8P01
 
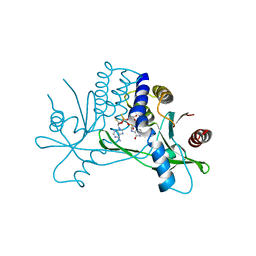 | | Crystal structure of human STING ectodomain in complex with BI 7446, a potent cyclic dinucleotide STING agonist with broad-spectrum variant activity for the treatment of cancer | | Descriptor: | 3-[(1~{R},3~{R},6~{R},8~{R},9~{R},10~{R},12~{R},15~{R},17~{R},18~{R})-8-(6-aminopurin-9-yl)-9-fluoranyl-18-oxidanyl-3,12-bis(oxidanylidene)-3,12-bis(sulfanyl)-2,4,7,11,13,16-hexaoxa-3$l^{5},12$l^{5}-diphosphatricyclo[13.2.1.0^{6,10}]octadecan-17-yl]-6~{H}-imidazo[4,5-d]pyridazin-7-one, Stimulator of interferon genes protein | | Authors: | Nar, H. | | Deposit date: | 2023-05-09 | | Release date: | 2023-07-26 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.094 Å) | | Cite: | Discovery of BI 7446: A Potent Cyclic Dinucleotide STING Agonist with Broad-Spectrum Variant Activity for the Treatment of Cancer.
J.Med.Chem., 66, 2023
|
|
5F2P
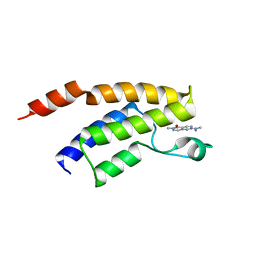 
 | | Crystal structure of the BRD9 bromodomain in complex with compound 3. | | Descriptor: | 2-(dimethylamino)-6-methyl-pyrido[4,3-d]pyrimidin-5-one, BRD9 | | Authors: | Nar, H, Fiegen, D, Zoephel, A, Bader, G. | | Deposit date: | 2015-12-02 | | Release date: | 2016-03-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-Based Design of an in Vivo Active Selective BRD9 Inhibitor.
J.Med.Chem., 59, 2016
|
|
3E5V
 
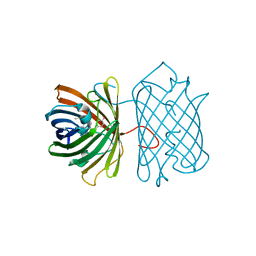 | | Crystal Structure Analysis of eqFP611 Double Mutant T122R, N143S | | Descriptor: | Red fluorescent protein eqFP611 | | Authors: | Nar, H, Nienhaus, K, Nienhaus, U, Wiedenmann, J. | | Deposit date: | 2008-08-14 | | Release date: | 2008-09-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Trans-cis isomerization is responsible for the red-shifted fluorescence in variants of the red fluorescent protein eqFP611.
J.Am.Chem.Soc., 130, 2008
|
|
3E5T
 
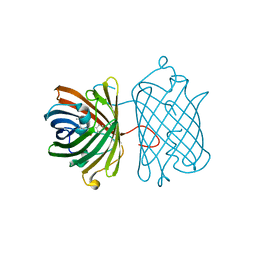 | | Crystal Structure Analysis of FP611 | | Descriptor: | Red fluorescent protein eqFP611 | | Authors: | Nar, H, Nienhaus, K, Nienhaus, U, Wiedenmann, J. | | Deposit date: | 2008-08-14 | | Release date: | 2008-09-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Trans-cis isomerization is responsible for the red-shifted fluorescence in variants of the red fluorescent protein eqFP611.
J.Am.Chem.Soc., 130, 2008
|
|
5LLS
 
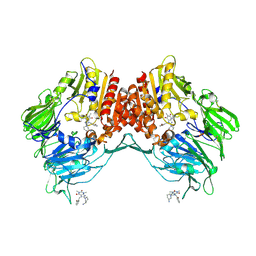 | | Porcine dipeptidyl peptidase IV in complex with 8-(3-aminopiperidin-1-yl)-7-[(2-bromophenyl)methyl]-1,3-dimethyl-2,3,6,7-tetrahydro-1H-purine-2,6-dione | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 8-[(3~{R})-3-azanylpiperidin-1-yl]-7-[(2-bromophenyl)methyl]-1,3-dimethyl-purine-2,6-dione, ... | | Authors: | Nar, H, Blaesse, M. | | Deposit date: | 2016-07-28 | | Release date: | 2016-09-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Comparative Analysis of Binding Kinetics and Thermodynamics of Dipeptidyl Peptidase-4 Inhibitors and Their Relationship to Structure.
J.Med.Chem., 59, 2016
|
|
4K8B
 
 | | Crystal structure of HCV NS3/4A protease complexed with inhibitor | | Descriptor: | N-(tert-butylcarbamoyl)-3-methyl-L-valyl-(4R)-N-[(1R,2S)-1-carboxy-2-ethenylcyclopropyl]-4-[(7-methoxy-2-phenylquinolin-4-yl)oxy]-L-prolinamide, NS3 protease, Nonstructural protein, ... | | Authors: | Nar, H. | | Deposit date: | 2013-04-18 | | Release date: | 2014-03-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Ligand bioactive conformation plays a critical role in the design of drugs that target the hepatitis C virus NS3 protease.
J.Med.Chem., 57, 2014
|
|
1ZUX
 
 | | EosFP Fluorescent Protein- Green Form | | Descriptor: | green to red photoconvertible GPF-like protein EosFP | | Authors: | Nar, H, Nienhaus, K, Wiedenmann, J, Nienhaus, G.U. | | Deposit date: | 2005-06-01 | | Release date: | 2005-06-28 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural basis for photo-induced protein cleavage and green-to-red conversion of fluorescent protein EosFP.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
3AZU
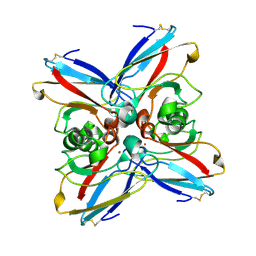 
 | |
2BTJ
 | | Fluorescent Protein EosFP - red form | | Descriptor: | Green to red photoconvertible GFP-like protein EosFP | | Authors: | Nar, H, Nienhaus, K, Wiedenmann, J, Nienhaus, G.U. | | Deposit date: | 2005-06-01 | | Release date: | 2005-06-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Photo-Induced Protein Cleavage and Green-to-Red Conversion of Fluorescent Protein Eosfp.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1KTT
 
 | | Thrombin inhibitor complex | | Descriptor: | 4-(5-BENZENESULFONYLAMINO-1-METHYL-1H-BENZOIMIDAZOL-2-YLMETHYL)-BENZAMIDINE, hirudin IIB, thrombin | | Authors: | Nar, H. | | Deposit date: | 2002-01-17 | | Release date: | 2002-02-06 | | Last modified: | 2013-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-based design of novel potent nonpeptide thrombin inhibitors.
J.Med.Chem., 45, 2002
|
|
1KTS
 
 | | Thrombin Inhibitor Complex | | Descriptor: | 3-({2-[(4-CARBAMIMIDOYL-PHENYLAMINO)-METHYL]-3-METHYL-3H-BENZOIMIDAZOLE-5-CARBONYL}-PYRIDIN-2-YL-AMINO)-PROPIONIC ACID ETHYL ESTER, hirudin IIB, thrombin | | Authors: | Nar, H. | | Deposit date: | 2002-01-17 | | Release date: | 2002-02-06 | | Last modified: | 2013-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure-based design of novel potent nonpeptide thrombin inhibitors.
J.Med.Chem., 45, 2002
|
|
7WJT
 
 | |
4C1W
 
 | | Carbohydrate binding domain from Streptococcus pneumoniae NanA sialidase complexed with 3'-sialyllactose | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose, NEURAMINIDASE | | Authors: | Yang, L, Connaris, H, Potter, J.A, Taylor, G.L. | | Deposit date: | 2013-08-14 | | Release date: | 2014-08-27 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Prevention of Influenza by Targeting Host Receptors Using Engineered Proteins.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
2K0N
 
 | |
1ORL
 
 | | 1H NMR structure determination of Viscotoxin C1 | | Descriptor: | Viscotoxin C1 | | Authors: | Molinari, H, Romagnoli, S, Fogolari, F, Catalano, M, Urech, K, Giannattasio, M, Ragona, L. | | Deposit date: | 2003-03-14 | | Release date: | 2003-04-01 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of viscotoxin C1 from Viscum album species Coloratum ohwi: toward a structure-function analysis of viscotoxins.
Biochemistry, 42, 2003
|
|
9FE6
 
 | | Short-chain dehydrogenase/reductase (SDR) from Thermus caliditerrae | | Descriptor: | MAGNESIUM ION, SDR family oxidoreductase | | Authors: | Kapur, B, Nar, H. | | Deposit date: | 2024-05-17 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.875 Å) | | Cite: | In silico enzyme screening identifies an SDR ketoreductase from Thermus caliditerrae as an attractive biocatalyst and promising candidate for protein engineering
Front Chem Biol, 2024
|
|
9FEB
 
 | |
1ILS
 
 | | X-RAY CRYSTAL STRUCTURE THE TWO SITE-SPECIFIC MUTANTS ILE7SER AND PHE110SER OF AZURIN FROM PSEUDOMONAS AERUGINOSA | | Descriptor: | AZURIN, COPPER (II) ION, NITRATE ION | | Authors: | Hammann, C, Nar, H, Huber, R, Messerschmidt, A. | | Deposit date: | 1995-10-12 | | Release date: | 1996-03-08 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray crystal structure of the two site-specific mutants Ile7Ser and Phe110Ser of azurin from Pseudomonas aeruginosa.
J.Mol.Biol., 255, 1996
|
|
6ZLE
 
 | |
8C7H
 
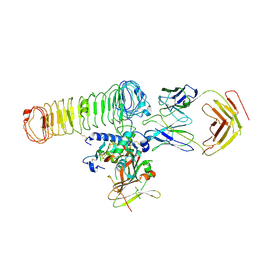 | | Cryo-EM Map of the latTGF-beta 28G11 Fab complex | | Descriptor: | 28G11 Fab heavy chain, 28G11 Fab light chain, Transforming growth factor beta activator LRRC32, ... | | Authors: | Ebenhoch, R, Nar, H. | | Deposit date: | 2023-01-16 | | Release date: | 2023-03-01 | | Last modified: | 2023-03-29 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Anti-GARP Antibodies Inhibit Release of TGF-beta by Regulatory T Cells via Different Modes of Action, but Do Not Influence Their Function In Vitro.
Immunohorizons, 7, 2023
|
|
7NM5
 | | Solution structure of MLKL executioner domain in complex with a fragment | | Descriptor: | 2-[(~{S})-methoxy-(4-phenylphenyl)methyl]-3~{H}-benzimidazole-5-carboxylic acid, Mixed lineage kinase domain-like protein | | Authors: | Ruebbelke, M, Bauer, M, Hamilton, J, Binder, F, Nar, H, Zeeb, M. | | Deposit date: | 2021-02-23 | | Release date: | 2021-09-22 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Discovery and Structure-Based Optimization of Fragments Binding the Mixed Lineage Kinase Domain-like Protein Executioner Domain.
J.Med.Chem., 64, 2021
|
|
7NM2
 
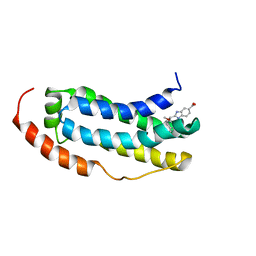 | | Solution structure of MLKL executioner domain in complex with a fragment | | Descriptor: | 2-[(~{S})-methoxy-(4-propan-2-ylphenyl)methyl]-3~{H}-benzimidazole-5-carboxylic acid, Mixed lineage kinase domain-like protein | | Authors: | Ruebbelke, M, Bauer, M, Hamilton, J, Binder, F, Nar, H, Zeeb, M. | | Deposit date: | 2021-02-23 | | Release date: | 2021-09-22 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Discovery and Structure-Based Optimization of Fragments Binding the Mixed Lineage Kinase Domain-like Protein Executioner Domain.
J.Med.Chem., 64, 2021
|
|
7NM4
 | | Solution structure of MLKL executioner domain in complex with a fragment | | Descriptor: | (~{S})-1~{H}-benzimidazol-2-yl-(4-propan-2-ylphenyl)methanol, Mixed lineage kinase domain-like protein | | Authors: | Ruebbelke, M, Bauer, M, Hamilton, J, Binder, F, Nar, H, Zeeb, M. | | Deposit date: | 2021-02-23 | | Release date: | 2021-09-22 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Discovery and Structure-Based Optimization of Fragments Binding the Mixed Lineage Kinase Domain-like Protein Executioner Domain.
J.Med.Chem., 64, 2021
|
|
6Z86
 | |
6Z88
 
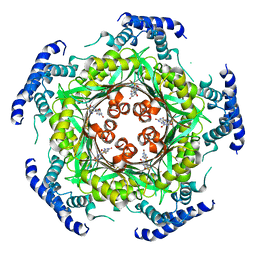 | | human GTP cyclohydrolase I in complex with allosteric inhibitor | | Descriptor: | 5-azanyl-[1,3]thiazolo[5,4-d]pyrimidine-2,7-dione, GTP cyclohydrolase 1, ZINC ION | | Authors: | Ebenhoch, R, Nar, H. | | Deposit date: | 2020-06-02 | | Release date: | 2020-12-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.687 Å) | | Cite: | A hybrid approach reveals the allosteric regulation of GTP cyclohydrolase I.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
