7EWU
 
 | |
7EX6
 
 | |
7EX8
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 27 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX9
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 28 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX2
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with L-742,001 (methyl ester form) | | Descriptor: | MANGANESE (II) ION, Replicase, methyl (Z)-4-[4-[(4-chlorophenyl)methyl]-1-(phenylmethyl)piperidin-4-yl]-2-oxidanyl-4-oxidanylidene-but-2-enoate | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX4
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 5 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX7
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 16 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
7EX3
 
 | | Crystal structure of Ebinur Lake virus cap snatching endonuclease in complex with inhibitor 3 | | Descriptor: | CHLORIDE ION, MANGANESE (II) ION, Replicase, ... | | Authors: | Kuang, W, Hu, Z, Gong, P. | | Deposit date: | 2021-05-26 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and Biochemical Basis for Development of Diketo Acid Inhibitors Targeting the Cap-Snatching Endonuclease of the Ebinur Lake Virus (Order: Bunyavirales ).
J.Virol., 96, 2022
|
|
8TBQ
 
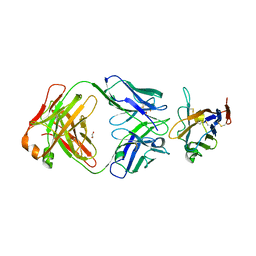 | | The crystal structure of human VISTA extra cellular domain in complex with Fab fragment of pH-selective anti-VISTA antibody | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, SNS-101 Fab heavy chain, ... | | Authors: | van der Horst, E.H, Mukherjee, A, Thisted, T. | | Deposit date: | 2023-06-29 | | Release date: | 2024-03-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.593 Å) | | Cite: | VISTA checkpoint inhibition by pH-selective antibody SNS-101 with optimized safety and pharmacokinetic profiles enhances PD-1 response.
Nat Commun, 15, 2024
|
|
8UV0
 
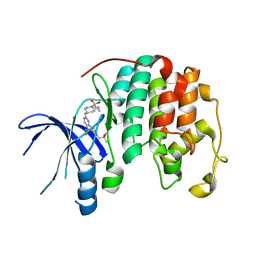 | |
7XXJ
 
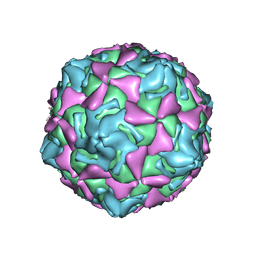 | | Echo 18 incubated with FcRn at pH5.5 | | Descriptor: | PALMITIC ACID, VP1, VP2, ... | | Authors: | Liu, C.C, Qu, X. | | Deposit date: | 2022-05-30 | | Release date: | 2022-06-29 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (3.33 Å) | | Cite: | Human FcRn Is a Two-in-One Attachment-Uncoating Receptor for Echovirus 18.
Mbio, 13, 2022
|
|
7XXA
 
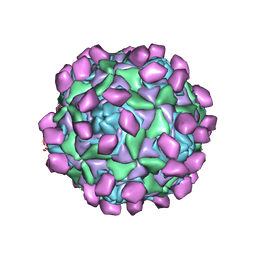 | | Complex of Echo 18 and FcRn at pH7.4 | | Descriptor: | IgG receptor FcRn large subunit p51, VP1, VP2, ... | | Authors: | Liu, C.C, Qu, X. | | Deposit date: | 2022-05-29 | | Release date: | 2022-06-29 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Human FcRn Is a Two-in-One Attachment-Uncoating Receptor for Echovirus 18.
Mbio, 13, 2022
|
|
7XXG
 
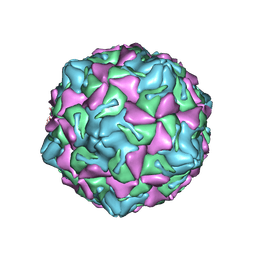 | | Echo 18 at pH5.5 | | Descriptor: | PALMITIC ACID, VP1, VP2, ... | | Authors: | Liu, C.C, Qu, X. | | Deposit date: | 2022-05-30 | | Release date: | 2022-06-29 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (3.37 Å) | | Cite: | Human FcRn Is a Two-in-One Attachment-Uncoating Receptor for Echovirus 18.
Mbio, 13, 2022
|
|
7BQA
 
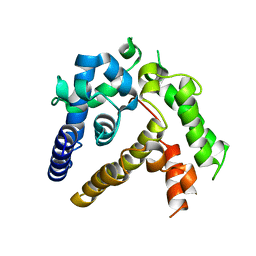 | | Crystal structure of ASFV p35 | | Descriptor: | 60 kDa polyprotein | | Authors: | Li, G.B, Fu, D, Chen, C, Guo, Y. | | Deposit date: | 2020-03-24 | | Release date: | 2020-06-24 | | Last modified: | 2021-05-05 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Crystal structure of the African swine fever virus structural protein p35 reveals its role for core shell assembly.
Protein Cell, 11, 2020
|
|
7F3Q
 
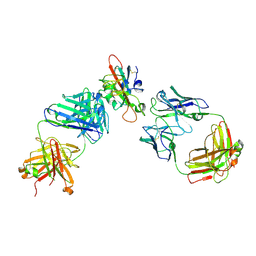 | | SARS-CoV-2 RBD in complex with A5-10 Fab and A34-2 Fab | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of A34-2 Fab, ... | | Authors: | Dou, Y, Wang, X, Liu, P, Lu, B, Wang, K. | | Deposit date: | 2021-06-16 | | Release date: | 2022-06-22 | | Last modified: | 2024-01-24 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Development of neutralizing antibodies against SARS-CoV-2, using a high-throughput single-B-cell cloning method.
Antib Ther, 6, 2023
|
|
7BZG
 
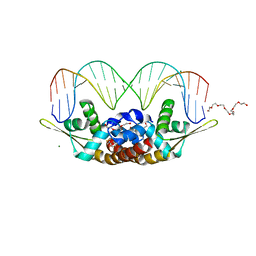 | | Structure of Bacillus subtilis HxlR, wild type in complex with formaldehyde and DNA | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, DI(HYDROXYETHYL)ETHER, DNA (5'-D(*CP*AP*GP*TP*AP*TP*CP*CP*TP*CP*GP*AP*GP*GP*AP*TP*AP*CP*TP*G)-3'), ... | | Authors: | Zhu, R, Chen, P.R. | | Deposit date: | 2020-04-27 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Genetically encoded formaldehyde sensors inspired by a protein intra-helical crosslinking reaction.
Nat Commun, 12, 2021
|
|
7BZD
 
 | | Structure of Bacillus subtilis HxlR, wild type | | Descriptor: | HTH-type transcriptional activator HxlR | | Authors: | Zhu, R, Chen, P.R. | | Deposit date: | 2020-04-27 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.612 Å) | | Cite: | Genetically encoded formaldehyde sensors inspired by a protein intra-helical crosslinking reaction.
Nat Commun, 12, 2021
|
|
7BZE
 
 | | Structure of Bacillus subtilis HxlR, K13A mutant | | Descriptor: | DI(HYDROXYETHYL)ETHER, HTH-type transcriptional activator HxlR | | Authors: | Zhu, R, Chen, P.R. | | Deposit date: | 2020-04-27 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.658 Å) | | Cite: | Genetically encoded formaldehyde sensors inspired by a protein intra-helical crosslinking reaction.
Nat Commun, 12, 2021
|
|
7CJM
 
 | | SARS CoV-2 PLpro in complex with GRL0617 | | Descriptor: | 5-amino-2-methyl-N-[(1R)-1-naphthalen-1-ylethyl]benzamide, Non-structural protein 3, ZINC ION | | Authors: | Fu, Z, Huang, H. | | Deposit date: | 2020-07-11 | | Release date: | 2020-09-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The complex structure of GRL0617 and SARS-CoV-2 PLpro reveals a hot spot for antiviral drug discovery.
Nat Commun, 12, 2021
|
|
7DSY
 
 | | Crystal Structure of RNase L in complex with KM05073 | | Descriptor: | 1-chloranyl-3-methylsulfinyl-6,7-dihydro-5H-2-benzothiophen-4-one, 5'-O-MONOPHOSPHORYLADENYLYL(2'->5')ADENYLYL(2'->5')ADENOSINE, PHOSPHATE ION, ... | | Authors: | Tang, J, Huang, H. | | Deposit date: | 2021-01-03 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.651 Å) | | Cite: | Identification of Small Molecule Inhibitors of RNase L by Fragment-Based Drug Discovery
J.Med.Chem., 65, 2022
|
|
7DTS
 
 | | Crystal structure of RNase L in complex with AC40357 | | Descriptor: | 5'-O-MONOPHOSPHORYLADENYLYL(2'->5')ADENYLYL(2'->5')ADENOSINE, PHOSPHATE ION, Ribonuclease L, ... | | Authors: | Tang, J, Huang, H. | | Deposit date: | 2021-01-06 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Identification of Small Molecule Inhibitors of RNase L by Fragment-Based Drug Discovery
J.Med.Chem., 65, 2022
|
|
7ELW
 
 | | Crystal structure of RNase L in complex with Myricetin | | Descriptor: | 3,5,7-TRIHYDROXY-2-(3,4,5-TRIHYDROXYPHENYL)-4H-CHROMEN-4-ONE, PHOSPHATE ION, Ribonuclease L, ... | | Authors: | Tang, J, Huang, H. | | Deposit date: | 2021-04-12 | | Release date: | 2021-11-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.55 Å) | | Cite: | Identification of Small Molecule Inhibitors of RNase L by Fragment-Based Drug Discovery
J.Med.Chem., 65, 2022
|
|
3UGL
 
 | | Structural and functional characterization of an anesthetic binding site in the second cysteine-rich domain of protein kinase C delta | | Descriptor: | PHOSPHATE ION, Proteine kinase C delta type, ZINC ION, ... | | Authors: | Shanmugasundararaj, S, Stehle, T, Miller, K.W. | | Deposit date: | 2011-11-02 | | Release date: | 2012-12-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.357 Å) | | Cite: | Structural and Functional Characterization of an Anesthetic Binding Site in the Second Cysteine-Rich Domain of Protein Kinase Cdelta
Biophys.J., 103, 2012
|
|
3UFF
 
 | |
3UGI
 
 | | Structural and functional characterization of an anesthetic binding site in the second cysteine-rich domain of protein kinase C delta | | Descriptor: | (methoxymethyl)cyclopropane, PHOSPHATE ION, Protein kinase C delta type, ... | | Authors: | Shanmugasundararaj, S, Stehle, T, Miller, K.W. | | Deposit date: | 2011-11-02 | | Release date: | 2012-12-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.361 Å) | | Cite: | Structural and Functional Characterization of an Anesthetic Binding Site in the Second Cysteine-Rich Domain of Protein Kinase Cdelta
Biophys.J., 103, 2012
|
|
