2QIC
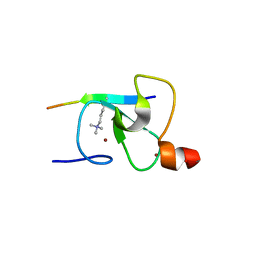 
 | | Crystal Structure of the ING1 PHD Finger in complex with a Histone H3K4ME3 peptide | | Descriptor: | H3K4ME3 PEPTIDE, Inhibitor of growth protein 1, ZINC ION | | Authors: | Pena, P.V, Champagne, K, Zhao, R, Kutateladze, T.G. | | Deposit date: | 2007-07-03 | | Release date: | 2008-05-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Histone H3K4me3 binding is required for the DNA repair and apoptotic activities of ING1 tumor suppressor.
J.Mol.Biol., 380, 2008
|
|
4X34
 
 | |
3C6W
 
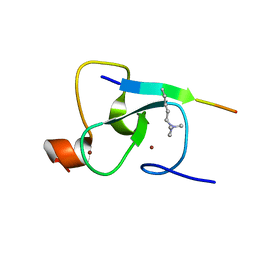
 | | Crystal structure of the ING5 PHD finger in complex with H3K4me3 peptide | | Descriptor: | H3K4me3 histone peptide, Inhibitor of growth protein 5, ZINC ION | | Authors: | Champagne, K.S, Pena, P.V, Johnson, K, Kutateladze, T.G. | | Deposit date: | 2008-02-05 | | Release date: | 2008-06-03 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The crystal structure of the ING5 PHD finger in complex with an H3K4me3 histone peptide.
Proteins, 72, 2008
|
|
2G6Q
 | |
6E83
 
 | |
3RCP
 
 | | Crystal structure of the FAPP1 pleckstrin homology domain | | Descriptor: | GLYCEROL, Pleckstrin homology domain-containing family A member 3 | | Authors: | Roy, S, He, J, Kutateladze, T.G. | | Deposit date: | 2011-03-31 | | Release date: | 2011-04-20 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular Basis of Phosphatidylinositol 4-Phosphate and ARF1 GTPase Recognition by the FAPP1 Pleckstrin Homology (PH) Domain.
J.Biol.Chem., 286, 2011
|
|
8E4V
 
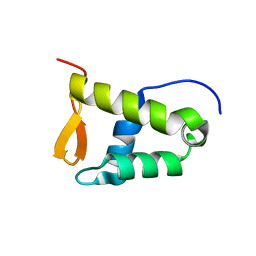 | | Solution structure of the WH domain of MORF | | Descriptor: | Isoform 3 of Histone acetyltransferase KAT6B | | Authors: | Zhang, Y, Kutateladze, T.G. | | Deposit date: | 2022-08-19 | | Release date: | 2023-05-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | MORF and MOZ acetyltransferases target unmethylated CpG islands through the winged helix domain.
Nat Commun, 14, 2023
|
|
7W9V
 
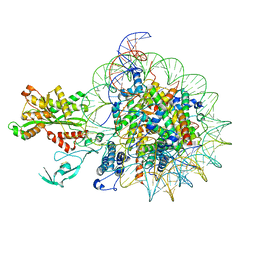 | | Cryo-EM structure of nucleosome in complex with p300 acetyltransferase catalytic core (complex I) | | Descriptor: | DNA (145-MER), Histone H2A type 1-B/E, Histone H2B type 1-J, ... | | Authors: | Hatazawa, S, Liu, J, Takizawa, Y, Zandian, M, Negishi, L, Kutateladze, T.G, Kurumizaka, H. | | Deposit date: | 2021-12-10 | | Release date: | 2022-07-13 | | Method: | ELECTRON MICROSCOPY (3.95 Å) | | Cite: | Structural basis for binding diversity of acetyltransferase p300 to the nucleosome.
Iscience, 25, 2022
|
|
4HCZ
 
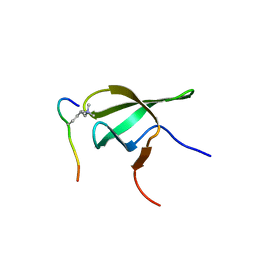 | | PHF1 Tudor in complex with H3K36me3 | | Descriptor: | H3L-like histone, PHD finger protein 1 | | Authors: | Musselman, C.A, Roy, S, Nunez, J, Kutateladze, T.G. | | Deposit date: | 2012-10-01 | | Release date: | 2012-11-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1.
Nat.Struct.Mol.Biol., 19, 2012
|
|
4OUF
 
 | | Crystal Structure of CBP bromodomain | | Descriptor: | 1,2-ETHANEDIOL, CREB-binding protein, DI(HYDROXYETHYL)ETHER | | Authors: | Roy, S, Das, C, Tyler, J.K, Kutateladze, T.G. | | Deposit date: | 2014-02-17 | | Release date: | 2014-03-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Binding of the histone chaperone ASF1 to the CBP bromodomain promotes histone acetylation.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
6X7B
 
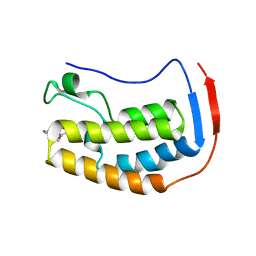 | | BRD4 Bromodomain 1 in complex with multi-action inhibitor SRX3212P | | Descriptor: | 7-[7-oxo-5-(piperazin-1-yl)-7H-thieno[3,2-b]pyran-3-yl]-N-[(pyridin-3-yl)methyl]-2,3-dihydro-1,4-benzodioxine-5-carboxamide, Bromodomain-containing protein 4 | | Authors: | Vann, K.R, Kutateladze, T.G. | | Deposit date: | 2020-05-29 | | Release date: | 2020-08-05 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.951 Å) | | Cite: | Design of thienopyranone-based BET inhibitors that bind multiple synthetic lethality targets.
Sci Rep, 10, 2020
|
|
6X7C
 
 | | BRD4 Bromodomain 1 in complex with multi-action inhibitor SRX3212 | | Descriptor: | 7-[5-(morpholin-4-yl)-7-oxo-7H-thieno[3,2-b]pyran-3-yl]-N-[(pyridin-3-yl)methyl]-2,3-dihydro-1,4-benzodioxine-5-carboxamide, Bromodomain-containing protein 4 | | Authors: | Vann, K.R, Kutateladze, T.G. | | Deposit date: | 2020-05-29 | | Release date: | 2020-08-05 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Design of thienopyranone-based BET inhibitors that bind multiple synthetic lethality targets.
Sci Rep, 10, 2020
|
|
6X7D
 
 | |
7K7T
 
 | | Crystal structure of human MORC4 ATPase-CW in complex with AMPPNP | | Descriptor: | Isoform 3 of MORC family CW-type zinc finger protein 4, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Klein, B.J, Tencer, A.H, Kutateladze, T.G. | | Deposit date: | 2020-09-24 | | Release date: | 2020-11-11 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.94 Å) | | Cite: | Molecular mechanism of the MORC4 ATPase activation.
Nat Commun, 11, 2020
|
|
6U04
 
 | |
8GE0
 
 | |
8GDX
 
 | |
6DS6
 
 | |
6E86
 
 | |
6AXJ
 
 | |
1R48
 
 | | Solution structure of the C-terminal cytoplasmic domain residues 468-497 of Escherichia coli protein ProP | | Descriptor: | Proline/betaine transporter | | Authors: | Zoetewey, D.L, Tripet, B.P, Kutateladze, T.G, Overduin, M.J, Wood, J.M, Hodges, R.S. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-23 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the C-terminal Antiparallel Coiled-coil Domain from Escherichia coli Osmosensor ProP.
J.Mol.Biol., 334, 2003
|
|
5ERC
 
 | |
5D7E
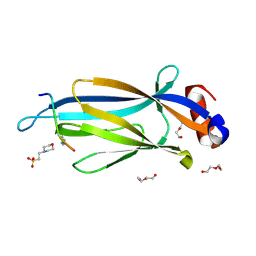 
 | | Crystal structure of Taf14 YEATS domain in complex with H3K9ac | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, H3K9ac, ... | | Authors: | Andrews, F.H, Shanle, E.K, Strahl, B.D, Kutateladze, T.G. | | Deposit date: | 2015-08-13 | | Release date: | 2015-09-23 | | Last modified: | 2019-12-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Association of Taf14 with acetylated histone H3 directs gene transcription and the DNA damage response.
Genes Dev., 29, 2015
|
|
7LKY
 
 | |
7MJU
 
 | |
