2CJS
 
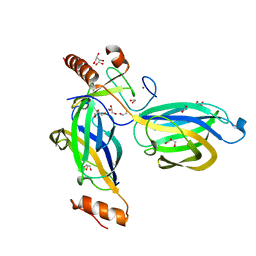 | | Structural Basis for a Munc13-1 Homodimer - Munc13-1 - RIM Heterodimer Switch: C2-domains as Versatile Protein-Protein Interaction Modules | | 分子名称: | 1,2-ETHANEDIOL, GLYCEROL, REGULATING SYNAPTIC MEMBRANE EXOCYTOSIS PROTEIN 2, ... | | 著者 | Lu, J, Machius, M, Dulubova, I, Dai, H, Sudhof, T.C, Tomchick, D.R, Rizo, J. | | 登録日 | 2006-04-06 | | 公開日 | 2006-06-07 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.78 Å) | | 主引用文献 | Structural Basis for a Munc13-1 Homodimer to Munc13-1/Rim Heterodimer Switch.
Plos Biol., 4, 2006
|
|
2F60
 
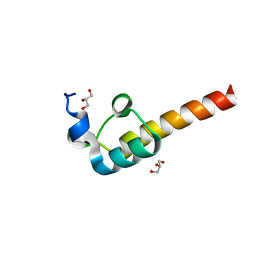 | | Crystal Structure of the Dihydrolipoamide Dehydrogenase (E3)-Binding Domain of Human E3-Binding Protein | | 分子名称: | GLYCEROL, Pyruvate dehydrogenase protein X component | | 著者 | Brautigam, C.A, Chuang, J.L, Wynn, R.M, Tomchick, D.R, Machius, M, Chuang, D.T. | | 登録日 | 2005-11-28 | | 公開日 | 2006-01-17 | | 最終更新日 | 2023-08-23 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | Structural Insight into Interactions between Dihydrolipoamide Dehydrogenase (E3) and E3 Binding Protein of Human Pyruvate Dehydrogenase Complex.
Structure, 14, 2006
|
|
2CJT
 
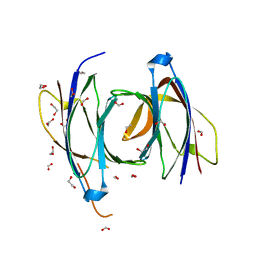 | | Structural Basis for a Munc13-1 Homodimer - Munc13-1 - RIM Heterodimer Switch: C2-domains as Versatile Protein-Protein Interaction Modules | | 分子名称: | 1,2-ETHANEDIOL, FORMIC ACID, UNC-13 HOMOLOG A | | 著者 | Lu, J, Machius, M, Dulubova, I, Dai, H, Sudhof, T.C, Tomchick, D.R, Rizo, J. | | 登録日 | 2006-04-06 | | 公開日 | 2006-06-07 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.44 Å) | | 主引用文献 | Structural Basis for a Munc13-1 Dimeric to Munc13-1/Rim Heterodimer Switch
Plos Biol., 4, 2006
|
|
2F5Z
 
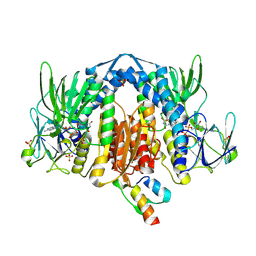 | | Crystal Structure of Human Dihydrolipoamide Dehydrogenase (E3) Complexed to the E3-Binding Domain of Human E3-Binding Protein | | 分子名称: | Dihydrolipoyl dehydrogenase, FLAVIN-ADENINE DINUCLEOTIDE, Pyruvate dehydrogenase protein X component, ... | | 著者 | Brautigam, C.A, Chuang, J.L, Wynn, R.M, Tomchick, D.R, Machius, M, Chuang, D.T. | | 登録日 | 2005-11-28 | | 公開日 | 2006-01-17 | | 最終更新日 | 2023-08-23 | | 実験手法 | X-RAY DIFFRACTION (2.18 Å) | | 主引用文献 | Structural Insight into Interactions between Dihydrolipoamide Dehydrogenase (E3) and E3 Binding Protein of Human Pyruvate Dehydrogenase Complex.
Structure, 14, 2006
|
|
2QYF
 
 | |
3BXE
 
 | |
7UCC
 
 | | Transcription factor FosB/JunD bZIP domain in the reduced form | | 分子名称: | CHLORIDE ION, ETHANOL, Protein fosB, ... | | 著者 | Kumar, A, Machius, M.C, Rudenko, G. | | 登録日 | 2022-03-16 | | 公開日 | 2023-01-25 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (1.94 Å) | | 主引用文献 | Chemically targeting the redox switch in AP1 transcription factor Delta FOSB.
Nucleic Acids Res., 50, 2022
|
|
7UCD
 | | Transcription factor FosB/JunD bZIP domain covalently modified with the cysteine-targeting alpha-haloketone compound Z2159931480 | | 分子名称: | 7-acetyl-4-methoxy-1-benzofuran-3(2H)-one, CHLORIDE ION, Protein fosB, ... | | 著者 | Kumar, A, Machius, M.C, Rudenko, G. | | 登録日 | 2022-03-16 | | 公開日 | 2023-01-25 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (3.21 Å) | | 主引用文献 | Chemically targeting the redox switch in AP1 transcription factor Delta FOSB.
Nucleic Acids Res., 50, 2022
|
|
3BEN
 
 | |
1HVX
 
 | | BACILLUS STEAROTHERMOPHILUS ALPHA-AMYLASE | | 分子名称: | ALPHA-AMYLASE, CALCIUM ION, SODIUM ION | | 著者 | Suvd, D, Fujimoto, Z, Takase, K, Matsumura, M, Mizuno, H. | | 登録日 | 2001-01-08 | | 公開日 | 2001-01-31 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Crystal structure of Bacillus stearothermophilus alpha-amylase: possible factors determining the thermostability.
J.Biochem., 129, 2001
|
|
4R8Q
 
 | |
5VPC
 | | Transcription factor FosB/JunD bZIP domain in its oxidized form, type-II crystal | | 分子名称: | CHLORIDE ION, Protein fosB, SODIUM ION, ... | | 著者 | Yin, Z, Machius, M.C, Rudenko, G. | | 登録日 | 2017-05-04 | | 公開日 | 2017-09-06 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.498 Å) | | 主引用文献 | Activator Protein-1: redox switch controlling structure and DNA-binding.
Nucleic Acids Res., 45, 2017
|
|
3OBV
 
 | |
3GMH
 
 | | Crystal Structure of the Mad2 Dimer | | 分子名称: | Mitotic spindle assembly checkpoint protein MAD2A, SULFATE ION | | 著者 | Ozkan, E, Luo, X, Machius, M, Yu, H, Deisenhofer, J. | | 登録日 | 2009-03-13 | | 公開日 | 2010-11-17 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (3.95 Å) | | 主引用文献 | Structure of an intermediate conformer of the spindle checkpoint protein Mad2.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
6MEP
 
 | | Crystal structure of the catalytic domain of the proto-oncogene tyrosine-protein kinase MER in complex with inhibitor UNC3437 | | 分子名称: | CHLORIDE ION, MAGNESIUM ION, Tyrosine-protein kinase Mer, ... | | 著者 | Da, C, Zhang, D, Stashko, M.A, Cheng, A, Hunter, D, Norris-Drouin, J, Graves, L, Machius, M, Miley, M.J, DeRyckere, D, Earp, H.S, Graham, D.K, Frye, S.V, Wang, X, Kireev, D. | | 登録日 | 2018-09-06 | | 公開日 | 2019-09-11 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.893 Å) | | 主引用文献 | Data-Driven Construction of Antitumor Agents with Controlled Polypharmacology.
J.Am.Chem.Soc., 141, 2019
|
|
5C7I
 
 | |
4OQQ
 
 | |
4OQP
 
 | |
3BXH
 
 | |
3BXF
 
 | | Crystal structure of effector binding domain of central glycolytic gene regulator (CggR) from Bacillus subtilis in complex with effector fructose-1,6-bisphosphate | | 分子名称: | 1,3-DIHYDROXYACETONEPHOSPHATE, 1,6-di-O-phosphono-beta-D-fructofuranose, CHLORIDE ION, ... | | 著者 | Rezacova, P, Otwinowski, Z. | | 登録日 | 2008-01-13 | | 公開日 | 2008-07-01 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Crystal structures of the effector-binding domain of repressor Central glycolytic gene Regulator from Bacillus subtilis reveal ligand-induced structural changes upon binding of several glycolytic intermediates.
Mol.Microbiol., 69, 2008
|
|
3BXG
 
 | |
2OKG
 
 | | Structure of effector binding domain of central glycolytic gene regulator (CggR) from B. subtilis | | 分子名称: | CHLORIDE ION, Central glycolytic gene regulator, GLYCERALDEHYDE-3-PHOSPHATE | | 著者 | Rezacova, P, Moy, S.F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | 登録日 | 2007-01-16 | | 公開日 | 2007-01-30 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Crystal structures of the effector-binding domain of repressor Central glycolytic gene Regulator from Bacillus subtilis reveal ligand-induced structural changes upon binding of several glycolytic intermediates.
Mol.Microbiol., 69, 2008
|
|
3EDV
 | |
3G59
 
 | |
3FWK
 
 | |
