7C7F
 
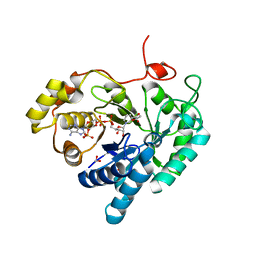 | | Crystal structures of AKR1C3 binary complex with NADP+ | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Aldo-keto reductase family 1 member C3, ... | | Authors: | Irie, K, Toyooka, N, Endo, S. | | Deposit date: | 2020-05-25 | | Release date: | 2020-09-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Development of Novel AKR1C3 Inhibitors as New Potential Treatment for Castration-Resistant Prostate Cancer.
J.Med.Chem., 63, 2020
|
|
7C7G
 
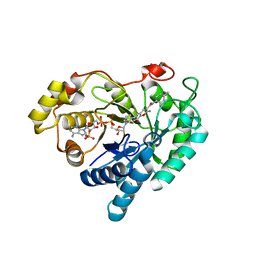 | |
7C7H
 
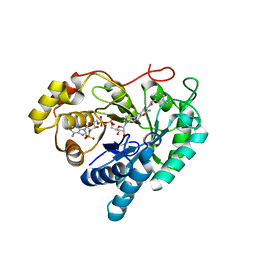 | |
6K9I
 
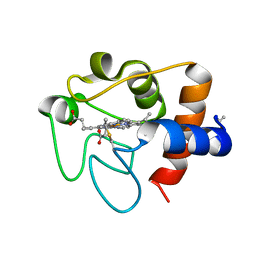 | |
6K9J
 
 | |
1X1P
 
 | | Crystal structure of Tk-RNase HII(1-197)-A(28-42) | | Descriptor: | Ribonuclease HII | | Authors: | Takano, K, Endo, S, Mukaiyama, A, Chon, H, Matsumura, H, Koga, Y, Kanaya, S. | | Deposit date: | 2005-04-11 | | Release date: | 2006-01-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of amyloid beta fragments in aqueous environments
Febs J., 273, 2006
|
|
2DDE
 
 | | Structure of cinnamycin complexed with lysophosphatidylethanolamine | | Descriptor: | (7S)-4,7-DIHYDROXY-10-OXO-3,5,9-TRIOXA-4-PHOSPHAUNDECAN-1-AMINIUM 4-OXIDE, LANTIBIOTIC CINNAMYCIN | | Authors: | Hosoda, K, Ohya, M, Kohno, T, Maeda, T, Endo, S, Wakamatsu, K. | | Deposit date: | 2006-01-27 | | Release date: | 2006-02-21 | | Last modified: | 2024-11-06 | | Method: | SOLUTION NMR | | Cite: | Structure determination of an immunopotentiator peptide, cinnamycin, complexed with lysophosphatidylethanolamine by 1H-NMR1.
J.Biochem., 119, 1996
|
|
1GEA
 
 | | RECEPTOR-BOUND CONFORMATION OF PACAP21 | | Descriptor: | PITUITARY ADENYLATE CYCLASE ACTIVATING POLYPEPTIDE | | Authors: | Inooka, H, Ohtaki, T, Kitahara, O, Ikegami, T, Endo, S, Kitada, C, Ogi, K, Onda, H, Fujino, M, Shirakawa, M. | | Deposit date: | 2000-10-20 | | Release date: | 2001-04-20 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Conformation of a peptide ligand bound to its G-protein coupled receptor.
Nat.Struct.Biol., 8, 2001
|
|
6CKX
 
 | |
7K41
 
 | | Bacterial O-GlcNAcase (OGA) with compound | | Descriptor: | 1,2-ETHANEDIOL, 4-(4-methylpiperidin-1-yl)-N-(2-phenylethyl)pyrimidin-2-amine, ACETATE ION, ... | | Authors: | Lane, W, Tjhen, R, Snell, G, Sang, B. | | Deposit date: | 2020-09-14 | | Release date: | 2021-01-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of a Novel and Brain-Penetrant O -GlcNAcase Inhibitor via Virtual Screening, Structure-Based Analysis, and Rational Lead Optimization.
J.Med.Chem., 64, 2021
|
|
2P5N
 
 | | Crystal structure of mouse 17-alpha hydroxysteroid dehydrogenase in complex with coenzyme NADPH | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Aldo-keto reductase family 1, member C21, ... | | Authors: | El-Kabbani, O, Dhagat, U. | | Deposit date: | 2007-03-15 | | Release date: | 2007-10-09 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of 3(17)alpha-hydroxysteroid dehydrogenase (AKR1C21) holoenzyme from an orthorhombic crystal form: an insight into the bifunctionality of the enzyme.
Acta Crystallogr.,Sect.F, 63, 2007
|
|
2O4U
 
 | | Crystal structure of Mammalian Dimeric Dihydrodiol Dehydrogenase | | Descriptor: | BETA-MERCAPTOETHANOL, Dimeric dihydrodiol dehydrogenase, GLYCEROL, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2006-12-04 | | Release date: | 2007-07-31 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of dimeric dihydrodiol dehydrogenase apoenzyme and inhibitor complex: probing the subunit interface with site-directed mutagenesis.
Proteins, 70, 2008
|
|
2O48
 
 | | Crystal structure of Mammalian Dimeric Dihydrodiol Dehydrogenase | | Descriptor: | BETA-MERCAPTOETHANOL, Dimeric dihydrodiol dehydrogenase, P-HYDROXYACETOPHENONE, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2006-12-04 | | Release date: | 2007-07-31 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Structures of dimeric dihydrodiol dehydrogenase apoenzyme and inhibitor complex: probing the subunit interface with site-directed mutagenesis.
Proteins, 70, 2008
|
|
3WFF
 
 | | Mineralocorticoid receptor ligand-binding domain with compound 2b | | Descriptor: | 6-[4-(2,4-difluorophenyl)-5-oxo-2,5-dihydrofuran-3-yl]-2H-1,4-benzoxazin-3(4H)-one, Mineralocorticoid receptor, PHOSPHATE ION | | Authors: | Sogabe, S, Habuka, N. | | Deposit date: | 2013-07-19 | | Release date: | 2013-08-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Design, synthesis, and structure-activity relationships of dihydrofuran-2-one and dihydropyrrol-2-one derivatives as novel benzoxazin-3-one-based mineralocorticoid receptor antagonists.
Bioorg.Med.Chem., 21, 2013
|
|
3WFG
 
 | | Mineralocorticoid receptor ligand-binding domain with compuond 2e | | Descriptor: | 1,2-ETHANEDIOL, 6-[(2S)-4-(4-fluorophenyl)-2-methyl-5-oxo-2,5-dihydrofuran-3-yl]-2H-1,4-benzoxazin-3(4H)-one, Mineralocorticoid receptor | | Authors: | Sogabe, S, Habuka, N. | | Deposit date: | 2013-07-19 | | Release date: | 2013-08-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Design, synthesis, and structure-activity relationships of dihydrofuran-2-one and dihydropyrrol-2-one derivatives as novel benzoxazin-3-one-based mineralocorticoid receptor antagonists.
Bioorg.Med.Chem., 21, 2013
|
|
3C3U
 
 | | Crystal structure of AKR1C1 in complex with NADP and 3,5-dichlorosalicylic acid | | Descriptor: | 3,5-dichloro-2-hydroxybenzoic acid, Aldo-keto reductase family 1 member C1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Dhagat, U, El-Kabbani, O. | | Deposit date: | 2008-01-28 | | Release date: | 2008-08-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Selectivity determinants of inhibitor binding to human 20alpha-hydroxysteroid dehydrogenase: crystal structure of the enzyme in ternary complex with coenzyme and the potent inhibitor 3,5-dichlorosalicylic acid
J.Med.Chem., 51, 2008
|
|
3CV6
 
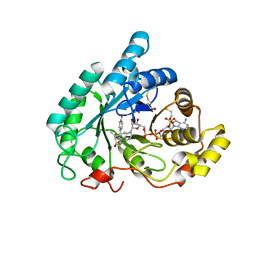 | | The crystal structure of mouse 17-alpha hydroxysteroid dehydrogenase GG225.226PP mutant in complex with inhibitor and cofactor NADP+. | | Descriptor: | 4-[(1R,2S)-1-ethyl-2-(4-hydroxyphenyl)butyl]phenol, Aldo-keto reductase family 1 member C21, BETA-MERCAPTOETHANOL, ... | | Authors: | Dhagat, U, El-Kabbani, O. | | Deposit date: | 2008-04-17 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the G225P/G226P mutant of mouse 3(17)alpha-hydroxysteroid dehydrogenase (AKR1C21) ternary complex: implications for the binding of inhibitor and substrate.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
4PF3
 
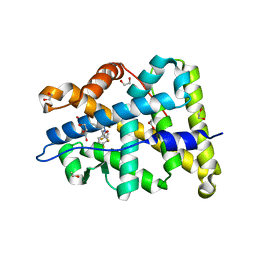 | | Mineralocorticoid receptor ligand-binding domain with compuond 37a | | Descriptor: | 1,2-ETHANEDIOL, 6-[1-(2,2-difluoro-3-hydroxypropyl)-5-(4-fluorophenyl)-3-methyl-1H-pyrazol-4-yl]-2H-1,4-benzoxazin-3(4H)-one, Mineralocorticoid receptor | | Authors: | Sogabe, S, Habuka, N. | | Deposit date: | 2014-04-28 | | Release date: | 2014-11-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Discovery of 6-[5-(4-fluorophenyl)-3-methyl-pyrazol-4-yl]-benzoxazin-3-one derivatives as novel selective nonsteroidal mineralocorticoid receptor antagonists
Bioorg.Med.Chem., 22, 2014
|
|
2D1Y
 
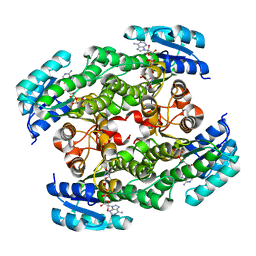 | |
5URM
 
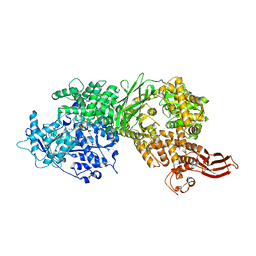 | | Crystal structure of human BRR2 in complex with T-1206548 | | Descriptor: | 3-(5-{[(2R)-5-amino-2-cyclohexyl-7-oxo-2,3-dihydro-7H-[1,3,4]thiadiazolo[3,2-a]pyrimidin-6-yl]methyl}furan-2-yl)benzoic acid, U5 small nuclear ribonucleoprotein 200 kDa helicase | | Authors: | Klein, M.G, Tjhen, R, Qin, L. | | Deposit date: | 2017-02-11 | | Release date: | 2017-07-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Discovery of Allosteric Inhibitors Targeting the Spliceosomal RNA Helicase Brr2.
J. Med. Chem., 60, 2017
|
|
5URJ
 
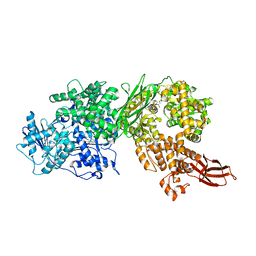 | | Crystal structure of human BRR2 in complex with T-3905516 | | Descriptor: | 6-benzyl-3-[(2R)-2-(3-fluoropyridin-2-yl)-6-methyl-3,4-dihydro-2H-1-benzopyran-7-yl]-4,6-dihydropyrido[4,3-d]pyrimidine-2,7(3H,8H)-dione, GLYCEROL, U5 small nuclear ribonucleoprotein 200 kDa helicase | | Authors: | Klein, M.G, Tjhen, R, Qin, L. | | Deposit date: | 2017-02-10 | | Release date: | 2017-07-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Discovery of Allosteric Inhibitors Targeting the Spliceosomal RNA Helicase Brr2.
J. Med. Chem., 60, 2017
|
|
5URK
 
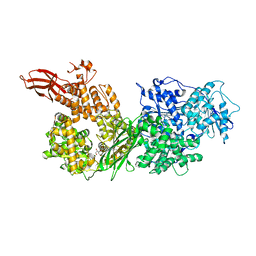 | | Crystal structure of human BRR2 in complex with T-3935799 | | Descriptor: | 6-benzyl-3-[3-(benzyloxy)phenyl]-4,6-dihydropyrido[4,3-d]pyrimidine-2,7(1H,3H)-dione, GLYCEROL, U5 small nuclear ribonucleoprotein 200 kDa helicase | | Authors: | Qin, L, Tjhen, R, Klein, M.G. | | Deposit date: | 2017-02-10 | | Release date: | 2017-07-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Discovery of Allosteric Inhibitors Targeting the Spliceosomal RNA Helicase Brr2.
J. Med. Chem., 60, 2017
|
|
3D3W
 
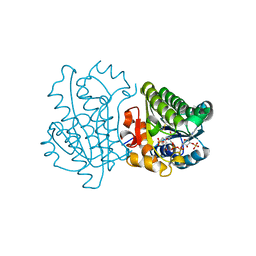 | |
1RCH
 
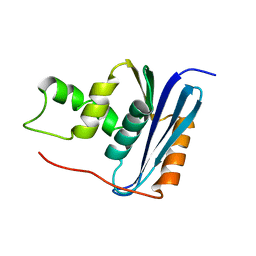 | | SOLUTION NMR STRUCTURE OF RIBONUCLEASE HI FROM ESCHERICHIA COLI, 8 STRUCTURES | | Descriptor: | RIBONUCLEASE HI | | Authors: | Yamazaki, T, Fujiwara, M, Kato, T, Yamasaki, K, Nagayama, K. | | Deposit date: | 1995-06-23 | | Release date: | 1997-02-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Ribonuclease Hi from Escherichia Coli
Biol.Pharm.Bull., 23, 2000
|
|
3CV7
 
 | | Crystal structure of porcine aldehyde reductase ternary complex | | Descriptor: | 3,5-dichloro-2-hydroxybenzoic acid, Alcohol dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2008-04-18 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.412 Å) | | Cite: | Structure of aldehyde reductase in ternary complex with coenzyme and the potent 20alpha-hydroxysteroid dehydrogenase inhibitor 3,5-dichlorosalicylic acid: Implications for inhibitor binding and selectivity
Arch.Biochem.Biophys., 479, 2008
|
|
