2G3M
 
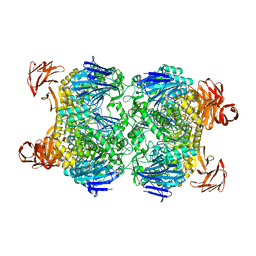 | | Crystal structure of the Sulfolobus solfataricus alpha-glucosidase MalA | | Descriptor: | Alpha-glucosidase | | Authors: | Ernst, H.A, Lo Leggio, L, Willemoes, M, Leonard, G, Blum, P, Larsen, S. | | Deposit date: | 2006-02-20 | | Release date: | 2006-05-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structure of the Sulfolobus solfataricus alpha-Glucosidase: Implications for Domain Conservation and Substrate Recognition in GH31.
J.Mol.Biol., 358, 2006
|
|
2G3N
 
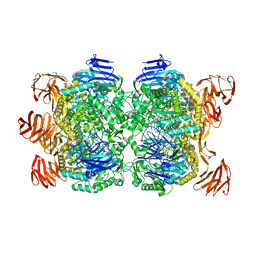 | | Crystal structure of the Sulfolobus solfataricus alpha-glucosidase MalA in complex with beta-octyl-glucopyranoside | | Descriptor: | Alpha-glucosidase, octyl beta-D-glucopyranoside | | Authors: | Ernst, H.A, Lo Leggio, L, Willemoes, M, Leonard, G, Blum, P, Larsen, S. | | Deposit date: | 2006-02-20 | | Release date: | 2006-05-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structure of the Sulfolobus solfataricus alpha-Glucosidase: Implications for Domain Conservation and Substrate Recognition in GH31.
J.Mol.Biol., 358, 2006
|
|
4B7W
 
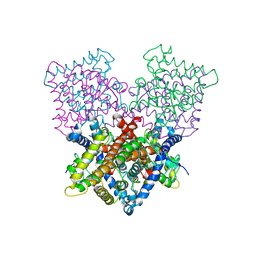 | | Ligand binding domain human hepatocyte nuclear factor 4alpha: Apo form | | Descriptor: | HEPATOCYTE NUCLEAR FACTOR 4-ALPHA | | Authors: | Dudasova, Z, Okvist, M, Kretova, M, Ondrovicova, G, Skrabana, R, LeGuevel, R, Salbert, G, Leonard, G, McSweeney, S, Barath, P. | | Deposit date: | 2012-08-24 | | Release date: | 2013-09-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Fatty Acids are not Essential Structural Components of Hepatocyte Nuclear Factor 4Alpha
To be Published
|
|
5NX9
 
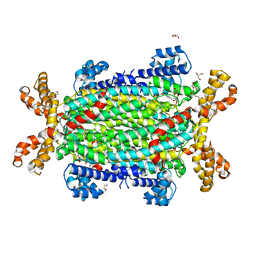 | | Crystal structure of Neanderthal Adenylosuccinate Lyase (ADSL) in complex with its products AMP and fumarate | | Descriptor: | 2-[9-(3,4-DIHYDROXY-5-PHOSPHONOOXYMETHYL-TETRAHYDRO-FURAN-2-YL)-9H-PURIN-6-YLAMINO]-SUCCINIC ACID, ADENOSINE MONOPHOSPHATE, Adenylosuccinate lyase, ... | | Authors: | Van Laer, B, Kapp, U, Soler-Lopez, M, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2017-05-09 | | Release date: | 2018-05-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular comparison of Neanderthal and Modern Human adenylosuccinate lyase.
Sci Rep, 8, 2018
|
|
5NXA
 
 | | Crystal structure of Neanderthal Adenylosuccinate Lyase (ADSL)in complex with its products AICAR and fumarate | | Descriptor: | AMINOIMIDAZOLE 4-CARBOXAMIDE RIBONUCLEOTIDE, Adenylosuccinate lyase, CHLORIDE ION, ... | | Authors: | Van Laer, B, Kapp, U, Soler-Lopez, M, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2017-05-09 | | Release date: | 2018-05-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Molecular comparison of Neanderthal and Modern Human adenylosuccinate lyase.
Sci Rep, 8, 2018
|
|
5NX8
 
 | | Crystal structure of Neanderthal Adenylosuccinate Lyase (ADSL) | | Descriptor: | Adenylosuccinate lyase, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Van Laer, B, Kapp, U, Soler-Lopez, M, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2017-05-09 | | Release date: | 2018-05-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Molecular comparison of Neanderthal and Modern Human adenylosuccinate lyase.
Sci Rep, 8, 2018
|
|
1EFC
 
 | | INTACT ELONGATION FACTOR FROM E.COLI | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, PROTEIN (ELONGATION FACTOR) | | Authors: | Song, H, Parsons, M.R, Rowsell, S, Leonard, G, Phillips, S.E.V. | | Deposit date: | 1998-11-24 | | Release date: | 1999-03-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structure of intact elongation factor EF-Tu from Escherichia coli in GDP conformation at 2.05 A resolution.
J.Mol.Biol., 285, 1999
|
|
2VF7
 
 | | Crystal structure of UvrA2 from Deinococcus radiodurans | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, EXCINUCLEASE ABC, SUBUNIT A., ... | | Authors: | Timmins, J, Gordon, E, Caria, S, Leonard, G, Kuo, M.S, Monchois, V, McSweeney, S. | | Deposit date: | 2007-10-31 | | Release date: | 2008-12-16 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and mutational analyses of Deinococcus radiodurans UvrA2 provide insight into DNA binding and damage recognition by UvrAs.
Structure, 17, 2009
|
|
2VF8
 
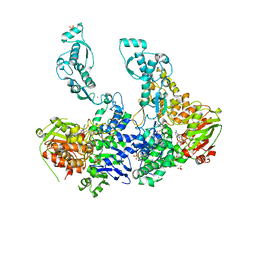 | | Crystal structure of UvrA2 from Deinococcus radiodurans | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, EXCINUCLEASE ABC SUBUNIT A, PHOSPHATE ION, ... | | Authors: | Timmins, J, Gordon, E, Caria, S, Leonard, G, Kuo, M.S, Monchois, V, McSweeney, S. | | Deposit date: | 2007-10-31 | | Release date: | 2008-12-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and Mutational Analyses of Deinococcus Radiodurans Uvra2 Provide Insight Into DNA Binding and Damage Recognition by Uvras.
Structure, 17, 2009
|
|
2XRH
 
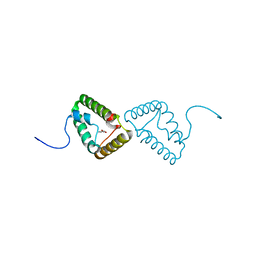 | | Crystal structure of the truncated form of HP0721 | | Descriptor: | NICOTINIC ACID, PROTEIN HP0721 | | Authors: | Cioci, G, Terradot, L, Dian, C, Muller-Dieckmann, C, Leonard, G. | | Deposit date: | 2010-09-15 | | Release date: | 2011-09-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of Hp0721, a Novel Secreted Protein from Helicobacter Pylori.
Proteins, 79, 2011
|
|
2BHV
 
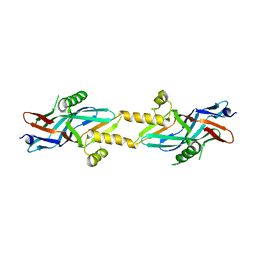 | | Structure of ComB10 of the Com Type IV secretion system of Helicobacter pylori | | Descriptor: | COMB10 | | Authors: | Terradot, L, Oomen, C, Bayliss, R, Leonard, G, Waksman, G. | | Deposit date: | 2005-01-18 | | Release date: | 2005-03-16 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structures of Two Core Subunits of the Bacterial Type Iv Secretion System, Virb8 from Brucella Suis and Comb10 from Helicobacter Pylori
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
6I34
 
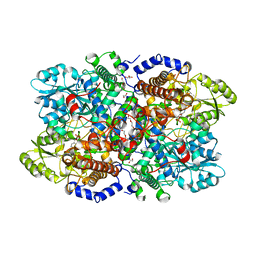 | | Crystal structure of Neanderthal glycine decarboxylase (P-protein) | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Van Laer, B, Kapp, U, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2018-11-05 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural insights in human glycine decarboxylase and comparison with the Neanderthal variant
To Be Published
|
|
6I33
 
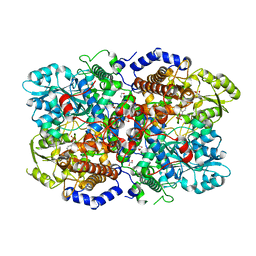 | | Crystal structure of human glycine decarboxylase (P-protein) | | Descriptor: | 1,2-ETHANEDIOL, BICARBONATE ION, Glycine dehydrogenase (decarboxylating), ... | | Authors: | Van Laer, B, Kapp, U, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2018-11-05 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural insights in human glycine decarboxylase and comparison with the Neanderthal variant
To Be Published
|
|
6I35
 
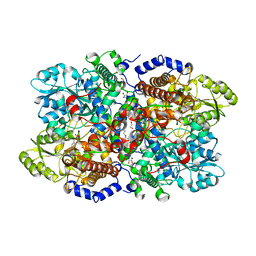 | | Crystal structure of human glycine decarboxylase (P-protein) bound with pyridoxyl-glycine-5'-monophosphate | | Descriptor: | 1,2-ETHANEDIOL, BICARBONATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Van Laer, B, Kapp, U, Leonard, G, Mueller-Dieckmann, C. | | Deposit date: | 2018-11-05 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights in human glycine decarboxylase and comparison with the Neanderthal variant
To Be Published
|
|
1USP
 
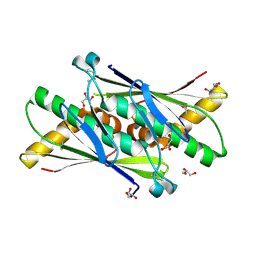 | | Organic Hydroperoxide Resistance Protein from Deinococcus radiodurans | | Descriptor: | GLYCEROL, ORGANIC HYDROPEROXIDE RESISTANCE PROTEIN | | Authors: | Meunier-Jamin, C, Kapp, U, Leonard, G, McSweeney, S. | | Deposit date: | 2003-11-27 | | Release date: | 2004-04-08 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Structure of the Organic Hydroperoxide Resistance Protein from Deinococcus Radiodurans: Do Conformational Changes Facilitate Recycling of the Redox Disulfide?
J.Biol.Chem., 279, 2004
|
|
1DZ3
 
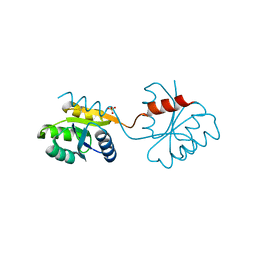 | | DOMAIN-SWAPPING IN THE SPORULATION RESPONSE REGULATOR SPO0A | | Descriptor: | SULFATE ION, Stage 0 sporulation protein A | | Authors: | Lewis, R.J, Brannigan, J.A, Muchova, K, Leonard, G, Barak, I, Wilkinson, A.J. | | Deposit date: | 2000-02-15 | | Release date: | 2000-04-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Domain swapping in the sporulation response regulator Spo0A.
J. Mol. Biol., 297, 2000
|
|
2BHY
 
 | | Crystal structure of Deinococcus radiodurans maltooligosyltrehalose trehalohydrolase in complex with trehalose | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BETA-MERCAPTOETHANOL, MAGNESIUM ION, ... | | Authors: | Timmins, J, Leiros, H.-K.S, Leonard, G, Leiros, I, McSweeney, S. | | Deposit date: | 2005-01-20 | | Release date: | 2005-03-31 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of Maltooligosyltrehalose Trehalohydrolase from Deinococcus Radiodurans in Complex with Disaccharides
J.Mol.Biol., 347, 2005
|
|
2BHU
 
 | | Crystal structure of Deinococcus radiodurans maltooligosyltrehalose trehalohydrolase | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MAGNESIUM ION, MALTOOLIGOSYLTREHALOSE TREHALOHYDROLASE, ... | | Authors: | Timmins, J, Leiros, H.-K.S, Leonard, G, Leiros, I, McSweeney, S. | | Deposit date: | 2005-01-18 | | Release date: | 2005-03-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal Structure of Maltooligosyltrehalose Trehalohydrolase from Deinococcus Radiodurans in Complex with Disaccharides
J.Mol.Biol., 347, 2005
|
|
2BHZ
 
 | | Crystal structure of Deinococcus radiodurans maltooligosyltrehalose trehalohydrolase in complex with maltose | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BETA-MERCAPTOETHANOL, MAGNESIUM ION, ... | | Authors: | Timmins, J, Leiros, H.-K.S, Leonard, G, Leiros, I, McSweeney, S. | | Deposit date: | 2005-01-20 | | Release date: | 2005-03-31 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal Structure of Maltooligosyltrehalose Trehalohydrolase from Deinococcus Radiodurans in Complex with Disaccharides
J.Mol.Biol., 347, 2005
|
|
1FSE
 
 | | CRYSTAL STRUCTURE OF THE BACILLUS SUBTILIS REGULATORY PROTEIN GERE | | Descriptor: | GERE, GLYCEROL, SULFATE ION | | Authors: | Ducros, V.M.-A, Lewis, R.J, Verma, C.S, Dodson, E.J, Leonard, G, Turkenburg, J.P, Murshudov, G.N, Wilkinson, A.J, Brannigan, J.A. | | Deposit date: | 2000-09-08 | | Release date: | 2001-03-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structure of GerE, the ultimate transcriptional regulator of spore formation in Bacillus subtilis.
J.Mol.Biol., 306, 2001
|
|
1KBZ
 
 | | Crystal Structure of apo-dTDP-6-deoxy-L-lyxo-4-hexulose reductase (RmlD) from Salmonella enterica serovar Typhimurium | | Descriptor: | MAGNESIUM ION, dTDP-glucose oxidoreductase | | Authors: | Blankenfeldt, W, Kerr, I.D, Giraud, M.F, McMiken, H.J, Leonard, G.A, Whitfield, C, Messner, P, Graninger, M, Naismith, J.H. | | Deposit date: | 2001-11-07 | | Release date: | 2002-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Variation on a theme of SDR. dTDP-6-deoxy-L- lyxo-4-hexulose reductase (RmlD) shows a new Mg2+-dependent dimerization mode.
Structure, 10, 2002
|
|
1KC3
 
 | | Crystal structure of dTDP-6-deoxy-L-lyxo-4-hexulose reductase (RmlD) in complex with NADPH and dTDP-L-rhamnose | | Descriptor: | 2'-DEOXY-THYMIDINE-BETA-L-RHAMNOSE, MAGNESIUM ION, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Blankenfeldt, W, Kerr, I.D, Giraud, M.F, McMiken, H.J, Leonard, G.A, Whitfield, C, Messner, P, Graninger, M, Naismith, J.H. | | Deposit date: | 2001-11-07 | | Release date: | 2002-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Variation on a theme of SDR. dTDP-6-deoxy-L- lyxo-4-hexulose reductase (RmlD) shows a new Mg2+-dependent dimerization mode.
Structure, 10, 2002
|
|
1KC1
 
 | | Crystal structure of dTDP-6-deoxy-L-lyxo-4-hexulose reductase (RmlD) in complex with NADPH | | Descriptor: | MAGNESIUM ION, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION, ... | | Authors: | Blankenfeldt, W, Kerr, I.D, Giraud, M.F, McMiken, H.J, Leonard, G.A, Whitfield, C, Messner, P, Graninger, M, Naismith, J.H. | | Deposit date: | 2001-11-07 | | Release date: | 2002-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Variation on a theme of SDR. dTDP-6-deoxy-L- lyxo-4-hexulose reductase (RmlD) shows a new Mg2+-dependent dimerization mode.
Structure, 10, 2002
|
|
6QQK
 | |
6QQI
 | |
