3QGN
 
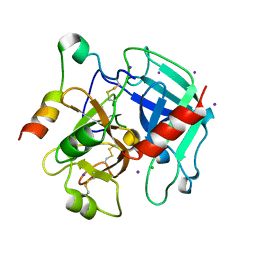 | | The allosteric E*-E equilibrium is a key property of the trypsin fold | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, IODIDE ION, ... | | 著者 | Niu, W, Gohara, D, Chen, Z, Di Cera, E. | | 登録日 | 2011-01-24 | | 公開日 | 2011-07-06 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Crystallographic and kinetic evidence of allostery in a trypsin-like protease.
Biochemistry, 50, 2011
|
|
3JZ1
 
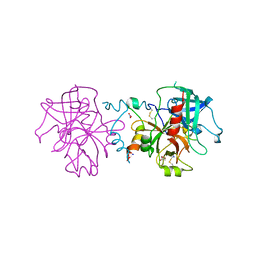 | | Crystal structure of human thrombin mutant N143P in E:Na+ form | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, NITRATE ION, ... | | 著者 | Niu, W, Chen, Z, Bush-Pelc, L.A, Bah, A, Gandhi, P.S, Di Cera, E. | | 登録日 | 2009-09-22 | | 公開日 | 2009-10-20 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Mutant N143P reveals how Na+ activates thrombin
J.Biol.Chem., 284, 2009
|
|
3JZ2
 
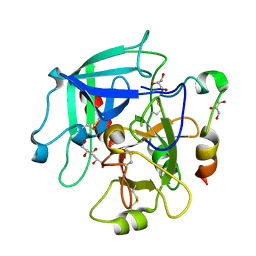 | | Crystal structure of human thrombin mutant N143P in E* form | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Thrombin heavy chain, ... | | 著者 | Niu, W, Chen, Z, Bush-Pelc, L.A, Bah, A, Gandhi, P.S, Di Cera, E. | | 登録日 | 2009-09-22 | | 公開日 | 2009-10-20 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Mutant N143P reveals how Na+ activates thrombin
J.Biol.Chem., 284, 2009
|
|
3S7K
 
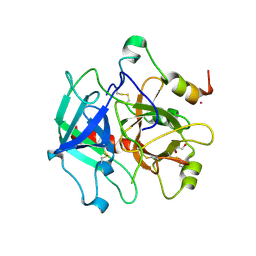 | | Structure of thrombin mutant Y225P in the E form | | 分子名称: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, POTASSIUM ION, Prothrombin | | 著者 | Niu, W, Chen, Z, Gandhi, P, Vogt, A, Pozzi, N, Pele, L.A, Zapata, F, Di Cera, E. | | 登録日 | 2011-05-26 | | 公開日 | 2011-07-06 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Crystallographic and Kinetic Evidence of Allostery in a Trypsin-like Protease.
Biochemistry, 50, 2011
|
|
3S7H
 
 | | Structure of thrombin mutant Y225P in the E* form | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Prothrombin | | 著者 | Niu, W, Chen, Z, Gandhi, P, Vogt, A, Pozzi, N, Pele, L.A, Zapata, F, Di Cera, E. | | 登録日 | 2011-05-26 | | 公開日 | 2011-07-06 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Crystallographic and Kinetic Evidence of Allostery in a Trypsin-like Protease.
Biochemistry, 50, 2011
|
|
3MVI
 
 | | Crystal structure of holo mADA at 1.6 A resolution | | 分子名称: | Adenosine deaminase, GLYCEROL, ZINC ION | | 著者 | Niu, W, Shu, Q, Chen, Z, Mathews, S, Di Cera, E, Frieden, C. | | 登録日 | 2010-05-04 | | 公開日 | 2010-11-03 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | The role of Zn2+ on the structure and stability of murine adenosine deaminase.
J.Phys.Chem.B, 114, 2010
|
|
3MVT
 
 | | Crystal structure of apo mADA at 2.2A resolution | | 分子名称: | Adenosine deaminase, CHLORIDE ION, GLYCEROL | | 著者 | Niu, W, Shu, Q, Chen, Z, Mathews, S, Di Cera, E, Frieden, C. | | 登録日 | 2010-05-04 | | 公開日 | 2010-10-13 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | The role of Zn2+ on the structure and stability of murine adenosine deaminase.
J.Phys.Chem.B, 114, 2010
|
|
4RN6
 
 | | Structure of prethrombin-2 mutant s195a bound to the active site inhibitor argatroban | | 分子名称: | (2R,4R)-4-methyl-1-(N~2~-{[(3S)-3-methyl-1,2,3,4-tetrahydroquinolin-8-yl]sulfonyl}-L-arginyl)piperidine-2-carboxylic acid, Thrombin heavy chain | | 著者 | Pozzi, N, Chen, Z, Zapata, F, Niu, W, Barranco-Medina, S, Pelc, L.A, Di Cera, E. | | 登録日 | 2014-10-23 | | 公開日 | 2014-11-05 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Autoactivation of thrombin precursors.
J.Biol.Chem., 288, 2013
|
|
4HZH
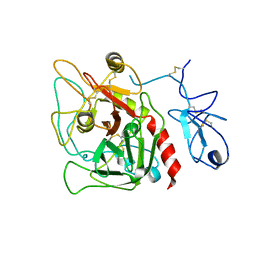 
 | | Structure of recombinant Gla-domainless prothrombin mutant S525A | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Prothrombin | | 著者 | Pozzi, N, Niu, W, Gohara, D.W, Chen, Z, Di Cera, E. | | 登録日 | 2012-11-15 | | 公開日 | 2013-06-26 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | Crystal structure of prothrombin reveals conformational flexibility and mechanism of activation.
J.Biol.Chem., 288, 2013
|
|
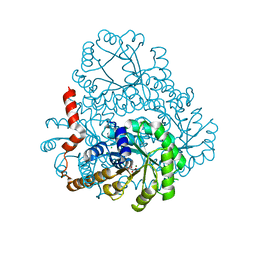
2DUA
 
 | |
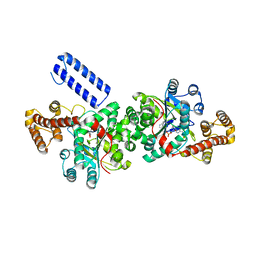
5UJJ
 
 | |
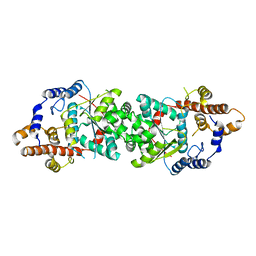
5UJI
 
 | |
2HRW
 
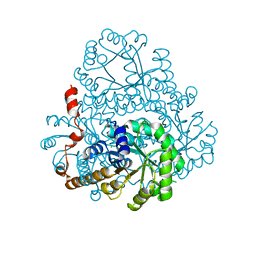 | | Crystal Structure of Phosphonopyruvate Hydrolase | | 分子名称: | CHLORIDE ION, Phosphonopyruvate hydrolase, SODIUM ION | | 著者 | Chen, C.C.H, Herzberg, O. | | 登録日 | 2006-07-20 | | 公開日 | 2006-10-03 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Structure and Kinetics of Phosphonopyruvate Hydrolase from Voriovorax sp. Pal2: New Insight into the Divergence of Catalysis within the PEP Mutase/Isocitrate Lyase Superfamily
Biochemistry, 45, 2006
|
|
2HJP
 
 | |
3B8I
 
 | |
6WN2
 
 | |
8TZX
 
 | | Ternary complex structure of Cereblon-DDB1 bound to WIZ(ZF7) and the molecular glue dWIZ-1 | | 分子名称: | (3S)-3-(5-{(1R)-1-[(2R)-1-ethylpiperidin-2-yl]ethoxy}-1-oxo-1,3-dihydro-2H-isoindol-2-yl)piperidine-2,6-dione, 1,2-ETHANEDIOL, DNA damage-binding protein 1, ... | | 著者 | Clifton, M.C, Ma, X, Ornelas, E. | | 登録日 | 2023-08-28 | | 公開日 | 2024-07-10 | | 最終更新日 | 2024-07-17 | | 実験手法 | X-RAY DIFFRACTION (3.15 Å) | | 主引用文献 | A molecular glue degrader of the WIZ transcription factor for fetal hemoglobin induction.
Science, 385, 2024
|
|
3FA4
 
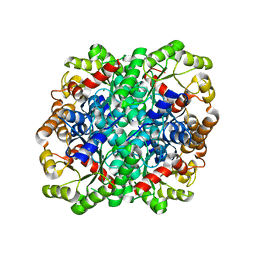 | | Crystal structure of 2,3-dimethylmalate lyase, a PEP mutase/isocitrate lyase superfamily member, triclinic crystal form | | 分子名称: | 2,3-dimethylmalate lyase, MAGNESIUM ION | | 著者 | Narayanan, B.C, Herzberg, O. | | 登録日 | 2008-11-14 | | 公開日 | 2009-01-27 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (2.18 Å) | | 主引用文献 | Structure and function of 2,3-dimethylmalate lyase, a PEP mutase/isocitrate lyase superfamily member.
J.Mol.Biol., 386, 2009
|
|
3FA3
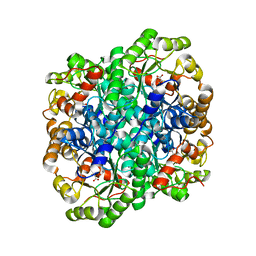 
 | | Crystal structure of 2,3-dimethylmalate lyase, a PEP mutase/isocitrate lyase superfamily member, trigonal crystal form | | 分子名称: | 2,2-difluoro-3,3-dihydroxybutanedioic acid, 2,3-dimethylmalate lyase, GLYCEROL, ... | | 著者 | Narayanan, B.C, Herzberg, O. | | 登録日 | 2008-11-14 | | 公開日 | 2009-01-27 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structure and function of 2,3-dimethylmalate lyase, a PEP mutase/isocitrate lyase superfamily member.
J.Mol.Biol., 386, 2009
|
|
7DAT
 
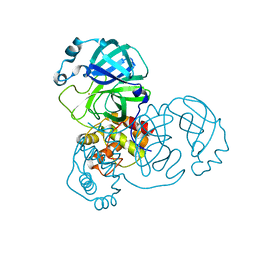 | | The crystal structure of COVID-19 main protease treated by AF | | 分子名称: | COVID-19 MAIN PROTEASE, GOLD ION | | 著者 | He, Z.S, He, B, Cao, P, Jiang, H.D, Gong, Y, Gao, X.Y. | | 登録日 | 2020-10-18 | | 公開日 | 2021-11-03 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.75 Å) | | 主引用文献 | A comparison of Remdesivir versus gold cluster in COVID-19 animal model: A better therapeutic outcome of gold cluster.
Nano Today, 44, 2022
|
|
7DAV
 
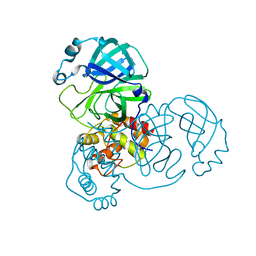 | | The native crystal structure of COVID-19 main protease | | 分子名称: | COVID-19 MAIN PROTEASE | | 著者 | He, Z.S, He, B, Cao, P, Jiang, H.D, Gong, Y, Gao, X.Y. | | 登録日 | 2020-10-18 | | 公開日 | 2021-11-03 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.77 Å) | | 主引用文献 | A comparison of Remdesivir versus gold cluster in COVID-19 animal model: A better therapeutic outcome of gold cluster.
Nano Today, 44, 2022
|
|
7DAU
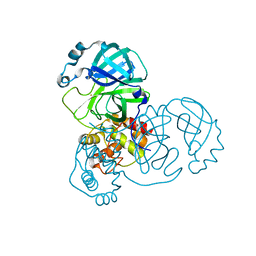 
 | | The crystal structure of COVID-19 main protease treated by GA | | 分子名称: | COVID-19 MAIN PROTEASE, GOLD ION | | 著者 | He, Z.S, He, B, Cao, P, Jiang, H.D, Gong, Y, Gao, X.Y. | | 登録日 | 2020-10-18 | | 公開日 | 2021-11-03 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.72 Å) | | 主引用文献 | A comparison of Remdesivir versus gold cluster in COVID-19 animal model: A better therapeutic outcome of gold cluster.
Nano Today, 44, 2022
|
|
4HFP
 
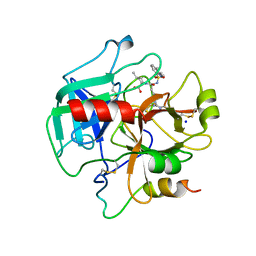 | | Structure of thrombin mutant S195a bound to the active site inhibitor argatroban | | 分子名称: | (2R,4R)-4-methyl-1-(N~2~-{[(3S)-3-methyl-1,2,3,4-tetrahydroquinolin-8-yl]sulfonyl}-L-arginyl)piperidine-2-carboxylic acid, Prothrombin, SODIUM ION | | 著者 | Pozzi, N, Chen, Z, Zapata, F, Lin, W, Barranco-Medina, S, Pelc, L.A, Di Cera, E. | | 登録日 | 2012-10-05 | | 公開日 | 2013-03-13 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Autoactivation of thrombin precursors.
J.Biol.Chem., 288, 2013
|
|
4H6T
 
 | | Crystal structure of prethrombin-2 mutant E14eA/D14lA/E18A/S195A | | 分子名称: | PHOSPHATE ION, Prothrombin | | 著者 | Pozzi, N, Chen, Z, Zapata, F, Pelc, L.A, Di Cera, E. | | 登録日 | 2012-09-19 | | 公開日 | 2013-03-13 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Autoactivation of thrombin precursors.
J.Biol.Chem., 288, 2013
|
|
4H6S
 
 | | Crystal structure of thrombin mutant E14eA/D14lA/E18A/S195A | | 分子名称: | Prothrombin, SODIUM ION | | 著者 | Pozzi, N, Chen, Z, Zapata, F, Pelc, L.A, Di Cera, E. | | 登録日 | 2012-09-19 | | 公開日 | 2013-03-13 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.19 Å) | | 主引用文献 | Autoactivation of thrombin precursors.
J.Biol.Chem., 288, 2013
|
|
