6ZZW
 
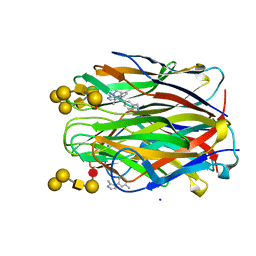 | | Structure of the N terminal domain of Bc2L-C lectin (1-131) in complex with Globo H (H-type 3) and CAS No 912569-62-1 | | Descriptor: | Lectin, SODIUM ION, [3-(2-methylimidazol-1-yl)phenyl]methanamine, ... | | Authors: | Lal, K, Bermeo, R, Imberty, A, Varrot, A. | | Deposit date: | 2020-08-05 | | Release date: | 2021-04-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Prediction and Validation of a Druggable Site on Virulence Factor of Drug Resistant Burkholderia cenocepacia*.
Chemistry, 27, 2021
|
|
7BFY
 
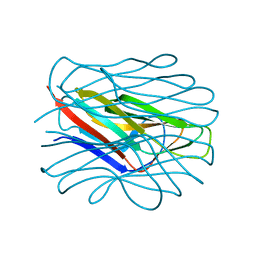 | |
8BLT
 
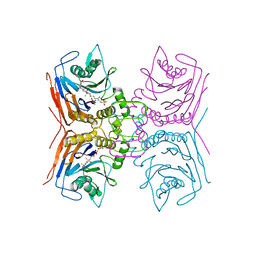 | | Structure of Lactobacillus salivarius (Ls) bile salt hydrolase(BSH) in complex with taurocholate (TCA) | | Descriptor: | Bile salt hydrolase, TAUROCHOLIC ACID | | Authors: | Karlov, D.S, Long, S.L, Zeng, X, Xu, F, Lal, K, Cao, L, Hayoun, K, Lin, J, Joyce, S.A, Tikhonova, I.G. | | Deposit date: | 2022-11-10 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Characterization of the mechanism of bile salt hydrolase substrate specificity by experimental and computational analyses.
Structure, 31, 2023
|
|
8BLS
 
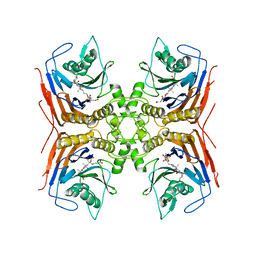 | | Structure of Lactobacillus salivarius (Ls) bile salt hydrolase(BSH) in complex with Glycocholate (GCA) | | Descriptor: | Bile salt hydrolase, GLYCOCHOLIC ACID | | Authors: | Karlov, D.S, Long, S.L, Zeng, X, Xu, F, Lal, K, Cao, L, Hayoun, K, Lin, J, Joyce, S.A, Tikhonova, I.G. | | Deposit date: | 2022-11-10 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Characterization of the mechanism of bile salt hydrolase substrate specificity by experimental and computational analyses.
Structure, 31, 2023
|
|
8E1A
 
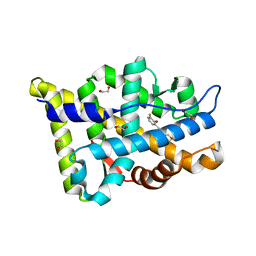 | | Structure-based study to overcome cross-reactivity of novel androgen receptor inhibitors | | Descriptor: | 1,2-ETHANEDIOL, 4-[4-(3-fluoro-2-methoxyphenyl)-1,3-thiazol-2-yl]morpholine, Androgen receptor | | Authors: | Lallous, N, Li, H, Radaeva, M, Dalal, K, Leblanc, E, Ban, F, Ciesielski, F, Chow, B, Morin, M, Singh, K, Rennie, P.S, Cherkasov, A. | | Deposit date: | 2022-08-10 | | Release date: | 2022-09-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structure-Based Study to Overcome Cross-Reactivity of Novel Androgen Receptor Inhibitors.
Cells, 11, 2022
|
|
8QWZ
 
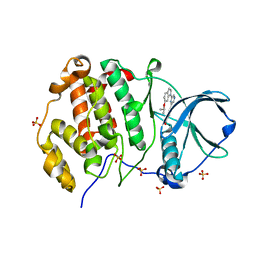 | | Crystal structure of human Casein Kinase II subunit alpha (CK2a1) in complex with 4-(6-(6-isopropoxy-1H-indol-1-yl)pyrazin-2-yl)benzoic acid | | Descriptor: | 4-[6-(6-propan-2-yloxyindol-1-yl)pyrazin-2-yl]benzoic acid, Casein kinase II subunit alpha, SULFATE ION | | Authors: | Kraemer, A, Galal, K, Willson, T, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-10-20 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of human Casein Kinase II subunit alpha (CK2a1) in complex with 4-(6-(6-isopropoxy-1H-indol-1-yl)pyrazin-2-yl)benzoic acid
To Be Published
|
|
8QWY
 
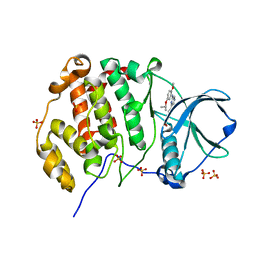 | | Crystal structure of human Casein Kinase II subunit alpha (CK2a1) in complex with 4-(6-((5-isopropoxy-2-methoxyphenyl)amino)pyrazin-2-yl)benzoic acid | | Descriptor: | 4-[6-[(2-methoxy-5-propan-2-yloxy-phenyl)amino]pyrazin-2-yl]benzoic acid, Casein kinase II subunit alpha, SULFATE ION | | Authors: | Kraemer, A, Galal, K, Willson, T, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-10-20 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of human Casein Kinase II subunit alpha (CK2a1) in complex with 4-(6-(6-isopropoxy-1H-indol-1-yl)pyrazin-2-yl)benzoic acid
To Be Published
|
|
3FFV
 
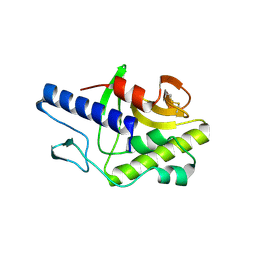 | | Crystal Structure Analysis of Syd | | Descriptor: | Protein syd | | Authors: | Maurus, R, Brayer, G.D. | | Deposit date: | 2008-12-04 | | Release date: | 2009-01-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure, Binding, and Activity of Syd, a SecY-interacting Protein
J.Biol.Chem., 284, 2009
|
|
7OLW
 
 | |
7OLU
 
 | |
7MMO
 
 | | LY-CoV1404 neutralizing antibody against SARS-CoV-2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, LY-CoV1404 Fab heavy chain, LY-CoV1404 Fab light chain, ... | | Authors: | Hendle, J, Pustilnik, A, Sauder, J.M, Coleman, K.A, Boyles, J.S, Dickinson, C.D. | | Deposit date: | 2021-04-30 | | Release date: | 2021-05-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.427 Å) | | Cite: | LY-CoV1404 (bebtelovimab) potently neutralizes SARS-CoV-2 variants.
Biorxiv, 2022
|
|
