4FLC
 
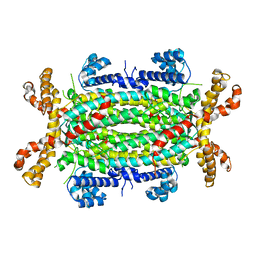 | | Structural and Biochemical Characterization of Human Adenylosuccinate Lyase (ADSL) and the R303C ADSL Deficiency Associated Mutation | | Descriptor: | Adenylosuccinate lyase | | Authors: | Deaton, M.K, Ray, S.P, Capodagli, G.C, Calkins, L.A.F, Sawle, L, Ghosh, K, Patterson, D, Pegan, S.D. | | Deposit date: | 2012-06-14 | | Release date: | 2012-08-01 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and Biochemical Characterization of Human Adenylosuccinate Lyase (ADSL) and the R303C ADSL Deficiency-Associated Mutation.
Biochemistry, 51, 2012
|
|
1LJ0
 
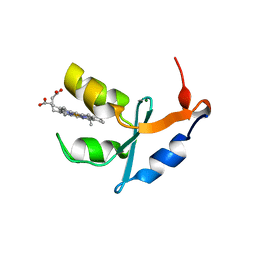 | | Structure of quintuple mutant of the rat outer mitocondrial cytochrome b5. | | Descriptor: | Cytochrome B5 outer mitochondrial membrane isoform, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Cowley, A.B, Altuve, A, Kuchment, O, Terzyan, S, Zhang, X.C, Rivera, M, Benson, D. | | Deposit date: | 2002-04-18 | | Release date: | 2002-11-20 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Toward engineering the stability and hemin binding properties
of microsomal cytochromes b5 into rat outer mitochondrial cytochrome b5:
Examining the influence of residues 25 and 71.
Biochemistry, 41, 2002
|
|
1C2U
 
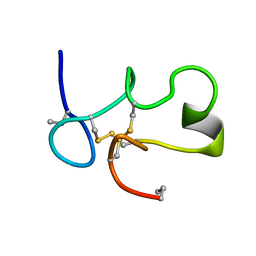 | | SOLUTION STRUCTURE OF [ABU3,35]SHK12-28,17-32 | | Descriptor: | SYNTHETIC PEPTIDE ANALOGUE OF SHK TOXIN | | Authors: | Pennington, M.W, Lanigan, M.D, Kalman, K, Manhir, V.M, Rauer, H, McVaugh, C.T, Behm, D, Donaldson, D, Chandy, K.G, Kem, W.R, Norton, R.S. | | Deposit date: | 1999-07-27 | | Release date: | 1999-11-10 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Role of disulfide bonds in the structure and potassium channel blocking activity of ShK toxin.
Biochemistry, 38, 1999
|
|
6LVF
 
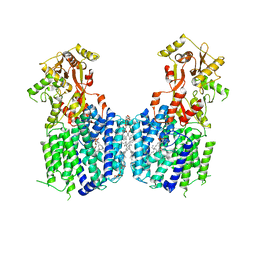 | | Cryo-EM structure of the multiple peptide resistance factor (MprF) loaded with one lysyl-phosphatidylglycerol molecule | | Descriptor: | (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, Low pH-inducible protein LpiA, ... | | Authors: | Song, D.F, Jiao, H.Z, Liu, Z.F. | | Deposit date: | 2020-02-02 | | Release date: | 2021-02-03 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Phospholipid translocation captured in a bifunctional membrane protein MprF.
Nat Commun, 12, 2021
|
|
6LV0
 
 | |
4UP3
 
 | | Crystal structure of the mutant C140S,C286Q thioredoxin reductase from Entamoeba histolytica | | Descriptor: | 1,2-ETHANEDIOL, FLAVIN-ADENINE DINUCLEOTIDE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Podust, L.M, Vieira, D.F. | | Deposit date: | 2014-06-11 | | Release date: | 2015-07-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | X-Ray Structures of Thioredoxin and Thioredoxin Reductase from Entamoeba Histolytica and Prevailing Hypothesis of the Mechanism of Auranofin Action.
J.Struct.Biol., 194, 2016
|
|
7XHG
 
 | |
7V4T
 
 | | Cryo-EM structure of Alphavirus M1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Capsid protein, E1 glycoprotein, ... | | Authors: | Gao, Y, Jia, X, Zhang, Q. | | Deposit date: | 2021-08-15 | | Release date: | 2022-08-17 | | Method: | ELECTRON MICROSCOPY (4.04 Å) | | Cite: | Cryo-EM evidence of viral N-glycosylation reveal receptor binding mechanisms
of alphavirus M1
To Be Published
|
|
7XHF
 
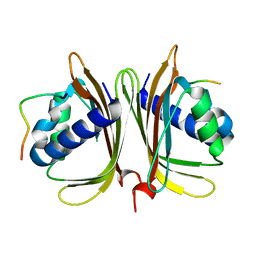 | |
4A5L
 
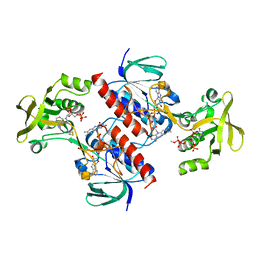 | | Crystal structure of the thioredoxin reductase from Entamoeba histolytica | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION, ... | | Authors: | Podust, L.M. | | Deposit date: | 2011-10-25 | | Release date: | 2012-10-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | X-Ray Structures of Thioredoxin and Thioredoxin Reductase from Entamoeba Histolytica and Prevailing Hypothesis of the Mechanism of Auranofin Action.
J.Struct.Biol., 194, 2016
|
|
4A65
 
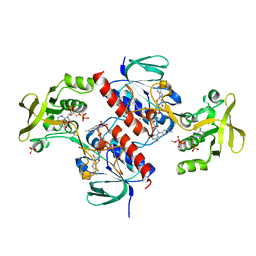 | | Crystal structure of the thioredoxin reductase from Entamoeba histolytica with AuCN | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GOLD ION, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Podust, L.M. | | Deposit date: | 2011-10-31 | | Release date: | 2012-11-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | X-Ray Structures of Thioredoxin and Thioredoxin Reductase from Entamoeba Histolytica and Prevailing Hypothesis of the Mechanism of Auranofin Action.
J.Struct.Biol., 194, 2016
|
|
3MBO
 
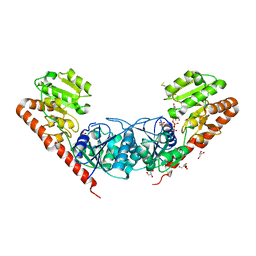 | | Crystal Structure of the Glycosyltransferase BaBshA bound with UDP and L-malate | | Descriptor: | D-MALATE, GLYCEROL, Glycosyltransferase, ... | | Authors: | Wallace, B.D, Claiborne, A, Redinbo, M.R. | | Deposit date: | 2010-03-25 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.307 Å) | | Cite: | Characterization of the N-Acetyl-alpha-d-glucosaminyl
l-Malate Synthase and Deacetylase Functions for Bacillithiol Biosynthesis
in Bacillus anthracis
Biochemistry, 49, 2010
|
|
7DUW
 
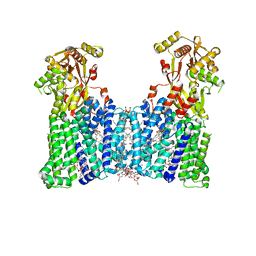 | | Cryo-EM structure of the multiple peptide resistance factor (MprF) loaded with two lysyl-phosphatidylglycerol molecules | | Descriptor: | (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE, (2~{R},3~{S},4~{S},5~{S},6~{S})-2-(hydroxymethyl)-6-[(2~{R},3~{S},4~{R},5~{R},6~{R})-2-(hydroxymethyl)-6-[2-[[(2~{R},3~{S},4~{R},5~{R},6~{S})-6-(hydroxymethyl)-5-[(2~{S},3~{R},4~{S},5~{S},6~{R})-6-(hydroxymethyl)-3,4,5-tris(oxidanyl)oxan-2-yl]oxy-3,4-bis(oxidanyl)oxan-2-yl]oxymethyl]-4-[(1~{R},2~{R},4~{S},5'~{R},6~{R},7~{R},8~{R},9~{S},12~{S},13~{R},16~{S})-5',7,9,13-tetramethylspiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icos-18-ene-6,2'-oxane]-16-yl]oxy-butoxy]-4,5-bis(oxidanyl)oxan-3-yl]oxy-oxane-3,4,5-triol, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, ... | | Authors: | Song, D.F, Jiao, H.Z, Liu, Z.F. | | Deposit date: | 2021-01-12 | | Release date: | 2021-04-21 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (2.96 Å) | | Cite: | Phospholipid translocation captured in a bifunctional membrane protein MprF.
Nat Commun, 12, 2021
|
|
7F9W
 
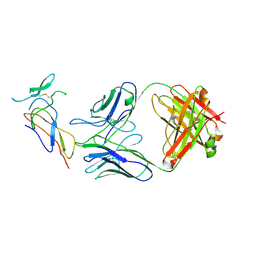 | | CD25 in complex with Fab | | Descriptor: | Heavy chain of Fab, Interleukin-2 receptor subunit alpha, Light chain of Fab | | Authors: | Liu, C. | | Deposit date: | 2021-07-05 | | Release date: | 2022-01-12 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Two novel human anti-CD25 antibodies with antitumor activity inversely related to their affinity and in vitro activity.
Sci Rep, 11, 2021
|
|
7CM4
 
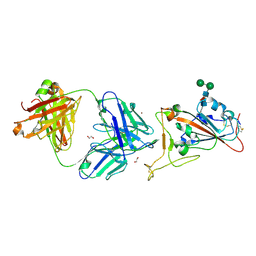 | | Crystal Structure of COVID-19 virus spike receptor-binding domain complexed with a neutralizing antibody CT-P59 | | Descriptor: | 1,2-ETHANEDIOL, IgG heavy chain, IgG light chain, ... | | Authors: | Kim, Y.G, Jeong, J.H, Bae, J.S, Lee, J. | | Deposit date: | 2020-07-24 | | Release date: | 2021-01-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | A therapeutic neutralizing antibody targeting receptor binding domain of SARS-CoV-2 spike protein.
Nat Commun, 12, 2021
|
|
6X81
 
 | |
6X85
 
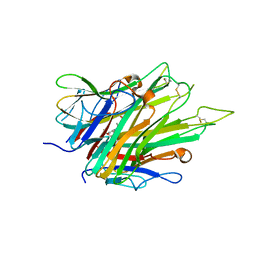 | | Crystal Structure of TNFalpha with indolinone compound 9 | | Descriptor: | 1-{[2-(difluoromethoxy)phenyl]methyl}-2,2-dimethyl-1,2-dihydro-3H-indol-3-one, Tumor necrosis factor | | Authors: | Longenecker, K.L, Stoll, V.S. | | Deposit date: | 2020-06-01 | | Release date: | 2021-01-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Development of Orally Efficacious Allosteric Inhibitors of TNF alpha via Fragment-Based Drug Design.
J.Med.Chem., 64, 2021
|
|
4ZVV
 
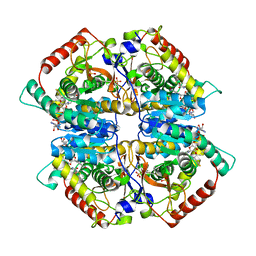 | | Lactate dehydrogenase A in complex with a trisubstituted piperidine-2,4-dione inhibitor GNE-140 | | Descriptor: | (2~{R})-5-(2-chlorophenyl)sulfanyl-2-(4-morpholin-4-ylphenyl)-4-oxidanyl-2-thiophen-3-yl-1,3-dihydropyridin-6-one, L-lactate dehydrogenase A chain, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Li, Y, Chen, Z, Eigenbrot, C. | | Deposit date: | 2015-05-18 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Metabolic plasticity underpins innate and acquired resistance to LDHA inhibition.
Nat.Chem.Biol., 12, 2016
|
|
7S4E
 
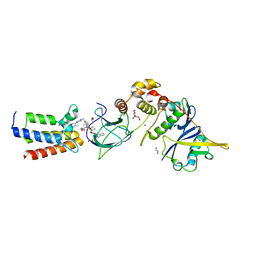 | |
1MPJ
 
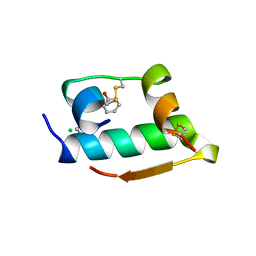 | | X-RAY CRYSTALLOGRAPHIC STUDIES ON HEXAMERIC INSULINS IN THE PRESENCE OF HELIX-STABILIZING AGENTS, THIOCYANATE, METHYLPARABEN AND PHENOL | | Descriptor: | CHLORIDE ION, PHENOL, PHENOL INSULIN, ... | | Authors: | Whittingham, J.L, Dodson, E.J, Moody, P.C.E, Dodson, G.G. | | Deposit date: | 1995-09-13 | | Release date: | 1996-01-29 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | X-ray crystallographic studies on hexameric insulins in the presence of helix-stabilizing agents, thiocyanate, methylparaben, and phenol.
Biochemistry, 34, 1995
|
|
6RZE
 
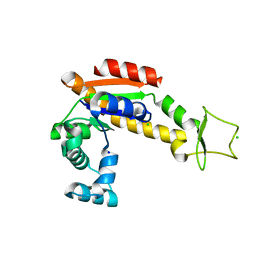 | | Crystal structure of E. coli Adenylate kinase R119A mutant | | Descriptor: | Adenylate kinase, CHLORIDE ION, SODIUM ION | | Authors: | Grundstrom, C, Rogne, P, Wolf-Watz, M, Sauer-Eriksson, A.E. | | Deposit date: | 2019-06-13 | | Release date: | 2019-08-07 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Nucleation of an Activating Conformational Change by a Cation-pi Interaction.
Biochemistry, 58, 2019
|
|
6XRL
 
 | | Crystal structure of human PI3K-gamma in complex with inhibitor IPI-549 | | Descriptor: | 2-amino-N-[(1S)-1-{8-[(1-methyl-1H-pyrazol-4-yl)ethynyl]-1-oxo-2-phenyl-1,2-dihydroisoquinolin-3-yl}ethyl]pyrazolo[1,5-a]pyrimidine-3-carboxamide, Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform, SULFATE ION | | Authors: | Walker, N.P, Jeffrey, J.L. | | Deposit date: | 2020-07-13 | | Release date: | 2020-09-16 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Discovery of Potent and Selective PI3K gamma Inhibitors.
J.Med.Chem., 63, 2020
|
|
1EAW
 
 | |
1EAX
 
 | | Crystal structure of MTSP1 (matriptase) | | Descriptor: | BENZAMIDINE, SULFATE ION, SUPPRESSOR OF TUMORIGENICITY 14 | | Authors: | Friedrich, R, Bode, W. | | Deposit date: | 2001-07-17 | | Release date: | 2002-01-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Catalytic Domain Structures of Mt-Sp1/Matriptase, a Matrix-Degrading Transmembrane Serine Proteinase.
J.Biol.Chem., 277, 2002
|
|
1EV6
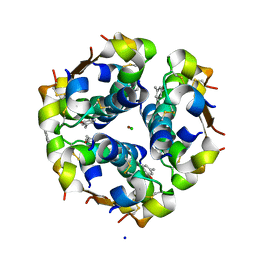 
 | | Structure of the monoclinic form of the M-cresol/insulin R6 hexamer | | Descriptor: | CHLORIDE ION, INSULIN, M-CRESOL, ... | | Authors: | Smith, G.D, Ciszak, E, Magrum, L.A, Pangborn, W.A, Blessing, R.H. | | Deposit date: | 2000-04-19 | | Release date: | 2000-12-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | R6 Hexameric Insulin Complexed with m-Cresol or Resorcinol
Biochem.Biophys.Res.Commun., 56, 2000
|
|
