008-12-21 | | Release date: | 2009-07-21 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Despite a conserved cystine knot motif, different cyclotides have different membrane binding modes.
Biophys.J., 97, 2009
|
|
2KVX
 
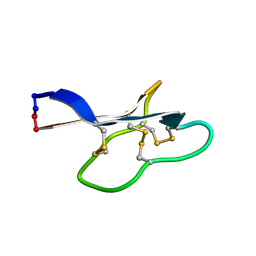 | | Solution structure of kalata B12 | | Descriptor: | Kalata-B12 | | Authors: | Wang, C.K. | | Deposit date: | 2010-03-29 | | Release date: | 2011-03-09 | | Last modified: | 2016-06-01 | | Method: | SOLUTION NMR | | Cite: | The role of conserved Glu residue on cyclotide stability and activity: a structural and functional study of kalata B12, a naturally occurring Glu to Asp mutant.
Biochemistry, 50, 2011
|
|
1AKG
 
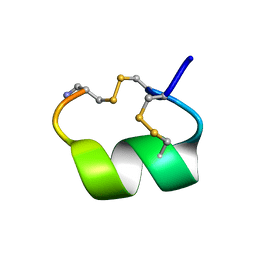 | | ALPHA-CONOTOXIN PNIB FROM CONUS PENNACEUS | | Descriptor: | ALPHA-CONOTOXIN PNIB | | Authors: | Hu, S.-H, Martin, J.L. | | Deposit date: | 1997-05-18 | | Release date: | 1998-05-20 | | Last modified: | 2015-06-10 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structure at 1.1 A resolution of alpha-conotoxin PnIB: comparison with alpha-conotoxins PnIA and GI.
Biochemistry, 36, 1997
|
|
2L1Q
 
 | |
2LWQ
 | |
2LWS
 | |
2LWU
 
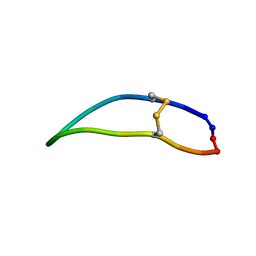 | |
2LWV
 | |
2MH1
 
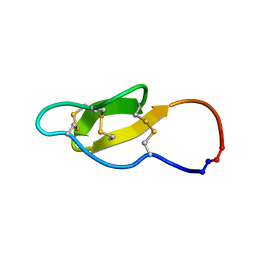 | |
2M7Z
 | | Structure of SmTSP2EC2 | | Descriptor: | CD63-like protein Sm-TSP-2 | | Authors: | Mulvenna, J, Jia, X. | | Deposit date: | 2013-05-02 | | Release date: | 2014-01-22 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Solution structure, membrane interactions, and protein binding partners of the tetraspanin Sm-TSP-2, a vaccine antigen from the human blood fluke Schistosoma mansoni
J.Biol.Chem., 289, 2014
|
|
2N7F
 
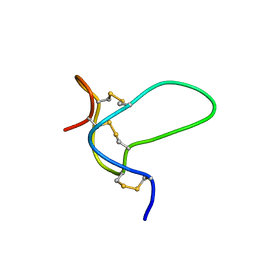 | | NMR solution structure of muO-conotoxin MfVIA | | Descriptor: | muO-conotoxin MfVIA | | Authors: | Schroeder, C.I, Mobli, M. | | Deposit date: | 2015-09-09 | | Release date: | 2016-04-06 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Development of a mu O-Conotoxin Analogue with Improved Lipid Membrane Interactions and Potency for the Analgesic Sodium Channel NaV1.8.
J.Biol.Chem., 291, 2016
|
|
2ND3
 
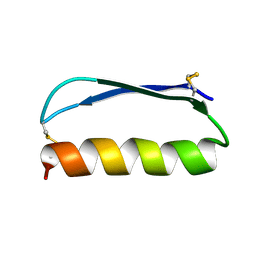 | | Solution structure of the de novo mini protein gEEH_04 | | Descriptor: | De novo mini protein EEH_04 | | Authors: | Pulavarti, S.V, Bahl, C.D, Gilmore, J.M, Eletsky, A, Buchko, G.W, Baker, D, Szyperski, T. | | Deposit date: | 2016-04-22 | | Release date: | 2016-09-21 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Accurate de novo design of hyperstable constrained peptides.
Nature, 538, 2016
|
|
2N9T
 
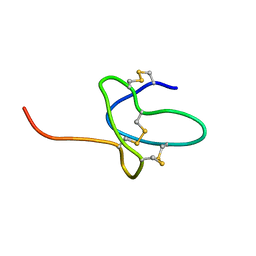 | | NMR solution structure of ProTx-II | | Descriptor: | Beta/omega-theraphotoxin-Tp2a | | Authors: | Schroeder, C.I. | | Deposit date: | 2015-12-08 | | Release date: | 2016-07-06 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Interaction of Tarantula Venom Peptide ProTx-II with Lipid Membranes Is a Prerequisite for Its Inhibition of Human Voltage-gated Sodium Channel NaV1.7.
J.Biol.Chem., 291, 2016
|
|
2ND2
 | | Solution structure of the de novo mini protein gHHH_06 | | Descriptor: | De novo mini protein HHH_06 | | Authors: | Pulavarti, S.V, Eletsky, A, Bahl, C.D, Buchko, G.W, Baker, D, Szyperski, T. | | Deposit date: | 2016-04-22 | | Release date: | 2016-09-21 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Accurate de novo design of hyperstable constrained peptides.
Nature, 538, 2016
|
|
2NB6
 | |
2NB5
 | | NMR solution structure of PawS Derived Peptide 9 (PDP-9) | | Descriptor: | Preproalbumin PawS1 | | Authors: | Armstrong, D.A, Franke, B, Elliott, A.G, Mylne, J.S, Rosengren, K.J. | | Deposit date: | 2016-01-24 | | Release date: | 2016-06-29 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Natural structural diversity within a conserved cyclic peptide scaffold.
Amino Acids, 49, 2017
|
|
2MV1
 | |
2MDQ
 
 | |
