5GIE
 
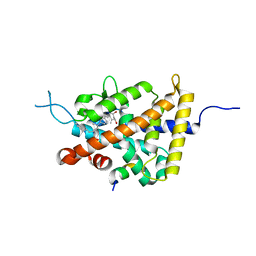 | | Crystal structure of VDR in complex with DLAM-4P (P21 form) | | Descriptor: | (3R,5S)-5-[(2R)-2-[(1R,3aS,4E,7aR)-7a-methyl-4-[(2Z)-2-[(3S,5R)-2-methylidene-3,5-bis(oxidanyl)cyclohexylidene]ethylidene]-2,3,3a,5,6,7-hexahydro-1H-inden-1-yl]propyl]-3-methyl-3-oxidanyl-1-(4-phenylbutyl)pyrrolidin-2-one, SRC1, Vitamin D3 receptor | | Authors: | Asano, L, Shimizu, T. | | Deposit date: | 2016-06-23 | | Release date: | 2016-12-07 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.387 Å) | | Cite: | Regulation of the vitamin D receptor by vitamin D lactam derivatives.
Febs Lett., 590, 2016
|
|
5GID
 
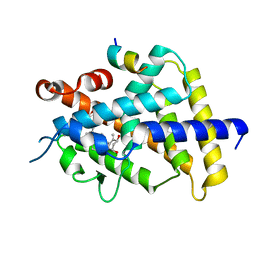 | | Crystal structure of VDR in complex with DLAM-4 (C2 form) | | Descriptor: | (3R,5S)-5-[(2R)-2-[(1R,3aS,4E,7aR)-7a-methyl-4-[(2Z)-2-[(3S,5R)-2-methylidene-3,5-bis(oxidanyl)cyclohexylidene]ethylidene]-2,3,3a,5,6,7-hexahydro-1H-inden-1-yl]propyl]-3-methyl-3-oxidanyl-1-(4-phenylbutyl)pyrrolidin-2-one, SRC1, Vitamin D3 receptor | | Authors: | Asano, L, Shimizu, T. | | Deposit date: | 2016-06-23 | | Release date: | 2016-12-07 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | Regulation of the vitamin D receptor by vitamin D lactam derivatives.
Febs Lett., 590, 2016
|
|
5GIC
 
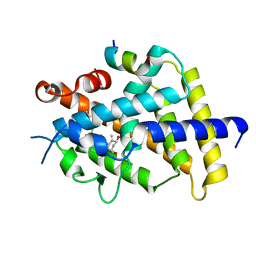 | | Crystal structure of VDR in complex with DLAM-2P | | Descriptor: | (3~{R},5~{S})-5-[(2~{R})-2-[(1~{R},3~{a}~{S},4~{Z},7~{a}~{R})-7~{a}-methyl-4-[(2~{E})-2-[(3~{S},5~{R})-2-methylidene-3,5-bis(oxidanyl)cyclohexylidene]ethylidene]-2,3,3~{a},5,6,7-hexahydro-1~{H}-inden-1-yl]propyl]-3-methyl-3-oxidanyl-1-(2-phenylethyl)pyrrolidin-2-one, SRC1, Vitamin D3 receptor | | Authors: | Shimizu, T, Asano, L. | | Deposit date: | 2016-06-23 | | Release date: | 2016-12-07 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.352 Å) | | Cite: | Regulation of the vitamin D receptor by vitamin D lactam derivatives.
Febs Lett., 590, 2016
|
|
6M4I
 
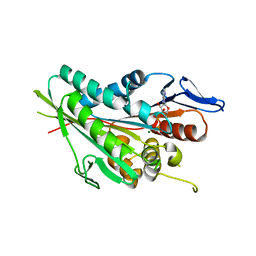 | |
3VI8
 
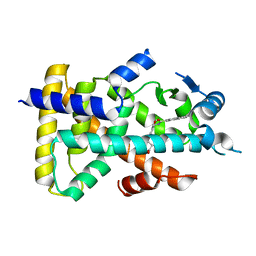 | | Human PPAR alpha ligand binding domain in complex with a synthetic agonist APHM13 | | Descriptor: | (2S)-2-(4-methoxy-3-{[(pyren-1-ylcarbonyl)amino]methyl}benzyl)butanoic acid, Peroxisome proliferator-activated receptor alpha | | Authors: | Oyama, T, Miyachi, H, Morikawa, K. | | Deposit date: | 2011-09-25 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Peroxisome proliferator-activated receptors (PPARs) have multiple binding points that accommodate ligands in various conformations: phenylpropanoic acid-type PPAR ligands bind to PPAR in different conformations, depending on the subtype
J.Med.Chem., 55, 2012
|
|
3VJI
 
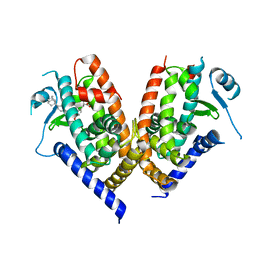 | | Human PPAR gamma ligand binding domain in complex with JKPL53 | | Descriptor: | (2S)-2-{4-butoxy-3-[({4-[(3S,5S,7S)-tricyclo[3.3.1.1~3,7~]dec-1-yl]benzoyl}amino)methyl]benzyl}butanoic acid, Peroxisome proliferator-activated receptor gamma | | Authors: | Tomioka, D, Kuwabara, N, Hashimoto, H, Sato, M, Shimizu, T. | | Deposit date: | 2011-10-20 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Peroxisome proliferator-activated receptors (PPARs) have multiple binding points that accommodate ligands in various conformations: phenylpropanoic acid-type PPAR ligands bind to PPAR in different conformations, depending on the subtype.
J.Med.Chem., 55, 2012
|
|
3VJH
 
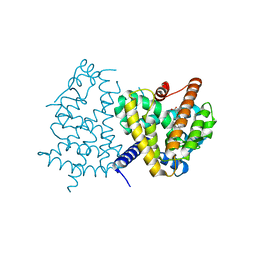 | | Human PPAR GAMMA ligand binding domain in complex with JKPL35 | | Descriptor: | (2S)-2-[4-methoxy-3-({[4-(trifluoromethyl)benzoyl]amino}methyl)benzyl]pentanoic acid, Peroxisome proliferator-activated receptor gamma | | Authors: | Tomioka, D, Kuwabara, N, Hashimoto, H, Sato, M, Shimizu, T. | | Deposit date: | 2011-10-20 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Peroxisome proliferator-activated receptors (PPARs) have multiple binding points that accommodate ligands in various conformations: phenylpropanoic acid-type PPAR ligands bind to PPAR in different conformations, depending on the subtype.
J.Med.Chem., 55, 2012
|
|
3WPN
 
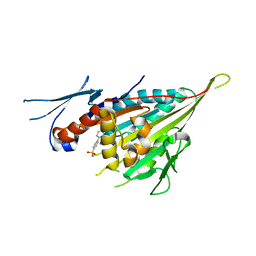 | |
1BHI
 
 | | STRUCTURE OF TRANSACTIVATION DOMAIN OF CRE-BP1/ATF-2, NMR, 20 STRUCTURES | | Descriptor: | CRE-BP1 | | Authors: | Nagadoi, A, Nakazawa, K, Uda, H, Maekawa, T, Ishii, S, Nishimura, Y. | | Deposit date: | 1998-06-09 | | Release date: | 1999-06-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the transactivation domain of ATF-2 comprising a zinc finger-like subdomain and a flexible subdomain.
J.Mol.Biol., 287, 1999
|
|
