4B6W
 
 | | Architecture of Trypanosoma brucei Tubulin-Binding cofactor B | | Descriptor: | 1,2-ETHANEDIOL, TUBULIN-SPECIFIC CHAPERONE | | Authors: | Fleming, J.R, Morgan, R.E, Fyfe, P.K, Kelly, S.M, Hunter, W.N. | | Deposit date: | 2012-08-15 | | Release date: | 2012-08-22 | | Last modified: | 2013-07-17 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | The Architecture of Trypanosoma Brucei Tubulin-Binding Cofactor B and Implications for Function.
FEBS J., 280, 2013
|
|
1UNQ
 
 | | High resolution crystal structure of the Pleckstrin Homology Domain Of Protein Kinase B/Akt Bound To Ins(1,3,4,5)-Tetrakisphophate | | Descriptor: | INOSITOL-(1,3,4,5)-TETRAKISPHOSPHATE, RAC-ALPHA SERINE/THREONINE KINASE | | Authors: | Milburn, C.C, Deak, M, Kelly, S.M, Price, N.C, Alessi, D.R, van Aalten, D.M.F. | | Deposit date: | 2003-09-12 | | Release date: | 2004-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (0.98 Å) | | Cite: | Binding of phosphatidylinositol 3,4,5-trisphosphate to the pleckstrin homology domain of protein kinase B induces a conformational change.
Biochem. J., 375, 2003
|
|
1UNP
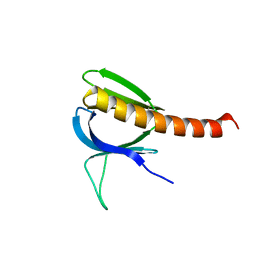 
 | | Crystal structure of the pleckstrin homology domain of PKB alpha | | Descriptor: | RAC-ALPHA SERINE/THREONINE KINASE | | Authors: | Milburn, C.C, Deak, M, Kelly, S.M, Price, N.C, Alessi, D.R, van Aalten, D.M.F. | | Deposit date: | 2003-09-12 | | Release date: | 2004-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Binding of phosphatidylinositol 3,4,5-trisphosphate to the pleckstrin homology domain of protein kinase B induces a conformational change.
Biochem. J., 375, 2003
|
|
1UNR
 
 | | Crystal structure of the PH domain of PKB alpha in complex with a sulfate molecule | | Descriptor: | RAC-ALPHA SERINE/THREONINE KINASE, SULFATE ION | | Authors: | Milburn, C.C, Deak, M, Kelly, S.M, Price, N.C, Alessi, D.R, van Aalten, D.M.F. | | Deposit date: | 2003-09-15 | | Release date: | 2004-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Binding of phosphatidylinositol 3,4,5-trisphosphate to the pleckstrin homology domain of protein kinase B induces a conformational change.
Biochem. J., 375, 2003
|
|
2KYA
 
 | | Solution structure of the leader sequence of the patellamide precursor peptide, PatE1-34 | | Descriptor: | Patellamide protein | | Authors: | Houssen, W.E, Wright, S.H, Kalverda, A.P, Thompson, G.S, Kelly, S.M, Jaspars, M. | | Deposit date: | 2010-05-21 | | Release date: | 2010-09-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Leader Sequence of the Patellamide Precursor Peptide, PatE(1-34).
Chembiochem, 11, 2010
|
|
1OGW
 
 | | Synthetic Ubiquitin with fluoro-Leu at 50 and 67 | | Descriptor: | UBIQUITIN | | Authors: | Alexeev, D, Ramage, R, Young, D.W, Sawyer, L. | | Deposit date: | 2003-05-13 | | Release date: | 2003-05-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Synthesis, Structural and Biological Studies of Ubiquitin Mutants Containing (2S, 4S)-5-Fluoroleucine Residues Strategically Placed in the Hydrophobic Core
Chembiochem, 4, 2003
|
|
7PVC
 
 | | The structure of Kbp.K from E. coli with potassium bound. | | Descriptor: | POTASSIUM ION, Potassium binding protein Kbp | | Authors: | Smith, B.O. | | Deposit date: | 2021-10-01 | | Release date: | 2021-10-13 | | Last modified: | 2023-10-11 | | Method: | SOLUTION NMR | | Cite: | Tuning the Sensitivity of Genetically Encoded Fluorescent Potassium Indicators through Structure-Guided and Genome Mining Strategies.
ACS Sens, 7, 2022
|
|
2JPS
 
 | | NAB2 N-terminal domain | | Descriptor: | Nuclear polyadenylated RNA-binding protein NAB2 | | Authors: | Grant, R, Marshall, N.J, Yang, J, Fasken, M, Kelly, S, Harreman, M.T, Neuhaus, D, Corbett, A.H, Stewart, M. | | Deposit date: | 2007-05-23 | | Release date: | 2008-03-18 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Structure of the N-terminal Mlp1-binding domain of the Saccharomyces cerevisiae mRNA-binding protein, Nab2
J.Mol.Biol., 376, 2008
|
|
3Q79
 
 | |
3Q7A
 
 | | Cryptococcus neoformans protein farnesyltransferase in complex with FPP and L-778,123 | | Descriptor: | (2R)-3-(cyclohexylamino)-2-hydroxypropane-1-sulfonic acid, 4-[(5-{[4-(3-CHLOROPHENYL)-3-OXOPIPERAZIN-1-YL]METHYL}-1H-IMIDAZOL-1-YL)METHYL]BENZONITRILE, FARNESYL DIPHOSPHATE, ... | | Authors: | Hast, M.A, Beese, L.S. | | Deposit date: | 2011-01-04 | | Release date: | 2011-08-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of Cryptococcus neoformans Protein Farnesyltransferase Reveal Strategies for Developing Inhibitors That Target Fungal Pathogens.
J.Biol.Chem., 286, 2011
|
|
3Q78
 
 | |
3Q75
 
 | |
3Q7F
 
 | |
3Q73
 
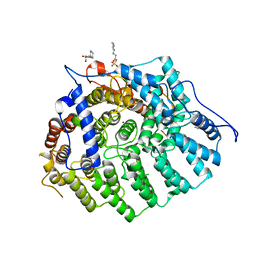 | | Cryptococcus neoformans protein farnesyltransferase, apo enzyme | | Descriptor: | (2S)-3-(cyclohexylamino)-2-hydroxypropane-1-sulfonic acid, Farnesyltransferase, alpha subunit, ... | | Authors: | Hast, M.A, Beese, L.S. | | Deposit date: | 2011-01-04 | | Release date: | 2011-08-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of Cryptococcus neoformans Protein Farnesyltransferase Reveal Strategies for Developing Inhibitors That Target Fungal Pathogens.
J.Biol.Chem., 286, 2011
|
|
3SFX
 
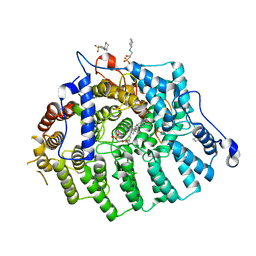 | | Cryptococcus neoformans protein farnesyltransferase in complex with FPT-II and tipifarnib | | Descriptor: | (2R)-3-(cyclohexylamino)-2-hydroxypropane-1-sulfonic acid, 6-[(S)-AMINO(4-CHLOROPHENYL)(1-METHYL-1H-IMIDAZOL-5-YL)METHYL]-4-(3-CHLOROPHENYL)-1-METHYLQUINOLIN-2(1H)-ONE, Cryptococcus neoformans protein farnesyltransferase alpha subunit, ... | | Authors: | Hast, M.A, Beese, L.S. | | Deposit date: | 2011-06-14 | | Release date: | 2011-08-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of Cryptococcus neoformans Protein Farnesyltransferase Reveal Strategies for Developing Inhibitors That Target Fungal Pathogens.
J.Biol.Chem., 286, 2011
|
|
3SFY
 
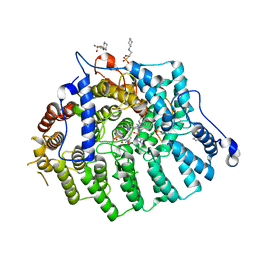 | |
4N58
 
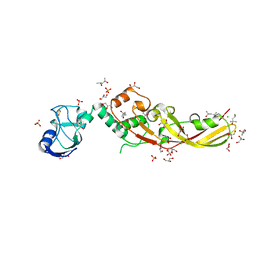 | | Crystal Structure of Pectocin M2 at 1.86 Angstroms | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Grinter, R, Roszak, A.W, Zeth, K, Cogdell, C.J, Walker, D. | | Deposit date: | 2013-10-09 | | Release date: | 2014-06-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structure of the atypical bacteriocin pectocin M2 implies a novel mechanism of protein uptake.
Mol.Microbiol., 93, 2014
|
|
4N59
 
 | | The Crystal Structure of Pectocin M2 at 2.3 Angstroms | | Descriptor: | CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, Pectocin M2, ... | | Authors: | Zeth, K, Grinter, R, Roszak, A.W, Cogdell, R.J, Walker, D. | | Deposit date: | 2013-10-09 | | Release date: | 2014-06-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the atypical bacteriocin pectocin M2 implies a novel mechanism of protein uptake.
Mol.Microbiol., 93, 2014
|
|
4ZHP
 
 | | The crystal structure of Potato ferredoxin I with 2Fe-2S cluster | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, Potato Ferredoxin I | | Authors: | Grinter, R, Josts, I, Roszak, A.W, Cogdell, R.J, Walker, D. | | Deposit date: | 2015-04-26 | | Release date: | 2016-08-31 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structure of the bacterial plant-ferredoxin receptor FusA.
Nat Commun, 7, 2016
|
|
4ZGV
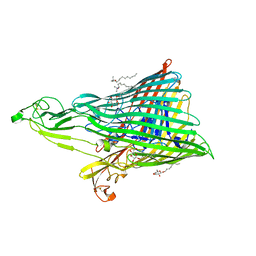 
 | | The Crystal Structure of the Ferredoxin Receptor FusA from Pectobacterium atrosepticum SCRI1043 | | Descriptor: | Ferredoxin receptor, LAURYL DIMETHYLAMINE-N-OXIDE, octyl beta-D-glucopyranoside | | Authors: | Grinter, R, Josts, I, Roszak, A.W, Cogdell, R.J, Walker, D. | | Deposit date: | 2015-04-24 | | Release date: | 2016-08-31 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure of the bacterial plant-ferredoxin receptor FusA.
Nat Commun, 7, 2016
|
|
4ZHO
 
 | | The crystal structure of Arabidopsis ferredoxin 2 with 2Fe-2S cluster | | Descriptor: | CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, Ferredoxin-2, ... | | Authors: | Grinter, R, Josts, I, Roszak, A.W, Cogdell, R.J, Walker, D. | | Deposit date: | 2015-04-26 | | Release date: | 2016-08-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structure of the bacterial plant-ferredoxin receptor FusA.
Nat Commun, 7, 2016
|
|
2V75
 
 | | N-terminal domain of Nab2 | | Descriptor: | NUCLEAR POLYADENYLATED RNA-BINDING PROTEIN NAB2 | | Authors: | Grant, R.P, Marshall, N.J, Stewart, M. | | Deposit date: | 2007-07-26 | | Release date: | 2008-01-29 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of the N-Terminal Mlp1-Binding Domain of the Saccharomyces Cerevisiae Mrna-Binding Protein, Nab2.
J.Mol.Biol., 376, 2008
|
|
1E6C
 
 | | K15M MUTANT OF SHIKIMATE KINASE FROM ERWINIA CHRYSANTHEMI | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, ... | | Authors: | Maclean, J, Krell, T, Coggins, J.R, Lapthorn, A.J. | | Deposit date: | 2000-08-10 | | Release date: | 2001-06-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Biochemical and X-Ray Crystallographic Studies on Shikimate Kinase: The Important Structural Role of the P-Loop Lysine
Protein Sci., 10, 2001
|
|
4D9S
 
 | | Crystal structure of Arabidopsis thaliana UVR8 (UV Resistance locus 8) | | Descriptor: | UVB-resistance protein UVR8 | | Authors: | Arvai, A.S, Christie, J.M, Pratt, A.J, Hitomi, K, Getzoff, E.D. | | Deposit date: | 2012-01-11 | | Release date: | 2012-04-04 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.701 Å) | | Cite: | Plant UVR8 Photoreceptor Senses UV-B by Tryptophan-Mediated Disruption of Cross-Dimer Salt Bridges.
Science, 335, 2012
|
|
4B6M
 
 | |
